Abstract
Wiskott Aldrich syndrome (WAS), an X-linked immunodeficiency, results from loss-of-function mutations in the human hematopoietic cytoskeletal regulator gene WAS. Many missense mutations in the Ena Vasp homology1 (EVH1) domain preserve low-level WAS protein (WASp) expression and confer a milder clinical phenotype. Although disrupted binding to WASp-interacting protein (WIP) leads to enhanced WASp degradation in vivo, the intrinsic function of EVH1-mutated WASp is poorly understood. In the present study, we show that, despite mediating enhanced actin polymerization compared with wild-type WASp in vitro, EVH1 missense mutated proteins did not support full biologic function in cells, even when levels were restored by forced overexpression. Podosome assembly was aberrant and associated with dysregulated lamellipodia formation and impaired persistence of migration. At sites of residual podosome-associated actin polymerization, localization of EVH1-mutated proteins was preserved even after deletion of the entire domain, implying that WIP-WASp complex formation is not absolutely required for WASp localization. However, retention of mutant proteins in podosomes was significantly impaired and associated with reduced levels of WASp tyrosine phosphorylation. Our results indicate that the EVH1 domain is important not only for WASp stability, but also for intrinsic biologic activity in vivo.
Key Points
Impaired WASp-WIP binding causes dysregulated WASp activity and reduced WASp retention in podosomes resulting in ineffective cell migration.
The intrinsic function of EVH1 mutant WASp is defective. Therapeutic strategies to increase patient mutant protein levels are likely to fail.
Introduction
Wiskott Aldrich syndrome (WAS) is a primary immunodeficiency characterized by a broad immune defect, thrombocytopenia, severe eczema, and an increased susceptibility to both autoimmunity and hematologic malignancy.1 The disease is caused by mutations in the gene encoding the WAS protein (WASp), a key regulator of the actin cytoskeleton in hematopoietic cells.2,3 Cytosolic WASp exists in an autoinhibited conformation promoted by constitutive binding of the EVH1 domain to WASp Interacting Protein (WIP).4-8 Activation induces a conformational change exposing the VCA (Verprolin homology, Central region, Acidic domain) domain, which binds the Arp2/3 complex,9,10 initiating actin filament nucleation and polymerization.11 WASp activity is regulated through the combined influence of many binding partners and posttranslational modification, of which the Rho GTPase Cdc42 and tyrosine phosphorylation have particular importance.1
Mutations resulting in complete loss of WASp expression are predictive of severe, classic WAS while missense mutations in the EVH1 domain generally result in residual low-level WASp expression and cause milder X-linked thrombocytopenia (XLT).12-15 Despite this, there is significant clinical heterogeneity between different EVH1 missense mutations. In particular, mutations within exon 4 are associated with a more severe clinical phenotype.14,15 A recently published multicenter analysis of the clinical outcomes of XLT patients revealed significantly increased rates of infections, autoimmunity, and malignancy resulting in a median severe-event-free survival of only 10.2 years.16 Therefore, further categorization and mechanistic understanding of mutations associated with more severe clinical features is important.
The common mechanism of action for disease-causing EVH1 missense mutations appears to be enhanced protein degradation as a result of reduced WIP binding. Identification of the EVH1-binding surface for WIP as a long coiled surface clarified why so many different mutations could have the same effect. Based on this model, many EVH1 mutations have either been demonstrated in vitro (R86H, A134T, R138P, and Y107C)17,18 or predicted (28 of 35 EVH1 missense mutants)19,20 to have impaired WIP binding. Aside from this, the functional consequences of these mutations for WASp activity have been little explored.21 Furthermore, because of the reduced stability of clinical mutant WASp, the function of these proteins has not been assessed in cellular systems.
In the present study, we have investigated the functional activity of 4 disease-causing missense EVH1 WASp mutants. When expressed in cells, all 4 mutants demonstrated reduced binding to WIP and enhanced degradation. Using overexpression systems to restore cellular protein levels equivalent to wild-type WASp, we found that EVH1 missense mutants were unable to support normal WASp function, even though actin-polymerizing activity in vitro was preserved. Our findings suggest a critical role for the EVH1 domain in regulating WASp activity in addition to protecting it from degradation.
Methods
Mice
C57BL/6 (B6) wild-type (Charles River Laboratories) and WAS KO mice (supplied by T. Strom, Memphis, TN) were housed in specified pathogen-free conditions and used at 6-18 weeks of age. Experiments were performed under a Home Office–approved project license (held by S.B.). Immature murine BM derived dendritic cells (BMDCs) were generated by culturing in RPMI medium supplemented with 10% FCS, 100 units/mL of penicillin, and 100 μg/mL of streptomycin in the presence of 20 ng/mL of GM-CSF (PeproTech) for 7 days.
In vitro actin polymerization, pull-down, and immunoprecipitation assays
Confocal microscopy
Cells were prepared and stained using standard protocols (detailed in the supplemental Methods). Slides were imaged using a Zeiss LSM710 laser scanning spectral confocal microscope and a 63× Plan-Apochromat NA 1.4 WD 190-mm oil immersion lens. For podosome reconstitution, enumeration, volume, distribution, and cell-shape analysis, cells were imaged using sequential channel scanning: resolution of 4.55 pixels/μm (frame size, 1024 × 1024) and optimal aperture. Z stacks of 3 slices were acquired through the adhesive surface of the cell. Laser and acquisition settings remained the same for all samples. Random areas within each slide were selected for imaging, images were assigned a randomized code, and subsequent analysis was performed blinded.
Image analysis
Image analysis was performed with ImageJ Version 1.45d. Representative images shown are of individual Z slices with adjustments for brightness, contrast, and γ only. For podosome and cell-shape analysis, maximal projections of actin and vinculin images were used and cells were subjectively assigned a cell shape and podosome distribution category. Quantification of enhanced green fluorescent protein (EGFP)–WASp in individual cells was determined as follows: cell outlines were generated by thresholding maximal projections of actin signals and manually separating adjacent cells on the resultant masks. Cell outlines were applied to maximal projections of the EGFP signal and the integrated density signal (EGFPID) was measured to approximate EGFP-WASp concentration. EGFPID signals from all untransduced cells (WAS and B6) were natural log transformed to normality and mean (meanln(CON)) and SD (SDLn(CON)) was calculated. EGFPID signals for experimental cells (Exp-EGFPID) were converted to Z scores (multiple of SD from the mean) of the untransduced population using the following formula: zexpt = (Ln(Exp-EGFPID) − meanln(CON))/SDLn(CON). Logistical regression was used to analyze the relationship between EGFP signal (WASp concentration) and the probability of a cell forming podosomes. Z scores were banded to facilitate graphical representation. Analysis was blind to experimental group and quantitative analysis was performed on unmanipulated images.
Fluorescence recovery after photobleaching
BMDCs were cotransduced with EGFP-WASp (or EGFP alone) and LifeAct-mCherry using lentiviral infection. Imaging was performed using a Zeiss LSM710 microscope with a 63× lens. Double-positive cells with podosomes were identified and imaged using an 8× zoom (frame size, 184 × 184 pixels; resolution, 10.9 pixel/μm). Fluorophores were excited at a 488-nm wavelength (EGFP-WASp) or a 561-nm wavelength (LifeAct-mCherry). Laser intensities of 1.5% (488-nm laser; maximum, 25 mW) and 1% (561-nm laser; maximum, 15 mW) were used to obtain a strong signal while minimizing photobleaching. Images were acquired every 110 millisecond (pixel dwell, 1.4μs) to minimize general bleaching while sampling recovery. A region of interest of 60 × 40 pixels was bleached with 5 iterations of 100% laser power on 405- and 488-nm lasers. Imaging was performed for 3 seconds before bleaching (for normalization of initial fluorescence signal) and for 32 seconds after bleaching to measure fluorescence recovery. Recovery of individual fluorescence recovery after photobleaching (FRAP) curves to half of the initial intensity was used to determine the halftime. Average recovery curves were generated by plotting the mean standardized percentage of fluorescence recovery for each time point.
Migration assay
A total of 2.5 × 104 BMDCs were plated onto fibronectin-coated coverslips for at least 1 hour at 37°C. Inner chamber wells of Dunn chambers (Hawksley) were filled with culture medium and outer wells with culture medium containing 100 ng/mL of CCL3 (PeproTech). Chambers were imaged at 37°C using an inverted microscope (Zeiss Axiovert 135), 10×/0.25 NA lens (Zeiss). Images (phase and green fluorescence) were acquired every 10 minutes for 6 hours using a Hamamatsu digital camera (C4742-95) and analyzed using Volocity Version 5.3 software (Improvision).
Statistical analysis
Analysis was performed using SPSS Statistics Version 18 software (IBM) or Prism Version 5.01 software (GraphPad). Unless otherwise stated, differences between constructs within experiments were modeled using 2-way ANOVA, correcting for differences between experiments and constructs. Raw data were analyzed for normality and, where skewed, were transformed to normality. Estimated mean, SEM, and 95% confidence intervals were computed from the ANOVA model for each construct. Unless otherwise stated, differences between constructs were analyzed as fixed contrasts between wild-type and other conditions. For experiments in which output was most appropriately expressed as a ratio of a control value, results were expressed as symmetrical percentage differences (sympercentages) of the control value.24 Briefly, raw data were transformed by taking natural logs and multiplying by 100 and then analyzed by ANOVA (correcting for differences between experiments and constructs). Differences and their significance were then calculated between each construct and the control comparator. Differences (and SDs of differences) in the 100× ln scale corresponded directly to sympercentages between the construct and the control variable (without back transformation).24 Experiments with categorical outcomes were analyzed by logistical regression (with compensation for variation between experiments).
Results
Missense mutations in the EVH1 domain of WASp enhance actin-polymerizing activity in vitro
To investigate the function of disease-causing EVH1 domain mutations, we generated 4 lentiviral vectors containing human WASp missense mutations known to result in residual protein expression in vivo either with amino-terminal GST or EGFP tags (Figure 1A and supplemental Figure 1a-b). WASpR86H and WASpT45M are “hot spot” mutations associated with mild clinical disease, whereas WASpA134T and WASpR138P have a more variable and severe phenotype. In addition, EVH1 and VCA domain deletion WASp mutants (WASpdEVH1 and WASpdVCA), WASpH246D (which fails to bind Cdc4225 ), constitutively activeWASpS272P,26 and wild-type WASp (WASpWT) were generated (supplemental Figure 1a-b). All vectors successfully expressed at similar levels in Cos7 cells (supplemental Figure 1c).
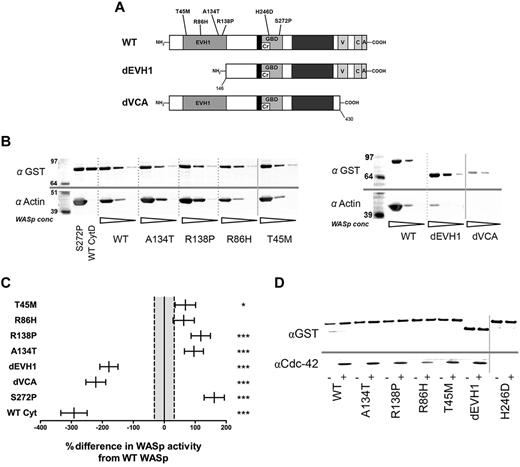
EVH1 missense mutations show increased actin polymerization in vitro but equivalent Cdc42 binding compared with WASpWT. (A) Structure of the WASp protein showing main functional domains and site of missense mutants generated for this study. (B) In vitro actin polymerization assay. Purified GST-WASp immobilized on Sepharose beads was incubated in U937 cell lysate supplemented with 5mM MgCl2 and 1mM ATP. Beads were resolved by SDS-PAGE and Western blotted for GST and actin. A representative experiment with 3 WASp concentrations for each construct is shown. All samples were performed simultaneously as part of a single experiment, but resolved on 2 blots (shown separately). (C) Summary of densitometry of GST-WASp and actin bands from 25 independent experiments. Differences between constructs were analyzed by ANCOVA of actin per unit GST-WASp (compensating for experiment and bead volume) and are expressed as the sympercentage difference in activity from WASpWT. Error bars represent SEM, and dotted lines show SEM of WT activity. Numbers of experimental samples analyzed per construct are as follows: WASpWT 49, WASpT45M 18, WASpR86H 19, WASpR138P 22, WASpA134T 22, WASpdEVH1 25, WASpdVCA 19, WASpS272P 19, WASpCytD 10. WT Cyt: WASpWT with 5nM cytochalasin D in lysate. (D) GST-WASp immobilized on Sepharose beads was incubated with U937 cell lysate supplemented with 125mM of the constitutively active Cdc42 12V mutant for 1 hour. Beads were resolved by SDS-PAGE and Western blotted for Cdc42 and GST. A representative blot from 3 experiments is shown. Nonadjacent lanes on Western blots are separated by solid gray lines. All lanes within each figure were on the same gel. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
EVH1 missense mutations show increased actin polymerization in vitro but equivalent Cdc42 binding compared with WASpWT. (A) Structure of the WASp protein showing main functional domains and site of missense mutants generated for this study. (B) In vitro actin polymerization assay. Purified GST-WASp immobilized on Sepharose beads was incubated in U937 cell lysate supplemented with 5mM MgCl2 and 1mM ATP. Beads were resolved by SDS-PAGE and Western blotted for GST and actin. A representative experiment with 3 WASp concentrations for each construct is shown. All samples were performed simultaneously as part of a single experiment, but resolved on 2 blots (shown separately). (C) Summary of densitometry of GST-WASp and actin bands from 25 independent experiments. Differences between constructs were analyzed by ANCOVA of actin per unit GST-WASp (compensating for experiment and bead volume) and are expressed as the sympercentage difference in activity from WASpWT. Error bars represent SEM, and dotted lines show SEM of WT activity. Numbers of experimental samples analyzed per construct are as follows: WASpWT 49, WASpT45M 18, WASpR86H 19, WASpR138P 22, WASpA134T 22, WASpdEVH1 25, WASpdVCA 19, WASpS272P 19, WASpCytD 10. WT Cyt: WASpWT with 5nM cytochalasin D in lysate. (D) GST-WASp immobilized on Sepharose beads was incubated with U937 cell lysate supplemented with 125mM of the constitutively active Cdc42 12V mutant for 1 hour. Beads were resolved by SDS-PAGE and Western blotted for Cdc42 and GST. A representative blot from 3 experiments is shown. Nonadjacent lanes on Western blots are separated by solid gray lines. All lanes within each figure were on the same gel. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
Although WAS-causing mutations are generally considered to result in loss of function, all EVH1 missense mutants (WASpEMM) were surprisingly found to induce more actin polymerization and bind more Arp2/3 than WASpWT (Figure 1B-C and supplemental Figure 1d) in an in vitro assay described previously.22,23 There was some variation, with WASpA138P and WASpA134T demonstrating approximately twice the actin-polymerization activity of WASpWT, whereas the increased activity of WASpR86H did not achieve statistical significance. These findings were not because of kinetic differences or saturation of the assay because similar differences were seen in response to increasing time increments and ATP concentration (supplemental Figure 1e-f). As expected, WASpS272P (Figure 1C) supported the highest levels of activity.22,26 In contrast, WASpdEVH1 demonstrated significantly reduced Arp2/3 binding and actin polymerization, but not to the same degree as WASpdVCA, which bound no detectable Arp2/3 and abolished all WASp-mediated actin-nucleating activity to a similar level as cytochalasin D (Figure 1B-C and supplemental Figure 1d,f-g). Because Cdc42 is a key activator of WASp,3,4 we investigated whether the enhanced activity of WASpEMM could result from differences in levels of Cdc42 binding. We found that all mutants, including WASpdEVH1, bound similar amounts of Cdc42 as WASpWT (Figure 1D) with the exception of WASpH246D, which, as expected, failed to bind Cdc42.25
EVH1 missense mutants are expressed at lower levels in cells and demonstrate increased rates of degradation and reduced WIP binding
To investigate intracellular activity, EGFP-WASp fusion proteins were expressed in U937 cells using lentiviral transduction at the same multiplicity of infection. All 4 EVH1 missense mutants were expressed at lower levels of protein than WASpWT (supplemental Figure 2a-c) and 35S protein–labeled pulse-chase experiments implied an increased rate of degradation (Figure 2A-B and supplemental Figure 2d). Interestingly, WASpdEVH1 showed considerable resistance to degradation, with a greater concentration of 35S-labeled protein at 24 hours than at the end of the pulse period (Figure 2A-B and supplemental Figure 2d).
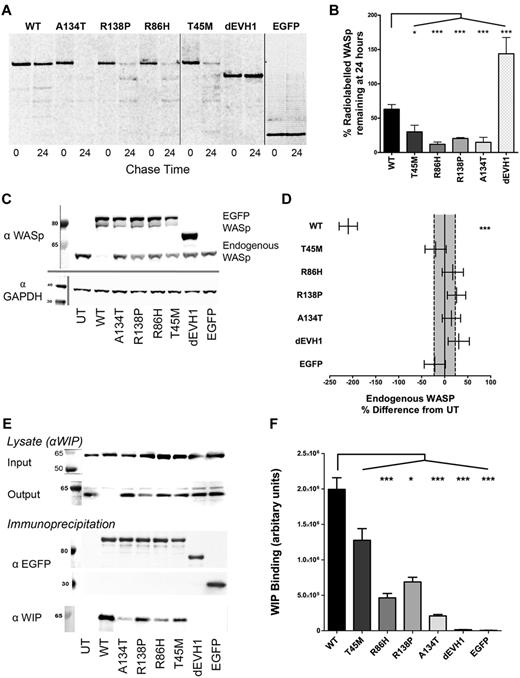
WASpEMM is degraded more rapidly and binds less WIP than WASpWT, but after forced expression with lentiviral transduction, can stably express at similar protein levels to endogenous WASp. (A-B) U937 cells were transduced with EGFP-WASp by lentiviral infection at an multiplicity of infection of 5 to generate stable cell lines. Cells were labeled with 35S cysteine/methionine (pulse), followed by 24 hours incubation in full RPMI with additional 2mM methionine and 2mM cysteine (chase). Samples were harvested before and after the chase period by lysis and anti-EGFP immunoprecipitation. Immunoprecipitates were resolved by SDS-PAGE and radio emission of dried gels was detected by phosphor screens and imaged using a Molecular Imager FX (Bio-Rad). Radio emission signals were quantified and the percent change in signal between 0 and 24 hours of chase was calculated. Differences between constructs in percent degradation from 3 experiments were analyzed by ANOVA. Nonadjacent lanes on Western blots are separated by solid gray lines. All lanes within each figure were on the same gel. (C) U937 cell lines stably transduced with EGFP-WASp constructs were sorted for similar EGFP expression. After 5 passages, cells were harvested and resolved lysates were immunoblotted for WASp and GAPDH (loading control). UT indicates untransduced. (D) Endogenous WASp bands were quantified by densitometry and differences between constructs from 4 experiments were analyzed by ANOVA. Absolute values were expressed as sympercentage differences from endogenous WASp levels in untransduced cells. Error bars represent SEM and dotted lines represent SEM of endogenous WASp concentration in untransduced cells. (E) U937 cells with standardized EGFP-WASp expression were harvested and lysates underwent anti-EGFP immunoprecipitation. Pre- and postimmunoprecipitation lysates were resolved by SDS-PAGE and immunoblotted for WIP. Immunoprecipitants were resolved and immunoblotted for EGFP and WIP. (F) EGFP and WIP bands from 6 experiments were quantified by densitometry. Differences between constructs were analyzed by ANOVA. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
WASpEMM is degraded more rapidly and binds less WIP than WASpWT, but after forced expression with lentiviral transduction, can stably express at similar protein levels to endogenous WASp. (A-B) U937 cells were transduced with EGFP-WASp by lentiviral infection at an multiplicity of infection of 5 to generate stable cell lines. Cells were labeled with 35S cysteine/methionine (pulse), followed by 24 hours incubation in full RPMI with additional 2mM methionine and 2mM cysteine (chase). Samples were harvested before and after the chase period by lysis and anti-EGFP immunoprecipitation. Immunoprecipitates were resolved by SDS-PAGE and radio emission of dried gels was detected by phosphor screens and imaged using a Molecular Imager FX (Bio-Rad). Radio emission signals were quantified and the percent change in signal between 0 and 24 hours of chase was calculated. Differences between constructs in percent degradation from 3 experiments were analyzed by ANOVA. Nonadjacent lanes on Western blots are separated by solid gray lines. All lanes within each figure were on the same gel. (C) U937 cell lines stably transduced with EGFP-WASp constructs were sorted for similar EGFP expression. After 5 passages, cells were harvested and resolved lysates were immunoblotted for WASp and GAPDH (loading control). UT indicates untransduced. (D) Endogenous WASp bands were quantified by densitometry and differences between constructs from 4 experiments were analyzed by ANOVA. Absolute values were expressed as sympercentage differences from endogenous WASp levels in untransduced cells. Error bars represent SEM and dotted lines represent SEM of endogenous WASp concentration in untransduced cells. (E) U937 cells with standardized EGFP-WASp expression were harvested and lysates underwent anti-EGFP immunoprecipitation. Pre- and postimmunoprecipitation lysates were resolved by SDS-PAGE and immunoblotted for WIP. Immunoprecipitants were resolved and immunoblotted for EGFP and WIP. (F) EGFP and WIP bands from 6 experiments were quantified by densitometry. Differences between constructs were analyzed by ANOVA. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
To control for protein expression levels, transduced cells were sorted to achieve comparable levels of WASp expression between mutants and WASpWT (supplemental Figure 2e-f). All proteins were stably expressed at equivalent levels to endogenous WASp, with no significant differences in EGFP-WASp levels between constructs (Figure 2C and supplemental Figure 2g) or differences in total cellular WIP levels (supplemental Figure 2h). EGFP-WASp mRNA expression in WASpEMM-sorted cell lines (as determined by real-time quantitative PCR of reverse transcribed total RNA) was greater or equivalent than that of WASpWT (1.0-5.2 × WASpWT expression), whereas WASpdEVH1 showed reduced relative expression (0.4 × WASpWT expression). This is consistent with previous studies showing that, when overexpressed in U937 cells, WASpEMM has increased susceptibility to degradation rather than impaired translation.12-15
Overexpression of WASpWT resulted in reduced expression of endogenous WASp (Figure 2C-D), suggesting that the total WASp concentration is controlled by the availability of free intracellular WIP, which prevents degradation.27-29 In contrast, endogenous WASp expression was relatively preserved in cells transduced with WASpEMM, most likely reflecting their reduced ability to form WIP-WASp complexes relative to endogenous WASp and subsequent preferential degradation. Consistent with this, WASp mutants immunoprecipitated significantly less WIP from U937 cells lysates than WASpWT (Figure 2E-F), demonstrating a reduced affinity for WIP. The degree of residual WIP affinity was variable, with WASpT45M demonstrating the highest and WASpA134T the lowest levels of binding. As predicted, WASpdEVH1 showed no detectable WIP binding (Figure 2E-F).
Even when overexpressed, EVH1 missense mutant WASp is unable to reconstitute podosomes as efficiently as WASpWT
WASp-mediated actin polymerization is required for several important immune cell functions, including podosome formation and migration in macrophages and dendritic cells (DCs).30,31 To test the function of WASpEMM in cells, we examined their ability to reconstitute podosome formation in WASp-null BMDCs (WASBMDCs). After adjusting for differences in Ab binding between human and murine WASp (supplemental Figure 3a) and transduction efficiency, we established an experimental model in which transduced WASBMDCs expressed WASpEMM at higher or similar levels than endogenous WASp (in B6 BMDCs) at all multiplicities of infection tested with equivalent transduction efficiency and cell viability (Figure 3A and supplemental Figure 3b).
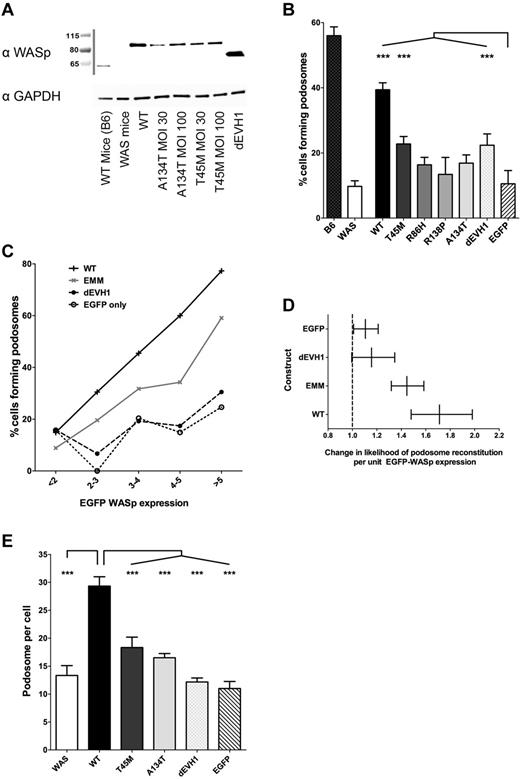
WASpEMM is unable to reconstitute podosomes as effectively as WASpWT, even at equivalent levels of WASp expression.WASBMDCs were transduced on day 2 of culture with EGFP-WASp by lentiviral infection (multiplicity of infection, 30 and 100). On day 7 of culture, cells were either lysed for Western blot analysis or harvested and plated on fibronectin-coated glass coverslips or chamber slides for 4 hours. Cells were then fixed with 4% paraformaldehyde, permeabilized with Triton X-100, stained with rabbit anti-EGFP and anti–rabbit Alexa Fluor 488, Alex Fluor 568 phalloidin, mouse anti-vinculin, and goat anti–mouse Cy5, and mounted in ProFade Gold. Slides were imaged using a Zeiss LSM710 laser scanning spectral confocal microscope and a 63× Plan-Apochromat NA 1.4 WD 190-mm oil immersion lens, and images were acquired with Zen 2009 software (Zeiss). All image analysis was performed using ImageJ Version 1.45d software. (A) Western blot of transduced WASBMDCs immunoblotted for WASp and GAPDH (loading control). (B) Confocal images of transduced cells were blindly analyzed to determine the proportion of cells that produce podosomes. The percentage of transduced cells producing podosomes for each construct was calculated from 3 independent experiments, with a total of 194-391 transduced cells analyzed for each condition. Differences between constructs were analyzed by ANOVA, with comparisons made with EGFP-only–transduced cells. (C) Likelihood of transduced cells forming podosomes increases with increasing expression of WASp protein. WASp expression within individual cells was estimated by calculating the integrated density of the EGFP signal within the cells outline. This value was converted to the number of SEMs of the EGFP signal in untransduced cells (Z score of negative population). Cells were grouped for EGFP intensity as follows: Z score < 2 (EGFP negative cells), 2-3, 3-4, 4-5, > 5, and the percentage of podosome-positive cells within each banding was calculated for each WASp construct. Data from all 4 EVH1 missense mutant WASp-transduced cells were combined and analyzed together (labeled EMM) to avoid statistical errors associated with small numbers of cells expressing high levels of EVH1 missense mutant WASp. The numbers of cells analyzed for each construct at each expression band is shown in supplemental Table 1. Results presented represent combined data from 2 independent experiments (total of 3094 cells analyzed). (D) Logistical regression to determine the size of the impact of EGFP-WASp expression on the likelihood of an individual cell forming podosomes for each individual construct (compensating for difference between experiments). EGFP-WASp expression was expressed as Z scores of untransduced EGFP signal and error bars represent 95% confidence intervals for the likelihood ratio. (E) Among cells that formed podosomes, the number of podosomes produced per cell was counted and the median was calculated for each construct. The average median podosome number per cell from 3 independent experiments is represented, with differences between constructs analyzed using ANOVA. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
WASpEMM is unable to reconstitute podosomes as effectively as WASpWT, even at equivalent levels of WASp expression.WASBMDCs were transduced on day 2 of culture with EGFP-WASp by lentiviral infection (multiplicity of infection, 30 and 100). On day 7 of culture, cells were either lysed for Western blot analysis or harvested and plated on fibronectin-coated glass coverslips or chamber slides for 4 hours. Cells were then fixed with 4% paraformaldehyde, permeabilized with Triton X-100, stained with rabbit anti-EGFP and anti–rabbit Alexa Fluor 488, Alex Fluor 568 phalloidin, mouse anti-vinculin, and goat anti–mouse Cy5, and mounted in ProFade Gold. Slides were imaged using a Zeiss LSM710 laser scanning spectral confocal microscope and a 63× Plan-Apochromat NA 1.4 WD 190-mm oil immersion lens, and images were acquired with Zen 2009 software (Zeiss). All image analysis was performed using ImageJ Version 1.45d software. (A) Western blot of transduced WASBMDCs immunoblotted for WASp and GAPDH (loading control). (B) Confocal images of transduced cells were blindly analyzed to determine the proportion of cells that produce podosomes. The percentage of transduced cells producing podosomes for each construct was calculated from 3 independent experiments, with a total of 194-391 transduced cells analyzed for each condition. Differences between constructs were analyzed by ANOVA, with comparisons made with EGFP-only–transduced cells. (C) Likelihood of transduced cells forming podosomes increases with increasing expression of WASp protein. WASp expression within individual cells was estimated by calculating the integrated density of the EGFP signal within the cells outline. This value was converted to the number of SEMs of the EGFP signal in untransduced cells (Z score of negative population). Cells were grouped for EGFP intensity as follows: Z score < 2 (EGFP negative cells), 2-3, 3-4, 4-5, > 5, and the percentage of podosome-positive cells within each banding was calculated for each WASp construct. Data from all 4 EVH1 missense mutant WASp-transduced cells were combined and analyzed together (labeled EMM) to avoid statistical errors associated with small numbers of cells expressing high levels of EVH1 missense mutant WASp. The numbers of cells analyzed for each construct at each expression band is shown in supplemental Table 1. Results presented represent combined data from 2 independent experiments (total of 3094 cells analyzed). (D) Logistical regression to determine the size of the impact of EGFP-WASp expression on the likelihood of an individual cell forming podosomes for each individual construct (compensating for difference between experiments). EGFP-WASp expression was expressed as Z scores of untransduced EGFP signal and error bars represent 95% confidence intervals for the likelihood ratio. (E) Among cells that formed podosomes, the number of podosomes produced per cell was counted and the median was calculated for each construct. The average median podosome number per cell from 3 independent experiments is represented, with differences between constructs analyzed using ANOVA. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
When plated on fibronectin, 56.0% of untransduced B6 control BMDCs assembled podosomes compared with 9.8% of WASBMDCs (Figure 3B). Transduction of WASBMDCs with WASpWT reconstituted podosome formation in 39.4% of cells, whereas EGFP alone resulted in indistinguishable podosome formation from WASp-null cells. WASpEMM was significantly less effective than WASpWT at supporting podosome formation (13.4%-22.7% of cells) and, in fact, only WASpT45M showed significantly increased reconstitution over EGFP alone (P < .001). Surprisingly, WASpdEVH1 was also able to reconstitute typical podosome structures in 22.4% of cells (significantly greater than EGFP alone, P < .005, Figure 3B).
To address the possibility that reduced podosome assembly by WASpEMM resulted from lower levels of protein expression compared with WASpWT, we analyzed individual cells matched for EGFP-WASp expression. Quantification of EGFP-WASp by measurement of fluorescent signal on confocal micrographs was validated by comparison with median fluorescent intensity on flow cytometry and densitometry of western blots over a range of expression levels (supplemental Figure 4a-f). Because it was difficult to achieve very high levels of expression of WASpEMM in WASBMDCs, the results of the missense mutants were combined (EMM) to achieve sufficient power for interpretation (supplemental Table 1). Both WASpWT and WASpEMM showed a positive correlation between WASp expression level and podosome reconstitution, but WASpWT supported higher levels of podosome reconstitution at every level of protein expression (Figure 3C) and yielded a greater increase in likelihood of podosome reconstitution per unit WASp expression than WASpEMM (Figure 3D). WASpdEVH1 showed no significant correlation between protein expression and likelihood of podosome formation (Figure 3D). Similarly, using logistical regression, we calculated the odds ratios for individual cells forming podosomes for each construct while compensating for EGFP-WASp expression level (Table 1). WASpWT-transduced cells were 3.4 times more likely to form podosomes than cells transformed with EGFP alone, whereas the ratios for WASpEMM-transduced cells were between 0.9 and 2.1 times. This demonstrates that the impaired ability of WASpEMM to reconstitute podosomes does not depend solely on the absolute level of WASp expression, but rather on loss of protein functionality. Consistent with this interpretation, WASpWT also reconstituted significantly more podosomes per cell than EVH1 missense mutant WASp (Figure 3E).
EVH1 mutant WASp localizes to podosomes and supports characteristic podosome morphology and dynamics
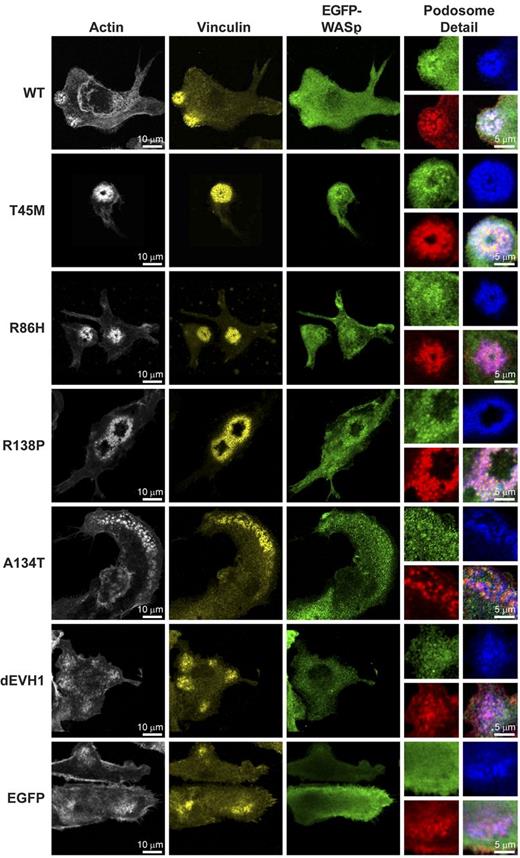
It has been reported previously that, in the absence of WIP, WASp activity is deregulated, resulting in localization of WASp to anomalous f-actin plaques.29 In contrast, we observed that when podosomes were formed by overexpression of any of the WASpEMM proteins, characteristic podosome morphology was seen, as demonstrated by discrete actin cores and surrounding vinculin rings, (Figure 4). Surprisingly, WASpdEVH1 was capable of forming podosomes with normal architecture, although it also resulted in assembly of abnormal f-actin plaques (supplemental Figure 5a) reminiscent of those formed by overexpression of WASp in WIP-deficient cells.29 Because localization of EGFP-WASp was better visualized in unfixed cells, WASBMDCs were cotransduced with EGFP-WASp and LifeAct-mCherry (a fluorescent f-actin–binding probe). WASpT45M was seen to colocalize strongly with f-actin in podosomes. Despite minimal WIP binding, WASpA134T also demonstrated significant podosome localization (supplemental Video). Surprisingly, even WASpdEVH1, which has no WIP binding, was clearly visible in podosomes (supplemental Video) and plaque structures (supplemental Figure 5a), indicating that WIP alone is not essential for WASp recruitment to actin structures at the cell membrane.
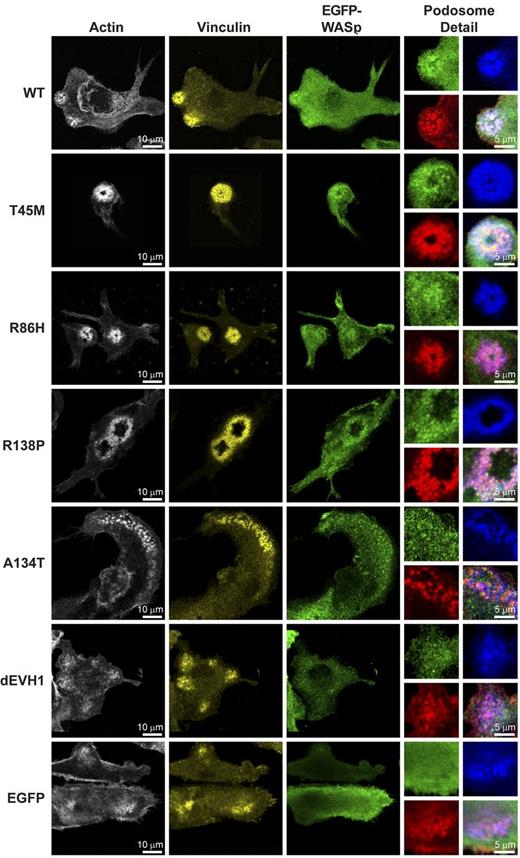
WASpEMM reconstitutes and localizes to structurally characteristic podosomes.WASBMDCs were plated onto fibronectin-coated glass coverslips on day 7 of culture. After 4 hours of adherence, cells were fixed and immunostained for EGFP (rabbit anti-EGFP and anti–rabbit Alexa Fluor 488), f-actin (Alexa Fluor 568 phalloidin), and vinculin (mouse anti-vinculin and goat anti–mouse Cy5). Images were taken using a Zeiss LSM710 laser scanning spectral confocal microscope and a 63× Plan-Apochromat NA 1.4 WD 190-mm oil immersion lens and processed using ImageJ Version 1.45d software. Images represent single Z stack slice through the adhesive surface of the cell, and adjustments were made to brightness, contrast, and γ only. For whole-cell images, actin is shown in white, vinculin in yellow, and EGFP-WASp in green. To aid the visualization of colocalization, in the podosomes detail panels, the colors are changed to actin in red, vinculin in blue, and EGFP-WASp in green. The bottom right panel for each construct represents a merged image of all 3 channels.
WASpEMM reconstitutes and localizes to structurally characteristic podosomes.WASBMDCs were plated onto fibronectin-coated glass coverslips on day 7 of culture. After 4 hours of adherence, cells were fixed and immunostained for EGFP (rabbit anti-EGFP and anti–rabbit Alexa Fluor 488), f-actin (Alexa Fluor 568 phalloidin), and vinculin (mouse anti-vinculin and goat anti–mouse Cy5). Images were taken using a Zeiss LSM710 laser scanning spectral confocal microscope and a 63× Plan-Apochromat NA 1.4 WD 190-mm oil immersion lens and processed using ImageJ Version 1.45d software. Images represent single Z stack slice through the adhesive surface of the cell, and adjustments were made to brightness, contrast, and γ only. For whole-cell images, actin is shown in white, vinculin in yellow, and EGFP-WASp in green. To aid the visualization of colocalization, in the podosomes detail panels, the colors are changed to actin in red, vinculin in blue, and EGFP-WASp in green. The bottom right panel for each construct represents a merged image of all 3 channels.
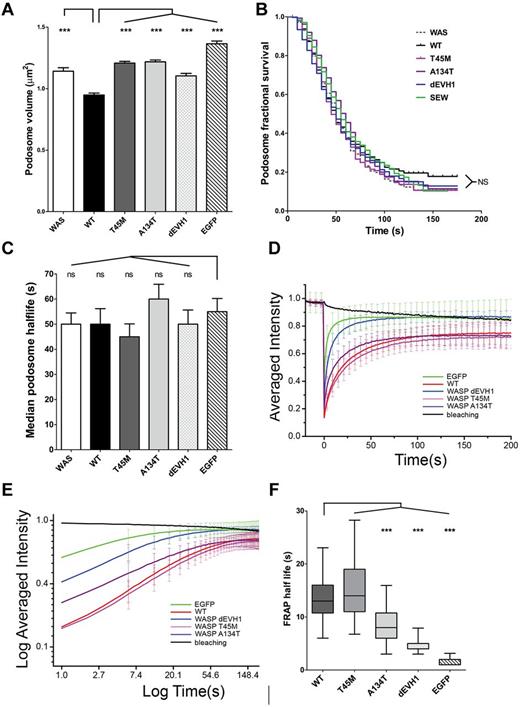
Further analysis revealed that podosomes formed in WASpEMM- and WASpdEVH1- transduced cells had a significantly larger volume than podosomes in WASpWT-transduced cells (1.21 vs 0.95 μm2, P < .001), but were significantly smaller than podosomes from cells transduced with EGFP alone (1.37 vs 1.10-1.21 μm2, P < .001; Figure 5A). These results suggest that WIP-WASp interaction plays a role in the organization of podosome structure. Detailed analysis of podosome dynamics by live imaging of WASBMDCs cotransduced with EGFP-WASp and LifeAct-mCherry revealed no significant differences in podosome survival curves, half-life, adhesion turnover index,32 or rate of podosome formation (Figure 5B-C and supplemental Figure 5b-c). Similarly, no differences were seen in the distribution of podosomes within individual cells (supplemental Figure 5d).
Podosome volume and WASp retention within podosomes is predicted by WIP binding affinity, but podosome dynamics are unaffected. (A) Podosome volume was analyzed in transduced WASBMDCs plated on fibronectin-coated glass coverslips, fixed, and immunostained for EGFP, f-actin, and vinculin. Podosomes were defined using actin and vinculin signals on maximal intensity projections and cross-sectional areas were measured in ImageJ Version 1.45d software. Differences between constructs were analyzed by ANOVA with a Bonferroni correction. (B-F) WASBMDCs were cotransduced with EGFP-WASp (or EGFP alone) and LifeAct-mCherry using lentiviral infection and plated onto fibronectin-coated glass-bottomed Petri dishes for 4 hours before imaging. All imaging was performed using a Zeiss LSM710 microscope with a 63× lens. (B-C) Cells with EGFP-WASp localized to podosomes were identified and imaged with EGFP and LifeAct signals for 3 minutes (single Z stack, 12 frames/min). Podosome life was defined as the time an individual podosome's actin intensity remained greater than 75% of its maximum intensity. Kaplan-Meier fractional survival curves were plotted (B) and used to calculate median podosome half-life for each construct (C). Differences between survival curves were analyzed using the log-rank (Mantel-Cox) test using both WT and EGFP as standards and no significant differences were seen. Median podosome half-life was calculated for each individual cell and differences between constructs were analyzed by ANOVA. (D-F) Podosome patches were identified using the LifeAct-mCherry signal and FRAP was performed on the EGFP signal. The fluorescence recovery signal was standardized to natural bleaching to give a percent fluorescence recovery for each time point. Data presented are combined from 18-30 FRAP experiments in each of 2 independent experiments. (D-E) WASpWT and WASpT45M show significantly longer retention in podosomes than EGFP alone, WASpdEVH1, and WASpA134T. Average recovery curves were generated by plotting mean (and SD) of standardized percent fluorescence recovery for each time point. (F) The half-life of individual FRAP curves was determined by the time taken when the FRAP recovery curve reached half of the initial intensity. Median, interquartile range (box), and range (whiskers) for each construct is presented and differences were analyzed by ANOVA with a Bonferroni correction. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
Podosome volume and WASp retention within podosomes is predicted by WIP binding affinity, but podosome dynamics are unaffected. (A) Podosome volume was analyzed in transduced WASBMDCs plated on fibronectin-coated glass coverslips, fixed, and immunostained for EGFP, f-actin, and vinculin. Podosomes were defined using actin and vinculin signals on maximal intensity projections and cross-sectional areas were measured in ImageJ Version 1.45d software. Differences between constructs were analyzed by ANOVA with a Bonferroni correction. (B-F) WASBMDCs were cotransduced with EGFP-WASp (or EGFP alone) and LifeAct-mCherry using lentiviral infection and plated onto fibronectin-coated glass-bottomed Petri dishes for 4 hours before imaging. All imaging was performed using a Zeiss LSM710 microscope with a 63× lens. (B-C) Cells with EGFP-WASp localized to podosomes were identified and imaged with EGFP and LifeAct signals for 3 minutes (single Z stack, 12 frames/min). Podosome life was defined as the time an individual podosome's actin intensity remained greater than 75% of its maximum intensity. Kaplan-Meier fractional survival curves were plotted (B) and used to calculate median podosome half-life for each construct (C). Differences between survival curves were analyzed using the log-rank (Mantel-Cox) test using both WT and EGFP as standards and no significant differences were seen. Median podosome half-life was calculated for each individual cell and differences between constructs were analyzed by ANOVA. (D-F) Podosome patches were identified using the LifeAct-mCherry signal and FRAP was performed on the EGFP signal. The fluorescence recovery signal was standardized to natural bleaching to give a percent fluorescence recovery for each time point. Data presented are combined from 18-30 FRAP experiments in each of 2 independent experiments. (D-E) WASpWT and WASpT45M show significantly longer retention in podosomes than EGFP alone, WASpdEVH1, and WASpA134T. Average recovery curves were generated by plotting mean (and SD) of standardized percent fluorescence recovery for each time point. (F) The half-life of individual FRAP curves was determined by the time taken when the FRAP recovery curve reached half of the initial intensity. Median, interquartile range (box), and range (whiskers) for each construct is presented and differences were analyzed by ANOVA with a Bonferroni correction. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
WASp retention in podosomes is disrupted by EVH1 mutations
Because WASpEMM was observed to localize to podosomes, we investigated the dynamics of WASpEMM localization within sites of actin polymerization using FRAP of EGFP-WASp proteins in transduced WASBMDCs. In these experiments, if EGFP-WASp binds strongly to podosomes, we expect fluorescence to recover slowly, with some proteins staying bound throughout the recovery period monitored (the unexchanged fraction). Conversely, if binding is weak, proteins exchange rapidly and fluorescence recovers rapidly. As expected, EGFP alone showed very rapid return to podosome patches after bleaching with no unexchanged protein (Figure 5D-F), demonstrating the rapid and purely diffusive dynamics of the EGFP tag. In contrast, WASpWT showed a much slower fluorescence recovery with an unexchanged fraction of 10% at equilibrium (Figure 5D-E), indicating a significantly longer half-life in the podosome than EGFP (median T1/2 = 13 seconds for WASpWT compared with median T1/2 = 2 seconds for EGFP, P < .001, Figure 5F). Consistent with our interpretation, WASpT45M, which retains the most WIP binding of all of the WASpEMM tested (Figure 2F), showed similar fluorescence recovery and half-life in the podosome to WASpWT, implying a comparable podosome-binding affinity. In contrast, WASpA134T, which preserves minimal WIP binding (Figure 2F), had a significantly faster fluorescence recovery rate, as reflected by a shortened half-life in the podosome (median T1/2 = 8.8 seconds, P < .001, compared with WASpWT), but recovery still remained significantly slower than EGFP alone. Unexchanged fractions for WASpWT and both WASpEMM were indistinguishable. These results demonstrate that disease-causing missense mutations in the EVH1 domain alter the dynamics of WASp-binding affinity and WASp turnover at podosome sites. This is not due simply to accelerated protein degradation, because WASpdEVH1, which is inherently resistant to degradation, had an even shorter half-life in podosomes than WASpA134T. A role for WIP in retaining WASp in podosomes is suggested instead.
Regulation of WASp activity in cells is disrupted by EVH1 mutations
Regulation of WASp activity in cells depends on interaction with Cdc42 at membrane sites and subsequent tyrosine phosphorylation at residue Y291.23,33,34 We hypothesized that reduced retention at sites of membrane-associated actin polymerization such as podosomes could affect the in vivo function of WASpEMM. Consistent with our in vitro bead assay data, which suggested intrinsic dysregulation, WASBMDCs expressing WASpEMM were generally found to have increased intracellular f-actin content after LPS stimulation than cells expressing WASpWT (Figure 6A). The effect was not observed for WASpT45M, which preserves most WIP binding. The significantly elevated f-actin levels observed in cells expressing WASpdEVH1 is likely to reflect resistance of this mutant to degradation, resulting in very high intracellular concentration of mutant protein (Figure 2A-B) that may sequester negative regulators of WASp activity. Furthermore, in cells, all WASpEMM showed less tyrosine phosphorylation than WASpWT (Figure 6B-C), suggesting reduced activation. In contrast, constitutive serine phosphorylation at S483/S48435 was similar between all WASp constructs (Figure 6B,D). These data further suggest that the EVH1 domain participates in regulation of intracellular WASp activity.
WASpEMM induces dysregulation of actin polymerization in vivo, which results in an increased susceptibility to form multiple lamellipodia and subsequent migration defect. (A) Transduced WASBMDCs were matured with lipopolysaccharide and harvested on day 9 of culture. F-actin within cells was quantified by flow cytometry after fixation and staining with Alexa Fluor 633 phalloidin. The sympercentage difference in median f-actin signal between transduced and simultaneously stained untransduced WASBMDCs is shown, with error bars representing the SEM. Differences between wild-type transduced and other constructs were analyzed by ANOVA. Data represent 8 independent experiments. (B) WASpEMM is less tyrosine phosphorylated than WASpWT in vivo. U937 cells with standardized EGFP-WASp expression were harvested and lysates underwent anti-EGFP immunoprecipitation. Immunoprecipitant were resolved and immunoblotted for EGFP, phosphotyrosine (4G10; pTyr), and WASp pSER433/434 (pSer). Representative immunoblots shown. (C-D) EGFP, pTyr, and pSer bands from 8 experiments were quantified by densitometry. The percent tyrosine phosphorylation was calculated by comparison with fully tyrosine phosphorylated standards, and the means and 95% confidence interval are graphically represented. Differences between wild-type and other constructs were analyzed by ANOVA (C). WASp serine phosphorylation was compared with WASpWT and mean differences (from 4 experiments) are expressed as sympercentage differences from WASpWT (with 95% confidence intervals). The 95% confidence interval for WASpWT is shown by dotted vertical lines. Differences from WASpWT were analyzed by ANOVA (D). (E) Transduction of WASBMDCs with WASpEMM results in a greater proportion of cells forming > 2 lamellipodia compared with WASpWT-transduced cells. Cell shape of fixed transduced WASBMDCs plated onto fibronectin-coated coverslips in 3 independent experiments was analyzed. EGFP-positive, spread cells with lamellipodia were scored as having > or ≤ 2 lamellipodia. (F) Relative odds (and 95% confidence interval) of an individual spread cell forming > 2 lamellipodia compared with wild-type transduced cells is shown. Odds ratios and significance of differences WT and other constructs were calculated using logistical regression. (G-I) WASBMDCs transduced with WASpEMM show similar velocity but reduced displacement and persistence during migration compared with WASpWT-transduced cells. Migration of transduced DCs plated onto fibronectin-coated coverslips toward a CCL3 gradient was assayed in Dunn chambers. Cells were imaged in phase and green fluorescence for 6 hours using an Zeiss Axiovert 135 (10 × /0.25 NA lens), and cell tracks were analyzed using Volocity Version 5.3.2 software. EGFP-negative cells on the same coverslip as transduced cells were analyzed as WAS controls. Average velocity (G), total cell displacement (H), and persistence of migration (meandering index; I) collated from 5 independent experiments are presented with 95% confidence intervals. Differences between wild-type and other construct transduced cells were analyzed by ANOVA with correction for variation between experiments. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
WASpEMM induces dysregulation of actin polymerization in vivo, which results in an increased susceptibility to form multiple lamellipodia and subsequent migration defect. (A) Transduced WASBMDCs were matured with lipopolysaccharide and harvested on day 9 of culture. F-actin within cells was quantified by flow cytometry after fixation and staining with Alexa Fluor 633 phalloidin. The sympercentage difference in median f-actin signal between transduced and simultaneously stained untransduced WASBMDCs is shown, with error bars representing the SEM. Differences between wild-type transduced and other constructs were analyzed by ANOVA. Data represent 8 independent experiments. (B) WASpEMM is less tyrosine phosphorylated than WASpWT in vivo. U937 cells with standardized EGFP-WASp expression were harvested and lysates underwent anti-EGFP immunoprecipitation. Immunoprecipitant were resolved and immunoblotted for EGFP, phosphotyrosine (4G10; pTyr), and WASp pSER433/434 (pSer). Representative immunoblots shown. (C-D) EGFP, pTyr, and pSer bands from 8 experiments were quantified by densitometry. The percent tyrosine phosphorylation was calculated by comparison with fully tyrosine phosphorylated standards, and the means and 95% confidence interval are graphically represented. Differences between wild-type and other constructs were analyzed by ANOVA (C). WASp serine phosphorylation was compared with WASpWT and mean differences (from 4 experiments) are expressed as sympercentage differences from WASpWT (with 95% confidence intervals). The 95% confidence interval for WASpWT is shown by dotted vertical lines. Differences from WASpWT were analyzed by ANOVA (D). (E) Transduction of WASBMDCs with WASpEMM results in a greater proportion of cells forming > 2 lamellipodia compared with WASpWT-transduced cells. Cell shape of fixed transduced WASBMDCs plated onto fibronectin-coated coverslips in 3 independent experiments was analyzed. EGFP-positive, spread cells with lamellipodia were scored as having > or ≤ 2 lamellipodia. (F) Relative odds (and 95% confidence interval) of an individual spread cell forming > 2 lamellipodia compared with wild-type transduced cells is shown. Odds ratios and significance of differences WT and other constructs were calculated using logistical regression. (G-I) WASBMDCs transduced with WASpEMM show similar velocity but reduced displacement and persistence during migration compared with WASpWT-transduced cells. Migration of transduced DCs plated onto fibronectin-coated coverslips toward a CCL3 gradient was assayed in Dunn chambers. Cells were imaged in phase and green fluorescence for 6 hours using an Zeiss Axiovert 135 (10 × /0.25 NA lens), and cell tracks were analyzed using Volocity Version 5.3.2 software. EGFP-negative cells on the same coverslip as transduced cells were analyzed as WAS controls. Average velocity (G), total cell displacement (H), and persistence of migration (meandering index; I) collated from 5 independent experiments are presented with 95% confidence intervals. Differences between wild-type and other construct transduced cells were analyzed by ANOVA with correction for variation between experiments. ***P < .005, **P < .01, *P < .05, and ns indicates P > .05.
EVH1 mutations impair DC migration
To determine whether the effects of EVH1 mutations on WASp activation and podosome assembly are sufficient to impair DC function, we analyzed migration in vitro. Directed cell migration depends on regulation of lamellipodia formation during cell polarization to select a dominant leading edge.36 After adhesion to fibronectin, no differences were seen in the proportion of spread or polarized cells or cell morphology between WASpWT and WASpEMM- or WASpdEVH1-transduced WASBMDCs (supplemental Figure 5e-g). However, polarized cells expressing WASpWT most frequently selected 1-2 lamellipodia, which is comparable with untransduced B6 BMDCs. In contrast, a higher percentage of polarized WASBMDCs expressing WASpEMM formed multiple (> 2) lamellipodia, as seen in untransduced or EGFP-transduced WASBMDCs (Figure 6E-F). In Dunn chamber migration assays, restoration of WASpWT in WASBMDCs did not influence cell velocity, but significantly increased cell displacement (distance traveled from the point of origin, Figure 6G-H and supplemental Table 2) as a result of improved directional persistence (movement in a straight line, Figure 6I). Although migration velocity was similar in WASBMDCs expressing WASpWT or any WASpEMM, displacement and persistence of migration were reduced in cells expressing mutant WASp (Figure 6G-I), with the greatest effect on WASpA134T and WASpdEVH1, which retain less WIP binding than WASpT45M. These data demonstrate that the EVH1 domain is required to regulate DC migration and that EVH1 missense mutations, even when overexpressed, do not fully support normal DC migration.
Discussion
A significant number of children affected by mutations in the WAS gene harbor missense mutations in the EVH1 domain that preserve some WASp expression and confer a milder form of the disease. It has been shown for WASpR86H and WASPA134T EVH1 domain mutations17,18 and predicted for others19,20 that reduced levels of protein are the result of impaired WIP binding leading to enhanced WASp degradation through proteasome- and calpain-mediated pathways.28,29,37 This has led to the prospect of novel treatments for patients with EVH1 domain mutations aimed at inhibiting mutant WASp degradation to restore physiologic levels of mutant WASp. In a recent study, expression of a 41–amino acid portion of the WASp-binding region of WIP restored WASp expression and T-cell spreading in cells isolated from patients with selected EVH1 mutations.38 However, the success of such strategies will depend on a relatively normal intrinsic function of WASp harboring such missense mutations, and, to our knowledge, there are virtually no studies specifically addressing this. In the present study, we used a combination of in vitro assays and intracellular overexpression to examine in detail the intrinsic function of a panel of disease-causing EVH1 missense mutations. As expected, missense mutants demonstrated variably impaired binding to WIP and enhanced degradation leading to reduced protein levels in cells, mimicking naturally occurring counterparts.12,13,15 Overexpression in WASBMDCs restored protein levels but did not restore full WASpWT function as measured by podosome formation, podosome volume, and migration. Furthermore, EVH1 mutants were retained in podosome sites for shorter periods and regulation of WASp protein activity was impaired both in terms of actin polymerization and tyrosine phosphorylation. These results demonstrate that: (1) the EVH1 domain is important not only for reducing WASp degradation, but also for regulating protein activity, and (2) that missense mutations in the EVH1 domain of WASp disrupt cellular function independently of protein levels.
WASp proteins bearing mutations in the EVH1 domain have been shown previously to retain biochemical function.21,29 To address this in more detail, we tested f-actin assembly mediated by 4 distinct missense EVH1 mutants and EVH1 domain–deleted WASp (dEVH1) using an in vitro bead-based assay described previously.22,23 This assay uses a physiologic whole-cell lysate as a source of actin and actin regulators rather than individual recombinant components. Surprisingly, we found that WASpEMM had a significantly enhanced ability to polymerize actin in vitro compared with WASpWT, which could not be explained by differences in binding to Cdc42, a key WASp activator.3 This may relate to a repressive effect of WIP on WASp-mediated actin polymerization through stabilization of autoinhibition, as has been suggested previously.7 However, deletion of the entire EVH1 domain (and therefore all WIP interaction) did not further enhance f-actin assembly, but rather resulted in substantial loss of actin-polymerizing capacity, suggesting that binding of other factors to parts of this region may also be important for regulation of activity.
It is already known that the WIP-WASp interaction is required for multiple WASp-dependent cell functions, including podosome and immune synapse formation.29,39 In addition to protecting WASp from degradation, WIP is also proposed to participate in localizing WASp family proteins to areas of actin polymerization,40 as evidenced by the presence of WIP-WASp complexes in podosomes and lamellipodia and the failure of WASp recruitment to these structures in the absence of WIP.29,41 It has been shown that podosome assembly is reduced in macrophages from patients harboring WASpV75M and WASpR86C EVH1 mutations,42 but analysis of intrinsic protein function is precluded by reduced WASp protein levels. Consistent with these studies, we also observed enhanced protein degradation resulting in reduced protein levels after expression of EGFP-tagged missense mutant proteins in cell lines, but, unexpectedly, loss of the whole EVH1 domain protected WASp from degradation, supporting the recent report that critical ubiquitination target sequences are contained in this region.37
To determine whether discrete EVH1 missense mutations confer disadvantage for intrinsic WASp function in vivo, we transduced primary WASp-deficient BMDCs using lentivirus to achieve expression of functional WASp levels equivalent to or greater than endogenous WASp. In this system, WASpEMM was unable to restore podosome formation to the same degree as WASpWT and assembled podosomes were significantly larger, suggesting impaired organization of podosome structure. These findings are unlikely to be simply because of reduced recruitment by WIP29 because WASpEMM retained significant localization to podosomes. Furthermore, WASpdEVH1 was capable of colocalizing with f-actin in podosome cores, implying that there are WIP-independent mechanisms for WASp recruitment and localization to sites of actin polymerization. Our findings suggest that other factors contribute to impaired podosome assembly. Even though WASpdEVH1 could form podosomes with characteristic morphology, it was also associated with large actin-vinculin–containing plaques as have also been observed after WASp overexpression in WIP-deficient cells,29 suggesting that WIP is an important regulator of WASp function after localization. This hypothesis is supported by our FRAP experiments demonstrating that WASpdEVH1 and WASpA134T (the missense mutant with the least WIP affinity) had a shortened half-life in the podosome compared with WASPT45M (which preserved the highest WIP affinity) and WASpWT, which were retained for longer. WIP therefore contributes to retention and stability of WASp at sites of actin polymerization. This finding in primary mammalian cells is in keeping with a recent observation in Drosophila that dissociation of the WIP-WASp complex, in that case by the Blown fuse protein, results in a rapid exchange of WASp from the myoblast podosome-like structure.43 Surprisingly, reduced retention of WASpEMM in podosomes did not alter podosome half-life or survival, implying that the WIP-WASp interaction is not essential for normal podosome dynamics.
Tyrosine phosphorylation of WASp at amino acid 291 in humans (293 in mice) has been shown to be important for both its regulation and stability. Although exceptions may occur, in vivo phosphorylation appears to occur downstream of Cdc42-induced release of WASp autoinhibition, suggesting that this modification takes place at membrane-associated sites of actin assembly.44,45 Constitutive tyrosine phosphorylation at this residue enhances WASp activity in vitro, but also targets WASp for accelerated degradation, complicating studies of effect in vivo.46 Conversely, the inability to phosphorylate this site reduces WASp-dependent actin polymerization in vitro and in vivo, conferring a WASp-deficiency state.46 In the present study, we have shown that despite normal serine phosphorylation, all WASpEMM and WASpdEVH1 showed reduced tyrosine phosphorylation in vivo and we hypothesize that this results from impaired WIP-mediated retention and spatial regulation at sites of actin polymerization. In addition, WASpEMM impaired the intrinsic regulation of WASp activity in vivo, as evidenced by increased total intracellular f-actin after stimulation. Consistent with our hypothesis that EVH1 missense mutants result in defects of spatially regulated actin polymerization, BMDCs expressing WASpEMM with minimal WIP binding exhibited impaired ability to polarize a dominant leading edge, as evidenced by the formation of excessive numbers of lamellipodia and reduced persistence of directed migration.36 WASpT45M, which preserves greater WIP interaction, had a lesser effect on leading edge formation and migration despite intrinsic protein dysfunction, possibly reflecting the sensitivity of these assays to specifically analyze WASp function.
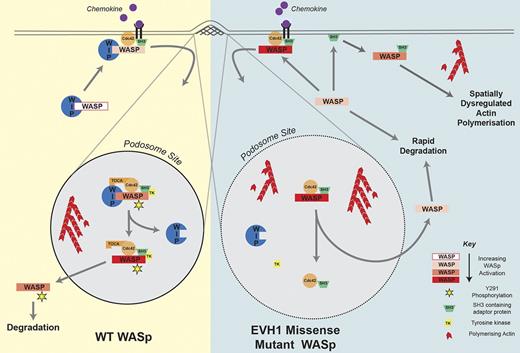
Our findings are consistent with the following model (Figure 7) in which WIP critically regulates the activation of WASp at specific membrane sites by 3 major mechanisms. First, WIP limits off-target WASp-mediated actin polymerization by favoring an auto-inhibited confirmation.7,8 Second, WIP functions to retain WASp for optimal times at sites of actin remodeling, which modulates activation though other WASp-binding partners such as Cdc42 (mediated through Toca-1)47 and SH3 adaptor proteins.48,49 These competitive influences, along with other modifications such as phosphorylation/dephosphorylation,34 allow WASp to switch between inhibited and active states after recruitment to the intended sites of actin polymerization. As shown recently, the dynamics of WASp retention are important for regulating Arp2/3-dependent actin nucleation, barbed end capping, and subsequent f-actin assembly.49 Third, WIP protects WASp from proteasome- and/or calpain-mediated degradation.27-29,50
Role of the WASp-WIP interaction in the formation of stable podosomes. WIP regulates the activation of WASp at specific sites such as podosomes by stabilizing the WASp autoinhibited conformation, retaining WASp at sites of active actin polymerization in the proximity of stimulatory binding partners and protecting WASp from degradation. In EVH1-mutated WASp, all 3 of these mechanisms are disrupted. See “Discussion” for further details.
Role of the WASp-WIP interaction in the formation of stable podosomes. WIP regulates the activation of WASp at specific sites such as podosomes by stabilizing the WASp autoinhibited conformation, retaining WASp at sites of active actin polymerization in the proximity of stimulatory binding partners and protecting WASp from degradation. In EVH1-mutated WASp, all 3 of these mechanisms are disrupted. See “Discussion” for further details.
This is the first comprehensive study of the intrinsic function of disease-causing EVH1 missense mutations. Although it has been postulated that the main role of WIP-WASp complex formation is to protect WASp from degradation and to localize WASp to sites of membrane associated actin polymerization,28,29,41 our data challenge this model by demonstrating that WASp can be localized to areas of new f-actin assembly even in the absence of WIP binding. Recruitment is likely to be mediated by the cooperative interaction of other binding partners such as Cdc42, Src kinases, and SH3 domain–containing adaptor proteins,45,48 whereas WIP plays a major role in retaining WASp within sites of active actin polymerization. In the present study, we have demonstrated that despite a preserved ability to polymerism actin, EVH1 missense mutants do not support normal intracellular WASp activity, even when expression levels are restored. Our findings suggest that treatment for XLT aimed simply at enhancing protein expression level may not fully restore biologic function and could potentially result in dysregulated actin assembly in vivo.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
The authors thank Professor Tim Cole (Professor of Medical Statistics, University College London Institute of Child Health, London, United Kingdom) for assistance with statistical analysis.
This work was supported by the Wellcome Trust (grant number 072725/Z/03/Z to A.J.J.W. and grant number 090233/Z/09/Z to G.B., D.M., and A.J.T.); the Primary Immunodeficiency Association and the Institute of Child Health Biomedical Research Centre (grant number 09MI29 to S.O.B.); Fundacao para a Ciencia and INOV Contacto, Portugal (to J.M.); the Pathological Society (to A.J.J.W.); the Mason Medical Foundation (to A.J.J.W.); the Human Frontier of Science Program (to M.F. and G.C. and a Young Investigator grant to G.C.); the Royal Society (University Research Fellowship to G.C.); and the Great Ormond Street Hospital Children's charity (to A.J.T.).
Wellcome Trust
Authorship
Contribution: A.J.J.W., J.M., G.B., D.M., M.F., and B.V. performed the research, collected, analyzed, and interpreted the data, and performed the statistical analysis; G.C., G.O.C.C., A.J.T., and S.O.B. designed and supervised the research; A.J.J.W. and S.O.B. wrote the manuscript; and all authors reviewed the manuscript and had access to primary data.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Dr Austen Jonathan Jacob Worth, Molecular Immunology Unit, Institute of Child Health, University College London, London, WC1N 1EH United Kingdom; e-mail: a.worth@ucl.ac.uk.