Abstract
BCR-ABL1 mutation testing is recommended for chronic myeloid leukemia (CML) patients who have suboptimal response and/or treatment failure with tyrosine kinase inhibitor (TKI) therapy. BCR-ABL1 mutations in the kinase domain (KD) of ABL1 account for at least 40-50% of all TKI resistant cases. Thus, detection of low-level mutations after development of resistance may offer critical information to guide subsequent therapy selection. The current gold standard for BCR-ABL1 mutation detection is Sanger sequencing (SS), which has an analytical sensitivity of approximately 10-20%. In this study, our aim was to detect low level BCR-ABL1 variants in follow up samples of CML patients with TKI resistance using next-generation sequencing (NGS) approach.
Eight patients with CML who were resistant to imatinib had been routinely sequenced with SS for BCR-ABL1 KD mutations between December 2009 and December 2012. We then retrospectively analyzed these samples with NGS. RT and long range PCR was performed to amplify BCR-ABL1 fusion transcripts and the PCR products sequenced bidirectional after library preparation. We performed a fusion transcript based BCR-ABL1 mutation assay using Roche 454 amplicon deep-sequencing technology that is suited for detecting low level variants in pooled amplicon samples. Sequencing data was analyzed with GS Amplicon Variant Analyzer (AVA) software, and the variant frequency cut-off was adjusted to 1%.
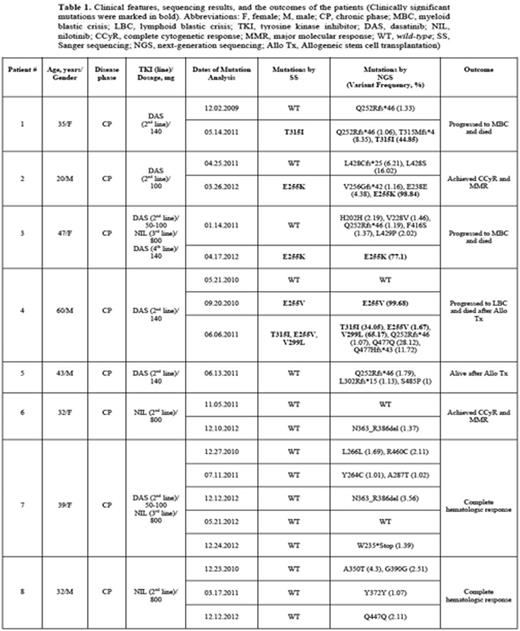
Clinical features, sequencing results, and the outcomes of the patients were summarized in Table 1. Four patients were male, and the median age was 37 years (range, 20-60 years). The patients were all in chronic phase at the time of the diagnosis. After imatinib resistance, 4 patients had received dasatinib (DAS), and 2 were given nilotinib (NIL) as second line TKI treatment. The remaining two patients had both received DAS and NIL (Table 1). In a set of 20 clinical samples, at different time points, NGS not only identified all the mutations detected by SS, but additionally identified low level variants present between 1 – 28.12 %. T315I and E255K/V were the most common mutations, which were detected in four patients, both by SS and NGS at the same time points (Table 1). Two patients (patient #1 and #4) had T315I, and they both progressed to blastic phase and died. E255K was detected in patients #2 and #3, and patient #2 had achieved and maintained complete cytogenetic and major molecular responses with 100 mg daily DAS, whereas patient #3 had received both NIL and DAS, but she was deceased due to myeloid blastic crisis. Among 4 patients (patients #5, #6, #7, and #8), mutation analysis was performed at eleven different time points, and these patients were wild-type with SS. We also did not detect any clinically significant mutations in these patients by NGS. Most probably mechanisms other than KD mutations were responsible for the TKI resistance among these four patients.
Polyclonal mutations in BCR-ABL1 KD are commonly identified in TKI resistant patients. Thus, detection of low-level mutations after development of resistance offers critical information to guide subsequent therapy selection. An inappropriate kinase inhibitor selection could highly increase the risk of treatment failure with clonal expansion of the resistant mutant. In our imatinib resistant cohort, we detected low level variants accompany to known mutations which may constitute background genetic variations. Although we had expected to detect mutations earlier by NGS (i.e. before these mutations can be detected by SS), we did not observe such finding in our patients. The patients' samples may not show a stable mutation spectrum between time points. Hence, it is not always possible to spot a mutation before patients show resistance to therapy. Regular NGS analysis might detect these mutations in earlier phases, which might help clinicians to choose the most suitable individual treatment modality for the patients.
The authors would like to thank the Interlaboratory Robustness of Next-generation sequencing (IRON) Phase II study group members, especially to Simona Soverini and Alexander Kohlmann who designed BCR-ABL primers and plates. We also would like to thank the Research Fund of the Istanbul University (Project no. 24244) and Turkish Society of Hematology for supporting the study.
Sayitoglu:Roche Diagnostics: Research Support Other.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal