Abstract
Abstract 2537
MLL gene rearrangements are the most common genetic events in infant leukemia. Up to date more than 100 various MLL rearrangements were described.
To evaluate the distribution of MLL rearrangements among infants (aged from 1 to 365 days) with both acute lymphoblastic leukemia (ALL) and acute myeloid leukemia (AML).
174 infants (117 ALL and 57 AML cases) were included in the current study. 11q23/MLL rearrangements were detected by chromosome banding analysis (CBA), fluorescence in-situ hybridization (FISH) and reverse-transcriptase PCR (RT-PCR). CBA was done according to standard procedure. FISH analysis using LSI MLL Dual Color, Break Apart Rearrangement Probe (Abbott Molecular, USA) was performed on at least 200 interphase nuclei and on all available metaphases. RT-PCR was performed as previously described (A. Borkhardt et al.,1994, N. Palisgaard et al., 1998, J. van Dongen et al., 1999). In 39 cases genomic DNA breakpoint was detected in MLL and translocation partner genes by long-distance inverse PCR (LDI-PCR). Exon-intron numbering of MLL gene was done according to I. Nilson et al, 1996.
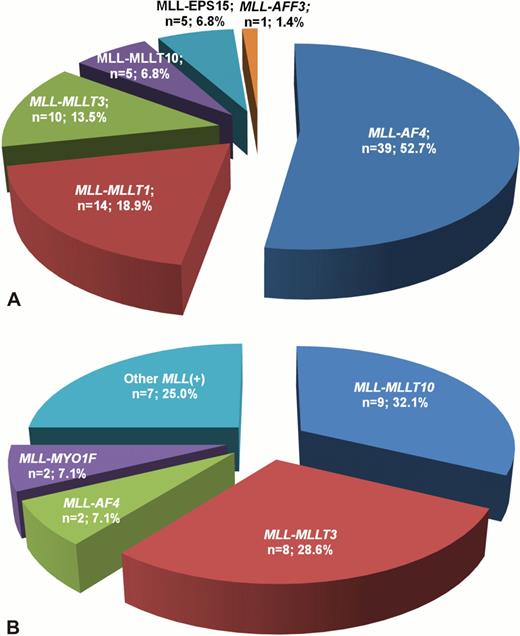
11q23/MLL rearrangements were revealed in 74 ALL patients (63.2%). Among this group MLL-AF4 was detected in the majority of cases (53.5%), less frequently were found MLL-MLLT1, MLL-MLLT3, MLL-MLLT10 and others (fig. 1a). Children with ALL under 6 months of age had significantly higher incidence of MLL rearrangements in comparison with older infants (84.0% vs. 47.8%, p<0.001). MLL-positive patients more frequently had BI-ALL and less frequently BII-ALL than infants without these rearrangements (p<0.001 for both). Fusion gene transcripts were sequenced in 26 MLL-rearranged ALL cases. Depending on breakpoint position within MLL and partner genes we detected 7 different types of MLL-AF4 fusion gene transcript, 3 types of MLL-MLLT1, 2 types of MLL-EPS15. The most common fusion site within MLL gene in ALL patients was exon 11, detected in 14 cases (53.8%). It was confirmed by LDI-PCR, that in addition to common breakpoint location in MLL gene (18 out of 27 cases in intron 11, 4 cases in intron 9) allowed to reveal less frequent breakpoint sites, like intron 12 (1 case), intron 10 (3 cases) and intron 7 (1 case). Interestingly, in the last case where LDI-PCR showed presence of MLL-AF4, this fusion gene transcript was not initially found by RT-PCR, because applied primer set did not cover exon 7. Moreover, due to lack of metaphases this patient was primary misclassified as MLL-rearranged, but MLL-AF4-negative. MLL rearrangements were found in 28 AML cases (49.1%). In AML patients the most common MLL rearrangements were MLL-MLLT10 (32% of cases) and MLL-MLLT3 (28%). Other ones were detected less frequently (fig. 1b). In AML patients frequency of MLL rearrangements was similar in children younger and older than 6 months (p=0.904). Among MLL-positive cases AML M5 were detected significantly more often and AML M7 significantly less frequent than in MLL-negative patients (p=0.024 and p=0.001, correspondingly). The most common breakpoint location within MLL gene in AML patients was intron 9, detected in 6 out of 12 cases (50%). Additional chromosomal abnormalities were revealed in 7 out of 21 MLL-positive AML patients with known karyotype (33%), while complex karyotype was detected in 5 cases (24%). Application of LDI-PCR allowed to verify rare MLL rearrangements, including MLL-AFF3 (1 ALL case), MLL-MYO1F (2 AML cases), MLL-SEPT6 (1 AML case), MLL-SEPT9 (1 AML case) In 4 ALL and 3 AML patients MLL rearrangements with concurrent 3'-deletion of MLL gene were found. 3'-deletion of MLL was not associated with breakpoint position in MLL gene and type of translocation partner gene. None of the patients with 3'-deletions had reciprocal fusion gene. Based on LDI-PCR data we assessed several mechanisms of fusion gene formation. Reciprocal translocations were detected in 29 cases, 3-way translocations in 3 cases, inversions in 5 cases, combination of inversion and insertion in 2 cases.
In the current study we precisely characterized large cohort of MLL-rearranged infant acute leukemia patients. Combination of all available techniques, including cytogenetics, FISH, RT-PCR and LDI-PCR can lead to detailed verification of every single MLL rearrangement.
Distribution of different MLL fusion partner genes in infant ALL (A) and AML (B) cases.
Distribution of different MLL fusion partner genes in infant ALL (A) and AML (B) cases.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.