Abstract
Novel gene mutation discovery has resulted in the increasing utility of targeted therapies. This is of particular relevance where traditional therapies have failed, resulting in increasing drug resistance and genetic instability. The incremental rise of subclonal populations of drug resistant cells is well recognized in CLL, however exactly how these subclones contribute to the overall disease course of the patient is unknown. Critical to further understanding the relevance of early minor subclones is the determination of the genetic profiles of these subclones and the identification of potential driver mutations.
While a high level of resolution of genetic mutations can be revealed using ultra-deep next generation sequencing of CLL cells, this method does not determine which actual subclone contain the mutations, and requires approximately 2000 fold coverage.
One of the most important prognostic markers in CLL is a deletion of the short arm of chromosome 17 (del17p), which includes deletion of TP53 gene. Whilst del17p is uncommon at diagnosis (only 5%-10% of all CLL patients), this proportion significantly increases to rougly 40-50% of chemo-refractory CLL. Therefore, we hypothesise that there are specific mutations in the del17p cells, including but not limited to TP53, which drive these subclones through clonal evolution, creating genetically unstable cells which are then refractory to treatment. We are particularly interested in those cases of CLL that carry a low frequency del17p subclone (<20% CLL cells), as these patients represent the greatest challenge to clinicians to decide the most appropriate course of treatment.
Current methods to detect these 17p-deleted cells, such as microscopy-based fluorescence in situ hybridization (FISH) and karyotyping, have restrictions on their lower limit of detection due to the low number of cells targeted. We have developed a sensitive method of detecting and flow sorting del17p cells to facilitate specific subclone analysis. FISH in suspension (FISH-IS) incorporates a flow cytometry-based imaging approach with automated analysis of thousands of cells, and is highly applicable to detecting del17p in CLL samples.
Methods:
The FISH-IS workflow was used with 17p locus-specific identifier (LSI) probes in CLL samples. A fluorescently labelled contig of multiple BAC clones covering the TP53 region was hybridised to CLL cells in suspension. Data was collected through the Image Stream X flow cytometer (Amnis) and IDEAS software was used to carry out the analysis.
Results:
In preliminary experiments CLL cells were mixed in fixed ratios with wild type 17p cells (wt 17p). We have shown that FISH-IS is able to accurately enumerate the 17p allele status (monoallelic vs biallelic) based on fluorescence intensity. Furthermore, the sensitivity of detection of del17p cells amongst 20,000 analysed cells was precisely identified to a 5% limit (Figure 1).
The second phase involved developing a methodology capable of enriching del17p low-frequency subclones in CLL samples by standard flow cytometry. Flow cytometry was used to sort cells based on their mean fluorescence intensity. Analysis of common polymorphisms within TP53 were used to demonstrate enrichment by collecting predefined fractions from the flow cytometer, based on fluorescence intensity and predicted 17p deletion status.
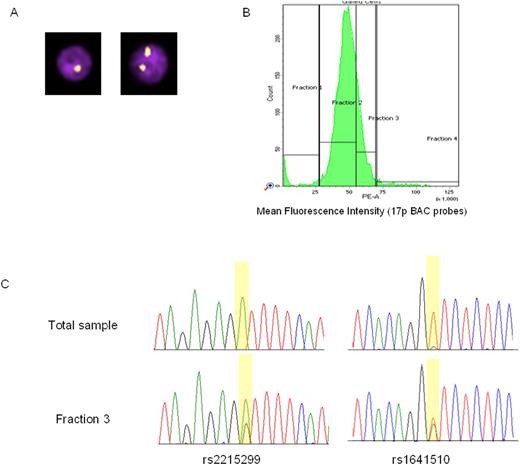
We confirmed this method on CLL samples carrying high-frequency del17p clones due to sample availability. Our data clearly shows that this method is able to enrich for the low frequency clone as evidenced by analysis of targeted heterozygous SNPs located in the deleted region of 17p (Figure 2).
Further sample analysis and exome sequencing is underway to determine sub-clonal mutation architecture. Original findings in this specific and novel approach to sub-clone analysis will be presented.
Conclusion:
This is the first time the genomic landscape of these low-frequency subclones has been interrogated in an unbiased manner. This data will enable a specific and in-depth genetic analysis of the untreated low-frequency del17p subclone, with a view to being able to identify the mechanisms of development of a chemorefractory and aggressive CLL phenotype.
Sensitivity of FISH-IS with a predictable mixing model.
Figure 2: Successful enrichment of low frequency CLL subclones based on 17p status. (A) FISH-IS images. (B) Flow sorting. (C) Validation of enrichment by SNPs within TP53.
Figure 2: Successful enrichment of low frequency CLL subclones based on 17p status. (A) FISH-IS images. (B) Flow sorting. (C) Validation of enrichment by SNPs within TP53.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.