Abstract
Abstract 2435
DLBCLs are the most prevalent lymphomas in adults and great advances have been made in understanding molecular effects on tumor cells as well as tissue environment, leading to determining gene prognosis signatures using transcriptional profiling techniques.
As blood cells interact with cells in almost all tissues in the body, blood-derived total RNA gene expressions have been investigated for the past years including for solid cancers [Clin Cancer Res. 2006:3374.], infections [Nature 2010;446:973.] and autoimmune disorders [Genes Immun. 2010;11:269., Immunity 2008;29:150.]. Blood-based microarray approaches were able to identify differentially expressed genes distinguishing patients from healthy volunteers.
Interested in the potential of this noninvasive and easy-to-use technique, we hypothesized that aggressive DLBCLs at diagnosis cause molecular perturbations on the whole blood allowing identifying gene expression differentiation compared to healthy controls.
Whole blood was collected into PAXgene™ Blood RNA tubes ensuring blood stabilization and sent within 24 hours to be stored at −80°C before extraction. Our study involved high-quality RNA samples from 75 DLBCL patients taken at diagnosis prior to any anti-cancer treatment and 87 healthy volunteers, sex and gender matched. All patients were less than 60 and enrolled in a multicentric & prospective clinical trial for aggressive form of DLBCL, GOELAMS 075, which compares the autologous stem cell treatment to regular R-CHOP procedure. The median age was 52 for patients and 48 for controls.
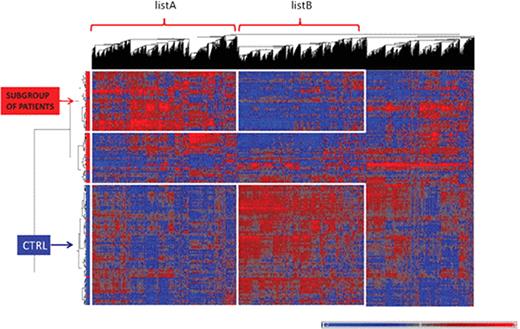
Gene expression profiling (GEP) was assessed using Affymetrix GeneChip® Human Exon 1.0 ST arrays. Unsupervised hierarchical clustering analysis (HCA) distinguished DLBCLs from controls. Two gene lists were identified based on HCA (Figure 1): listA consisted in 3,323 upregulated genes for a subgroup of patients and inversely, listB in 2,966 upregulated genes for controls. Canonical pathways were generated for both lists for genes meeting p<5% and FC >1.2 through the use of IPA (Ingenuity® Systems). The upregulated genes for patients (listA) were found associated with cytokine signaling pathways (Interleukins, NF-κB) while the down-regulated genes (listB) were implicated in T lymphocytes signaling pathways. Further investigations of the dataset by univariate analysis (Mann-Whitney test, FDR<5%, FC >1.5) found 1047 differentially expressed genes, confirming the systemic alteration. A set of 20 genes, selected as the best predictive genes for which the misclassification error rate is minimal, was able to discriminate DLBCLs from control samples (sensitivity= 88% & specificity=95%). No correlation was found between genes and biological parameters such as hemoglobin, leucocytes, lymphocytes, platelets or polynuclear neutrophils. The down-regulated genes were located in the nucleus and involved in transcription deregulation, DNA repair and apoptosis. Upregulated genes were related to the immune response as well as the inflammatory response with for instance S100 proteins which are implicated in myeloid-derived suppressor cell biology.
Despite the complex mixture of cell types in blood, whole blood has shown strong systemic perturbations in DLBCLs at diagnosis. Biological investigation indicated an over-expression of the inflammation and immune responses combined to perturbations of the T-lymphocyte pattern. Our findings concerning inflammation-related gene expression including NF-κB activation and upregulated cytokine transcripts, with for instance IL-1, IL-6 & IL-10, invite us to determine whether a specific DLBCL-induced inflammation process exists compared to other nonmalignant diseases [Clin Microbiol Rev. 2002 Jul; 15(3):414-29]. Comparison to other lymphoma and inflammatory diseases as well as with tumor status are under way allowing to better characterizing DLBCL-specific biomarkers.
These results shed new lights on DLBCL biology deciphering disease's heterogeneity at the RNA whole blood level. They encourage us to investigate whole blood GEP for prognosis and as a new parameter useful for disease classification.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal