Abstract
Introduction
Acute myeloid leukemia (AML) is an aggressive disease with genetic and phenotypic heterogeneity that results in a highly variable response to standard chemotherapy. Azacitidine (AZA) is a hypomethylating agent (HMA) and has been investigated in combination with intensive chemotherapy as an epigenetic primer to sensitize leukemic cells to treatment. In a phase 1 trial, this regimen was safe and well-tolerated with overall response rate (CR+CRi) of 61% and complete remission rate of 41% (Cahill et al, Blood Adv 2020). Predictive biomarkers for response to this treatment strategy have not yet been identified. Since 5-hydroxymethylcytosine (5hmC) is an epigenetic biomarker in cancer, we hypothesized that Nano-5hmC-Seal sequencing technology may serve as a novel approach to identifying 5hmC profiles predictive of treatment response to epigenetic priming.
Methods
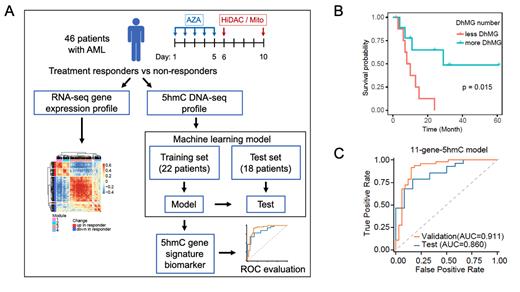
We performed RNA-seq gene expression and Nano-5hmC-Seal DNA profiling from peripheral blood/bone marrow samples of patients with high-risk AML to identify potential 5hmC profile biomarkers and gene expression changes (Figure 1A). Patients (n=46) were treated in a 3+3 dose-escalation scheme of AZA (37.5 mg/m 2, 50 mg/m 2, or 75 mg/m 2) on days 1-5 followed by high-dose cytarabine (3000 mg/m 2) and mitoxantrone (30 mg/m 2) (AZA-HiDAC-Mito) on day 6 and day 10 in a phase 1 trial previously reported (Cahill et al, Blood Adv 2020). We compared pre-treatment RNA-seq gene expression and 5hmC DNA profiles between responders (CR+CRi) and non-responders, as well as between pre-treatment and after 5 days of AZA for individual patients. We used an XGBoost machine learning model in Python based on a training set of patients to develop a 5hmC gene signature to predict response to AZA-HiDAC-Mito in an independent test set of patients. We compared continuous variables with two-tailed Student's t-test and used the Kaplan-Meier method with log-rank test for survival analysis.
Results
Thirty-three patients (72%) had adequate RNA samples for RNA-seq gene expression analysis. Eighteen responded to treatment (CR +CRi) and were enriched with gene expression patterns involved in cell-cell interaction and activation of cell cycle, while non-responders (n=15) had a higher expression of leukemic stem cell (LSC) signatures. There was no difference in gene expression profile when comparing pre-treatment samples to day 5 samples after AZA exposure. From the 5hmC profiling [n=40 (87%) patients with adequate samples], increased 5hmC in LSC genes was associated with treatment resistance to AZA-HiDAC-Mito (p=0.044). The number of differentially hydroxy-methylated genes (DhMGs) increased with higher doses of AZA exposure suggesting a dose-dependent epigenetic effect from AZA. Patients with a greater number of DhMGs following 5 days of AZA treatment had improved survival (p=0.015) (Figure 1B). Using the 5hmC-based XGBoost machine learning model comparing 5hmC profiles between responders to non-responders from a training set of patients (n=22), we developed an 11-gene 5hmC pre-treatment signature (including SKP1, WNT8A, CYP2E1, and NBPF9) to predict treatment response. The model was highly effective in predicting response to therapy, with an area under the curve (AUC) of 0.86 in an independent test set of patients (n=18) treated with AZA-HiDAC-Mito (Figure 1C).
Conclusion
In patients with AML treated with AZA-HiDAC-Mito, a pre-treatment LSC gene expression signature enriched with 5hmC was associated with treatment resistance. More DhMGs at day 5 appear to be a dose-dependent epigenetic effect that is induced by AZA and is associated with longer survival despite the absence of an immediate change in gene expression levels. An 11-gene 5hmC pre-treatment signature may be a predictive biomarker for AZA-HiDAC-Mito therapy and other HMA-based approaches. These findings warrant validation in a larger prospective trial.
Zhang: Bristol-Myers Squibb: Current Employment. Stock: Pfizer: Consultancy, Honoraria, Research Funding; amgen: Honoraria; agios: Honoraria; jazz: Honoraria; kura: Honoraria; kite: Honoraria; morphosys: Honoraria; servier: Honoraria; syndax: Consultancy, Honoraria; Pluristeem: Consultancy, Honoraria. Odenike: Celgene, Incyte, AstraZeneca, Astex, NS Pharma, AbbVie, Gilead, Janssen, Oncotherapy, Agios, CTI/Baxalta, Aprea: Research Funding; AbbVie, Celgene, Impact Biomedicines, Novartis, Taiho Oncology, Takeda: Consultancy. He: Epican Genetech: Current holder of individual stocks in a privately-held company, Current holder of stock options in a privately-held company.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal