Abstract
The genomic landscapes of acute myeloid leukemia (AML), myelodysplastic syndrome (MDS), myeloproliferative disorders (MPD) and other related myeloid malignancies are now amongst the best characterized cancer genomes. These malignancies share most of their somatic driver mutations, many of which have therapeutic and prognostic significance (Patel et al, NEJM 2012). Patient prognostication and clinical decision-making can be greatly facilitated by testing for these mutations in parallel with established diagnostic assays. Here, we describe and validate RISCL (Rearrangements, Indels, Substitutions, Copy number and Loss-of-heterozygosity), a novel methodological and bioinformatic tool for the molecular diagnosis of myeloid malignancies. This tool employs targeted DNA capture to simultaneously: 1) identify coding mutations in 49 genes, 2) detect the four most important translocations in AML and 3) derive genome-wide copy number and zygosity data.
Samples & methods
1. Samples
Genomic DNA was extracted from bone marrow samples of 62 patients with AML (n=86 samples, including 24 remission samples) and 68 patients with MDS; and from blood granulocytes and mononuclear cells from 5 cord blood samples and 18 adults with normal hematopoiesis.
2. cRNA baits and sequencing
The bait library (Agilent) contained 53,613 probes to capture: 1) all exons from 49 genes 2) intronic breakpoint sites for PML-RARA, CBFb-MYH11 and RUNX1-RUNX1T1 and MLL breakpoints 3) 9958 SNPs (minor allele frequency 0.40-0.45) for genome wide copy number and zygosity analysis. Barcoded sequencing was performed using Illumina HiSeq 2000 (100bp paired-end).
3. Bioinformatic analysis
We used bespoke bioinformatics for detecting coding substitutions and indels (MIDAS; Conte et al, Leukemia 2013), chromosomal translocations (SMALT-FIT), copy number analysis (Avadis software) and detection of specific mutations such as MLL-PTD and FLT3-ITD (in-house scripts).
4. Verification of results
To validate the sensitivity and specificity of our approach, we compared our findings to conventional diagnostic data and are also validating 30% of randomly selected variants.
Results
A mean of 94% of targeted bases were covered at least by 30x. In AML samples, the four most common coding mutations identified affected NPM1 (n=10), CEBPA (n=8), IDH1 (n=8) and NRAS (n=7). By comparison to conventional diagnostics, we detected 5/5 IDH1R132, 4/4 CEBPA, 1/1 IDH2R172K and 8/9 NPM1 mutations. In MDS samples, the top four mutations affected TP53 (n=15), TET2 (n=13), SRSF2 (n=9), ASXL1 (n=8) and mutations affecting spliceosome genes (n=18) that were mutually exclusive, as previously described (Yoshida et al, Nature 2011).
RISCL detected 100% of known translocations (28/28) in AML patients, namely CBFb-MYH11 (n=8/8), PML-RARA (n=9/9), RUNX1-RUNX1T1 (n=4/4) and rearrangements of MLL (n=7/7). In every case of MLL rearrangement the gene partner was identified and in one case with t(X;11) we identified a novel gene partner to MLL, DIAPH2. Furthermore, we identified one patient with an MLL rearrangement not identified at diagnosis.
Copy number analysis efficiently detected known large chromosomal deletions or monosomies in chromosome 5 (18/18) and 7 (10/12). Overall 47/54 large deletions were detected using Avadis software. Furthermore in one MDS patient we were able to detect a submicroscopic heterozygous deletion in chromosome 4 which included TET2. However this method was much less sensitive for detecting trisomies (13/27 trisomies detected overall). The reasons for this disparity between detection of deletions and amplifications using a standardized depth of coverage algorithm are unclear, but may include subclonal mutations, selection of karyotypically abnormal cells during metaphase preparation or limitations of our bioinformatic analysis, which we are currently investigating.
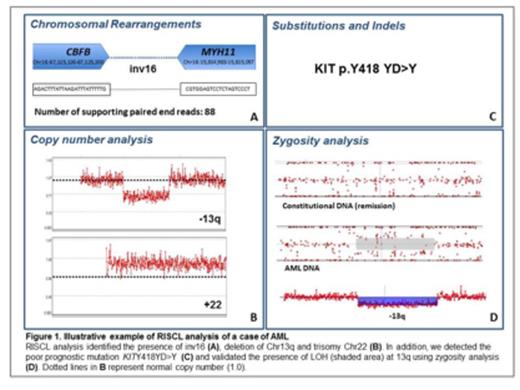
Figure 1 shows an example of the results of our holistic analysis using RISCL from an informative case of AML.
In summary we describe RISCL, a novel powerful holistic NGS tool for detailed characterization of myeloid malignancies that can be used for patient stratification and a personalized approach to malignancy in the molecular era. The same approach can be extended to other malignancies.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.