Abstract
Acute myeloid leukemia (AML) is a highly heterogeneous disorder characterized by the rapid clonal proliferation of blasts derived from hematopoietic progenitor cells, leading to failure of normal hematopoiesis. Although standard therapy, usually including idarubicin and cytarabine, has been used to achieve remission, the long-term survival rates remain low.
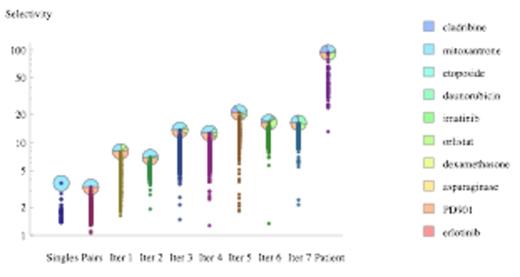
To show that it is possible to improve existing AML standard therapy, we have used experimental algorithms we have recently developed (iterative algorithms that find effective combinations without a full combinatorial screening) to find selective drug combinations for AML cells. The first search was done using the AML cell line KG-1 for the initial steps and cells from an AML patient for the final step. A normal fibroblast cell line (IMR-90) and normal blood progenitor cells were used as controls for toxicity. Drugs were chosen from those already approved or in clinical trials for AML. As shown in Figure 1, this experimental search was able to achieve selective killing of AML patient cells with a ratio close to 100:1 with respect to non-cancer control cells.
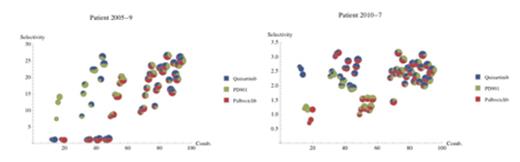
To refine the choice of drugs to be used in the search, we recently developed a network model of AML intracellular signal transduction (see www.leukemianetworks.org). The present version (AML 2.1) includes 5667 proteins and 22,218 interactions. The network can be used to integrate genomic and gene expression information from individual patients and to provide a shortlist of drugs to be tested further. Using literature data and our network model we identified three compounds as promising for AML drug combinations: the MEK inhibitor PD0325901, the FLT3 inhibitor Quizartinib and the CDK 4/6 inhibitor Palbociclib. For this experiment, normal progenitor blood cells were the controls. Figure 2 shows the results of 64 experimental tests of the response to different combinations of these drugs. The two AML patients clearly exhibited different drug sensitivities, further supporting the necessity of personalized drug combinations.
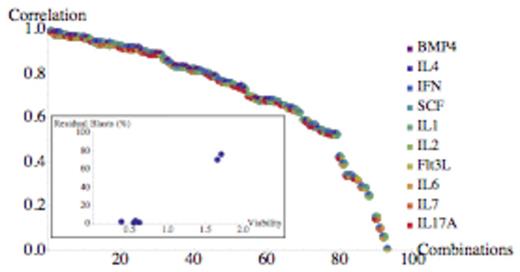
To improve the clinical relevance of these in vitro studies, we investigated the correlation between clinical and in vitro response to standard AML therapy, using a set of media containing 16 cytokines/growth factors in different combinations. These molecules have been selected using three methods: ligands acting on receptors that are part of gene signatures with prognostic significance in AML; cytokines that promote hematopoietic differentiation of stem cells; and ligands of receptors identified as playing an important role in AML in a recent RNAi screen. During several iterations of our combinatorial search algorithm we studied 185 different media. Figure 3 shows the correlation between clinical response (residual blasts after 28 days of therapy) and response to the same therapy in vitro after 72 h achieved in the last iteration of the search, in a study of six patients. The medium with the highest correlation had a correlation coefficient of 0.99 (p=0.0002). The in vitro response in this medium is clearly able to distinguish patients that respond to standard therapy vs those that do not, as can be seen from the inset of Fig. 3. Some cytokines (BMP-4 and IL-4) were always absent in the group of 14 media with lower correlation shown on the right of Figure 3. The remarkable variation in correlation with clinical response of these media, spanning correlation coefficient values from 0.99 to 0.004, clearly show the importance of optimizing the in vitro microenvironment for prediction of patient drug response.
In summary, we identified drug combinations that selectively kill AML primary cells and we optimized media that improve the in vitro prediction of clinical drug response. These methods can together assist in the personalization of AML therapy using patientÕs cells.
Selective drug combinations for AML primary cells. Selectivity is the ratio between AML and control cell viability.
Selective drug combinations for AML primary cells. Selectivity is the ratio between AML and control cell viability.
Selective drug combinations for AML primary cells from two patients, using multiple doses and combinations of Quizartinib, PD-0325901 and Palbociclib. Patient 2005-9 responded to MEK inhibitor PD-0325901 whereas Patient 2010-7 responded to FLT3 inhibitor Quizartinib.
Selective drug combinations for AML primary cells from two patients, using multiple doses and combinations of Quizartinib, PD-0325901 and Palbociclib. Patient 2005-9 responded to MEK inhibitor PD-0325901 whereas Patient 2010-7 responded to FLT3 inhibitor Quizartinib.
A pilot study showing the clinically predictive power of optimized media. The inset shows the best medium. Please note the strong effect of media composition on clinical correlation.
A pilot study showing the clinically predictive power of optimized media. The inset shows the best medium. Please note the strong effect of media composition on clinical correlation.
Piermarocchi:Salgomed Inc.: Equity Ownership. Paternostro:Salgomed Inc.: Equity Ownership.
Author notes
Asterisk with author names denotes non-ASH members.