Background: Circulating tumor DNA (ctDNA) is an emerging biomarker in non-Hodgkin lymphomas (NHLs). Current methods for ctDNA minimal residual disease (MRD) are limited by two factors - low input DNA amounts and high background error rates. VDJ sequencing (i.e., IgHTS) has low background but is limited by low cell-free DNA (cfDNA). Tracking multiple mutations via CAPP-Seq improves sensitivity, but detection is limited by background errors. Clustered mutations have been described in multiple cancers including NHLs and potentially have lower error rates. We explored clustered mutations from whole-genome sequencing (WGS) to identify 'phased variants' (PVs), defined as multiple mutations on a single DNA molecule (Fig 1A). We designed a method to capture PVs for improved ctDNA detection and explored its utility for MRD in DLBCL.
Methods: We reanalyzed WGS from 1455 tumors across 11 cancer types. We identified genomic regions recurrently containing PVs and designed an assay for deep cfDNA sequencing. We applied this assay to 171 patients with large B-cell lymphomas. We compared the performance of PVs for disease detection to current ctDNA techniques, including CAPP-Seq and duplex sequencing.
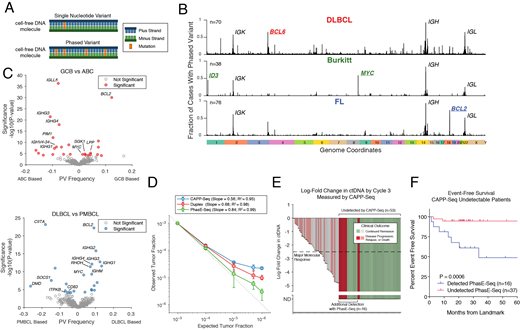
Results: To utilize PVs, mutations must occur within a typical cfDNA strand (~170bp). We measured the frequency of putative PVs in WGS, focusing on pairs of mutations occurring within <170bp. PVs were more frequent in NHLs than any other histology (median: DLBCL, 642; FL, 307; Burkitt, 89.5; CLL, 34; breast, 46; lung, colorectal, melanoma, bladder, cervical, head & neck < 10 per case; P < 0.001 for NHLs vs others). PVs in NHLs were enriched in single base substitution mutational signatures associated with activation-induced cytidine deaminase (AID) (SBS84 & 85). PVs in NHLs occurred in stereotyped regions, including canonical AID targets such as IGH, IGK, and IGL, as well as 44 other AID targets (Schmitz, NEJM 2018) (Fig 1B). We additionally identified novel regions not previously implicated as targets of AID, including LPP, XBP1, BZRAP1, and HLA-DQ.
We designed an approach for enriching PVs from ~115kb (Phased variant Enrichment Sequencing, PhasE-Seq) and other regions recurrently mutated in B-NHLs. We compared PhasE-Seq and CAPP-Seq using tumor and plasma samples from 16 patients. Compared to CAPP-Seq, PhasE-Seq yielded more SNVs and PVs per case (median SNVs: 331 vs 114, P<0.001; PVs: 729 vs 222.5, P <0.001). We next applied PhasE-Seq to 171 patients with untreated lymphomas (DLBCL, 148; primary mediastinal B-cell lymphoma, PMBCL, 23) profiling 58 tumor and 171 plasma samples with matched germline. We observed significant differences in the distribution of PVs between subtypes - for example, GCB-DLBCL had more PVs in BCL2, MYC, and SGK1, while ABC-DLBCL had more in PIM1 and IGHV4-34 (Fig 1C). Similarly, we noted enrichment in PVs in PMBCL in CIITA, SOCS1, CD83, and ITPKB.
We then compared PhasE-Seq to alternative methods for MRD detection. We used limiting dilutions of patient ctDNA down to 1:1,000,000 to establish the detection limit (LOD, Fig 1D). PhasE-Seq outperformed CAPP-Seq and duplex sequencing for recovery of expected tumor content, with a high degree of linearity down to ~1:1,000,000.
We applied standard CAPP-Seq and PhasE-Seq to patient cfDNA samples after two cycles of front-line therapy (n=92). We previously reported a 2.5-log reduction in ctDNA as prognostic at this time-point (Kurtz, JCO 2018). Using CAPP-Seq, 58% (53/92) of samples were undetectable. Using PhasE-Seq, 30% (16/53) of samples not detected by CAPP-Seq had evidence of MRD, with levels as low as 2:1,000,000. In patients with ctDNA undetected by CAPP-Seq, detection by PhasE-Seq significantly stratified outcomes (Fig 1E).
Conclusions: PVs are frequent in NHLs, likely due to AID, and correlate with disease biology. PhasE-Seq allows for superior detection of ctDNA, including MRD detection in the majority of patients after 2 cycles. Targeted sequencing of ctDNA should consider PVs to maximize detection and guide precision approaches.
Figure 1: A) Structure of phased variants B) Distribution of putative PVs from WGS data C) Genomic enrichment in PVs in lymphoma subtypes D) Dilution series comparing PhasE-Seq, CAPP-Seq, and duplex sequencing E) Waterfall plot showing ctDNA level vs outcome; undetectable ctDNA by CAPP-Seq is highlighted F) EFS of patients with undetectable ctDNA by CAPP-Seq after 2 cycles, stratified by PhasE-Seq
Kurtz:Roche: Consultancy. Diehn:Roche: Consultancy; AstraZeneca: Consultancy; Novartis: Consultancy; BioNTech: Consultancy; Quanticell: Consultancy. Alizadeh:Pharmacyclics: Consultancy; Janssen: Consultancy; Genentech: Consultancy; Roche: Consultancy; Gilead: Consultancy; Celgene: Consultancy; Chugai: Consultancy; Pfizer: Research Funding.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal