Abstract
We studied the efficiency of indirect tumor cell purging via enrichment of CD34+ hematopoietic progenitor cells from leukapheresis products (LP) in breast cancer patients based on immunomagnetic selection of CD34+ cells. Detection of tumor cells was made by immunocytochemical staining. In addition, we evaluated the capacity of cytokeratin 19 (CK19)- and a novel epidermal growth factor receptor (EGF-R)-specific reverse transcriptase–polymerase chain reaction (RT-PCR) for monitoring tumor cell depletion. LP from 13 breast cancer patients were analyzed. Twenty-three CD34 selection procedures were performed. A median of 1.4 × 1010 total nucleated cells ([TNC] range, 0.88 to 3.5 × 1010) with a median CD34 purity of 2.5% (range, 0.4% to 6.3%) were entered into the selection procedure. Immunomagnetic CD34 enrichment resulted in a median purity of 83.3% (range, 45% to 95.4%) and a median recovery of 73.2% (range, 22% to 95%). Retransfusion of CD34-selected cells after high-dose chemotherapy resulted in a rapid and sustained hematologic recovery, reaching an absolute neutrophil count of 500/μL at day +10 and platelet count of 20,000/μL at day +11. Tumor cell depletion was quantified by immunocytochemical detection of CK19-positive cells. By this method, a median tumor cell depletion of 1.9 log (range, 0.7 to <3 log) could be demonstrated. Immunocytochemical detection of tumor cells was more sensitive than RT-PCR, yielding positive results in 81% of LP (17 to 21) versus 58% positive LP (10 of 17). However, EGF-R–based RT-PCR was much more sensitive than CK19-based RT-PCR (10 of 17 v 1 of 17). Despite highly efficient CD34 selection, tumor cells were still detectable after CD34 enrichment using immunocytochemistry and EGF-R–specific RT-PCR. Thus, this novel EGF-R–specific RT-PCR appears to be of value as an additional method to detect contaminating breast cancer cells within LP.
RECENTLY, transplantation of CD34-enriched human hematopoietic progenitor cells has found widespread application in the setting of autologous transplantation. Using different methods for CD34 selection, it can be demonstrated that CD34-selected autografts are capable of long-term engraftment.1,2 The use of peripheral blood progenitor cell (PBPC) enrichment via CD34 selection in transplantation has enabled studies to investigate the capacity of ex vivo expansion of these cells.3 Furthermore, these cells can be used as targets for gene therapy strategies in malignant and genetic diseases.4-7 One of the potential benefits of CD34 enrichment of PBPC or bone marrow autografts is the concomitant depletion of tumor cells. So far, the relevance of residual tumor cells within the autograft is still unclear. Thus, the clinical importance of tumor cell purging of stem cell products remains open to discussion. However, it can be demonstrated that mobilization of stem cells by chemotherapy plus growth factor leads to concomitant tumor cell release into the peripheral blood.8 Gene-marking studies in patients with acute myelogenous leukemia, neuroblastoma, or chronic myelogenous leukemia have clearly demonstrated that contaminating tumor cells within the transplant contribute to relapse after autologous bone marrow transplantation or peripheral blood progenitor cell transplantation (PBPCT).9-11
Gribben et al12 observed that patients with follicular non-Hodgkin's lymphoma receiving autografts containing a t(14; 18) translocation had a higher probability of relapse than patients whose autografts were devoid of t(14; 18)-positive cells after purging. The sensitive and specific detection of tumor cells is a prerequisite for monitoring purging efficiency. Contaminating tumor cells have also been observed in autografts from breast cancer patients.8,13,14 Due to the fact that breast cancer does not appear to have a tumor-specific chromosomal aberration, tumor cell detection by polymerase chain reaction (PCR) is based on the amplification of tissue-specific transcripts (ie, genes specific for epithelial differentiation). Several genes have been analyzed for suitability as target genes for detection of minimal residual disease in breast cancer patients by PCR. The most prominent genes were muc-1,15 cytokeratin 8, and cytokeratin 18. However, cytokeratin 19 (CK19) appears to be a more suitable target gene,16 since it has been demonstrated that there is promiscuous expression of those previously mentioned in nonepithelial tissues. However, the application of CK19-specific RT-PCR as a sensitive method for tumor cell detection is hampered by the existence of CK19 pseudogenes, underlining the need for additional tissue-specific target genes. The aim of this study was to evaluate the immunomagnetic CD34 selection procedure with respect to the capacity for tumor cell purging, and to compare standard immunocytochemical staining, RT-PCR specific for CK19, and a novel epidermal growth factor receptor (EGF-R)-specific RT-PCR with regard to the ability to detect residual tumor cells.
SUBJECTS AND METHODS
Patient characteristics.Patients with metastatic breast cancer (stage IV) and high-risk patients (stage II/III ≥10 lymph nodes) were entered into the transplantation program after provision of informed consent. Induction and mobilization therapy consisted of three cycles of epirubicin (120 mg/m2), ifosfamide (7.5 g/m2), and granulocyte colony-stimulating factor ([GCS-F] 5 μg/kg subcutaneously, Neupogen; Amgen, Thousand Oaks, CA). PBPC harvest was started at a white blood cell count of more than 1,000/μL and a CD34 cell number more than 20/μL using an AS104 cell separator (Fresenius, Bad Homburg, Germany). Twelve liters of blood was processed using acid-citrate-dextrose (Fresenius) as an anticoagulant. Patients with metastatic breast cancer responding to conventional chemotherapy were treated by tandem autologous transplantation. A first cycle of high-dose chemotherapy consisting of melphalan (140 mg/m2) was followed by autologous stem cell rescue. After hematologic reconstitution, patients received a second high-dose treatment according to the ICE conditioning regimen (ifosfamide 14 g/m2, carboplatin 1.5 g/m2, and etoposide 1.5 g/m2) and subsequent stem cell transplantation. Patients received 5 μg/kg subcutaneously G-CSF posttransplant. Patient characteristics are shown in Table 1.
Patient Characteristics
| Patient No. . | Age (yr) . | Disease Stage . | Metastatic Sites . |
|---|---|---|---|
| 1 | 53 | IV | Liver, ST |
| 2 | 45 | IV | Lung, ST |
| 3 | 38 | IV | Bone, ST |
| 4 | 55 | III | — |
| 5 | 40 | II | — |
| 6 | 54 | IV | Liver, lung, ST |
| 7 | 49 | IV | Liver, bone |
| 8 | 61 | IV | Lung |
| 9 | 56 | IV | Liver |
| 10 | 36 | IV | ST |
| 11 | 46 | IV | Liver, lung |
| 12 | 37 | II | — |
| 13 | 54 | IV | Bone, lung, ST |
| Patient No. . | Age (yr) . | Disease Stage . | Metastatic Sites . |
|---|---|---|---|
| 1 | 53 | IV | Liver, ST |
| 2 | 45 | IV | Lung, ST |
| 3 | 38 | IV | Bone, ST |
| 4 | 55 | III | — |
| 5 | 40 | II | — |
| 6 | 54 | IV | Liver, lung, ST |
| 7 | 49 | IV | Liver, bone |
| 8 | 61 | IV | Lung |
| 9 | 56 | IV | Liver |
| 10 | 36 | IV | ST |
| 11 | 46 | IV | Liver, lung |
| 12 | 37 | II | — |
| 13 | 54 | IV | Bone, lung, ST |
Abbreviation: ST, soft tissue.
RNA preparation.For RNA preparation and subsequent PCR analysis, 1 to 2 × 106 mononuclear cells from leukapheresis products ([LP] after Ficoll-Hypaque separation) or after CD34 selection were lysed in TRI-zol reagent (GIBCO-BRL, Grand Island, NY) and sheared to homogeneity, following the one-step guanidinium isothiocyanate/phenol RNA preparation as originally described.17 Following phase separation by the addition of chloroform, the RNA contained in the aqueous upper phase was precipitated in isopropanol at −80°C, washed in 80% ethanol, and resuspended in RNAse-free water. To eliminate contaminating DNA, RNA was exposed to RNAse-free DNAse I (1 U/μg RNA; Boehringer, Mannheim, Germany) at 37°C for 30 minutes. After heat inactivation of the enzyme at 95°C for 5 minutes, the RNA was reprecipitated and resuspended. Degradation of genomic DNA was verified in PCR analysis using β2 -microglobulin primers spanning several exons18 and yielding a PCR product 700 bp larger than β2 -microglobulin mRNA in the presence of genomic DNA.
cDNA synthesis and PCR.Detection of tumor cells by PCR was performed using the primers specific for CK19 as reported by Datta et al.16 We have recently shown that breast cancer tumor cells can also be detected by an EGF-R-specific RT-PCR.19 RNA (10 μg) was incubated with 50 ng 15(dT) primer at 60°C for 10 minutes. After addition of deoxynucleotide mix (10 mmol/L), DTT (10 mmol/L), and first-strand reaction buffer, 100 U MMLV Reverse Transcriptase (GIBCO-BRL) was added to a final volume of 50 μL. The cDNA synthesis reaction was performed at 37°C for 40 minutes. After heat inactivation at 95°C for 5 minutes and subsequent maintenance at 4°C, 5 μL of each reaction product was subjected to PCR analysis. The PCR reaction was performed at 94°C for 40 seconds, 60°C for 1 minute, and 72°C for 40 seconds for 40 cycles, followed by a final 5 minutes at 72°C. All reagents used for PCR analysis were purchased from Promega Biotech, Inc (Heidelberg, Germany). Twenty microliters of the final PCR products was separated on a 1% agarose gel (Seakem ME-agarose; FMC Bioproducts, Rockland, ME). RNA preparation and PCR were performed in separate locations to avoid any contamination. The following primers were used in this study: CK19 (as described elsewhere), (bp 272 to 1342) 5′-AAGCTA-ACC-ATG-CAG-AAC-CTC-AAC-GAC-CGC-3′ (sense) and 5′-TTA-TTG-GCA-GGT-CAG-GAG-AAG-AGCC-3′ (antisense); EGF-R (bp 702 to 1175), 5′-TCT-CAG-CAA-CAT-GTC-GATGG-3′ (sense) and 5′-TCGCACTTCTTACACTTGCG-3′ (antisense); and β2 -microglobulin (bp 73 to 341), 5′-CGA-GCA-GAG-AATGGA-AAGTC-3′ (sense) and 5′-GAT-GCT-GCT-TAC-ATGTCTCG-3′ (antisense).
Immunocytochemical staining and quantitation.Mononuclear cells from LP were isolated by density centrifugation through Ficoll-Hypaque (Seromed, Hamburg, Germany), washed twice with phosphate-buffered saline (PBS), and resuspended at 1 × 106/mL. Thereafter, cells were attached to the slides. Each slide contained 10 spots, each 0.7 cm in size. A quantity of 5 × 104 cells in 50 μL were attached to poly-L-lysine–coated ICN slides on each spot for 1 hour in a humidified chamber at 37°C, air-dried, and stored at −20°C. Two slides with 5 × 105 cells per slide were analyzed. After CD34 selection, cells could be directly attached to the slides without prior Ficoll-Hypaque separation. After thawing, the slides were allowed to dry and fixed in acetone. The breast cancer cell line MCF-7 was used as a positive control for each immunostaining experiment. A murine pancytokeratin monoclonal antibody (MoAb) (A45; kindly provided by Dr U. Karsten, Max Delbrück Center for Molecular Medicine, Berlin, Germany) was used for immunocytochemical staining. For antibody staining, an alkaline phosphatase–anti-alkaline phosphatase staining procedure20 was applied. In brief, the slides were preincubated with nonspecific human IgG (5 mg/mL) for 30 minutes at room temperature. After three washing steps with Tris hydrochloride, one drop of a solution containing the primary antibody in 10% ABO serum and 1% bovine serum albumin/PBS per well was applied. Tris hydrochloride instead of the primary antibody served as a negative control. Subsequently, a secondary (rabbit-anti mouse, RAM; Dako, Hamburg, Germany) and tertiary (swine-anti rabbit, SAR; Dako) antibody was added, followed by wash steps. The slides were exposed to chromogenic substrate solution containing 0.3 mg/mL Naphthol-As-Tris-phosphate, 0.24 mg/mL Levamisole, and 0.4 mg/mL Fast-red ITR salt, and then counterstained with hematoxylin. All these chemicals were purchased from Sigma Chemical Co (München, Germany). Tumor cells were considered immunocytochemically positive when staining on greater than 70% of the cell membrane or cytoplasm, respectively, was observed, and cell morphology showed malignant features. Generally, an overall number of 1 × 106 mononuclear leukocytes (ie, two ICN slides) were analyzed, tumor cells per 106 mononuclear leukocytes were counted, and the total number of tumor cells was extrapolated. If greater than five immunocytochemically positive cells per 5 × 104 were detected, tumor cells were quantified.
CD34 selection.Enrichment of CD34 cells from LP was performed using the ISOLEX 300 device (Baxter Biotech, München, Germany) as previously described.21 22 After washing (3× RPMI 1640 containing 5% ACD-A and 1% human albumin at room temperature, 200g for 10 minutes), cells were incubated with anti-CD34 monoclonal antibody 9C5 (0.5 μg per 1 × 106 cells for 30 minutes at 4°C). Unbound antibody was removed by washing as mentioned above. Thereafter, sensitized cells were incubated with immunomagnetic beads coated with goat anti-mouse antibodies (Dynal, Oslo Norway). Subsequently, cells were captured by a magnet in the ISOLEX 300SA chamber. The bead-cell complex was cleaved by treatment with chymopapain. After a final washing, samples were drawn for immunophenotypic analysis, PCR, immunocytochemical tumor cell staining, and colony assays. The positively selected cell fraction was cryopreserved and stored within the vapor phase of liquid nitrogen.
Immunophenotype analysis and quantitative determination of CD34 cells.Immunophenotyping of cells was performed by three-color immunofluorescence using fluorescein isothiocyanate (FITC)-, phycoerythrin (PE)-, or PerCP-labeled antibodies. Samples of whole blood containing 106 nucleated cells were stained with directly conjugated monoclonal antibodies for 30 minutes at room temperature. Lysis buffer (FACS lysing solution, dilution 1:10; Becton Dickinson, Heidelberg, Germany) was added for 8 minutes. Subsequently, cells were washed three times with 1× PBS (400g for 10 minutes at 4°C). The following antibodies were used: CD34-PE, CD34-FITC, HLA-DR-PerCP, CD45-PE (Becton Dickinson), CD33-FITC, CD38-FITC, and Thy-1-FITC (Pharmingen, Hamburg, Germany). Immunofluorescence was measured using a FACScan flow cytometer (Becton Dickinson).
Quantitative analyses of CD34 cells in peripheral blood and in LP were performed by “live gating” on CD34+ cells falling into the lymphocyte/blast region coexpressing CD45 with a low side scatter. A minimum of 50,000 events and a minimum of 100 CD34+ events (within the live-gate region) were analyzed. CD34 enumeration was performed in triplicate.
Colony-forming unit assays.Hematopoietic colony formation was analyzed using standard methylcellulose CFU-GM colony assays (Stem Cell Technologies, Vancouver, Canada). For analysis of LP, 2 to 5 × 104 cells/mL were seeded into colony assays. Enriched CD34 cells were seeded out at 1,500 cells/mL. Analyses were performed in triplicate.
RESULTS
Mobilization of PBPC and enrichment of CD34 cells from LP.Thirteen patients with breast cancer were entered into the mobilization regimen. Ten patients had stage IV disease and extensive tumor burden; three patients were in the high-risk group (stage II/III >10 positive lymph nodes). The median age was 49 years (Table 1). Mobilization of CD34 cells was performed with a combination of myelosuppressive chemotherapy (ifosfamide 2.5 g/m2 days 1 to 3 and epirubicin 60 mg/m2 days 2 to 3) and G-CSF 5 μg/kg/body weight (Neupogen; Amgen). G-CSF was administered starting on day 4 until the end of stem cell harvest. Quantitative determination of CD34 cells within LP demonstrated a median percentage of 2.3% (Table 2). Aliquots were drawn from these preparations for immunocytochemical staining and PCR analysis. Since LP were purged, stem cell harvesting was also performed following the first cycle of induction therapy and subsequent cycles. As indicated, in the beginning, LP from some patients were directly processed after leukapheresis and CD34 selection was performed. Later on, LP were stored up to 48 hours at 4°C, combined, and processed together. Viability testing using trypan blue staining showed that more than 95% of cells were still viable after storage. In addition, no difference was observed with regard to colony-forming capacity after storage (data not shown).
CD34 Selection Efficiency
| Patient . | Cycle . | Before CD34 Selection . | After CD34 Selection . | ||
|---|---|---|---|---|---|
| . | . | TNC × 1010 . | % CD34 . | % CD34 . | % Recovery . |
| 1 | 1.7* | 5.6 | 92.2 | 89.2 | |
| 2 | 1.4* | 3.4 | 81.8 | 73.2 | |
| 3 | 1 | 1.0 | 2.5 | 81.9 | 64 |
| 2 | 2.6* | 1.2 | 83.3 | 38.8 | |
| 3 | 1.7 | 0.9 | 92.1 | 56.5 | |
| 4 | 2.5* | 4.3 | 70.7 | 85 | |
| 5 | 1 | 1.4 | 3.5 | 94.2 | 47.5 |
| 2 | 1.7 | 1.3 | 91.1 | 95 | |
| 6 | 1 | 1.04 | 3.03 | 88.2 | 67.7 |
| 2 | 1.2 | 2.6 | 89.8 | 95 | |
| 3 | 1.6 | 3.8 | 87.5 | 22 | |
| 7 | 3.1* | 2.8 | 82.6 | 95 | |
| 8 | 2.2* | 1.6 | 76.6 | 95 | |
| 9 | 2.5* | 2.1 | 68.8 | 29.5 | |
| 10 | 3.5* | 0.9 | 89.3 | 66.3 | |
| 11 | 1 | 1.3 | 2.6 | 82.2 | 30.9 |
| 2 | 1.2 | 1.4 | 89.3 | 81.1 | |
| 12 | 1 | 1.1 | 2.1 | 51.3 | 90 |
| 2 | 2.7* | 3.1 | 79 | 64 | |
| 13 | 1 | 1.1 | 1.1 | 45 | 80.8 |
| 2 | 0.88 | 6.3 | 65.1 | 30.7 | |
| 3 | 1.4 | 1.8 | 94.3 | 87.1 | |
| 4 | 1.3 | 0.4 | 95.4 | 92.7 | |
| Median (range) | 1.4 (0.88-3.5) | 2.5 (0.4-6.3) | 83.3 (45-95.4) | 73.2 (22-95) | |
| Mean | |||||
| (±SD) | 1.7 (±0.7) | 2.5 (±1.5) | 81.4 (±13.4) | 69.0 (±24.6) | |
| Patient . | Cycle . | Before CD34 Selection . | After CD34 Selection . | ||
|---|---|---|---|---|---|
| . | . | TNC × 1010 . | % CD34 . | % CD34 . | % Recovery . |
| 1 | 1.7* | 5.6 | 92.2 | 89.2 | |
| 2 | 1.4* | 3.4 | 81.8 | 73.2 | |
| 3 | 1 | 1.0 | 2.5 | 81.9 | 64 |
| 2 | 2.6* | 1.2 | 83.3 | 38.8 | |
| 3 | 1.7 | 0.9 | 92.1 | 56.5 | |
| 4 | 2.5* | 4.3 | 70.7 | 85 | |
| 5 | 1 | 1.4 | 3.5 | 94.2 | 47.5 |
| 2 | 1.7 | 1.3 | 91.1 | 95 | |
| 6 | 1 | 1.04 | 3.03 | 88.2 | 67.7 |
| 2 | 1.2 | 2.6 | 89.8 | 95 | |
| 3 | 1.6 | 3.8 | 87.5 | 22 | |
| 7 | 3.1* | 2.8 | 82.6 | 95 | |
| 8 | 2.2* | 1.6 | 76.6 | 95 | |
| 9 | 2.5* | 2.1 | 68.8 | 29.5 | |
| 10 | 3.5* | 0.9 | 89.3 | 66.3 | |
| 11 | 1 | 1.3 | 2.6 | 82.2 | 30.9 |
| 2 | 1.2 | 1.4 | 89.3 | 81.1 | |
| 12 | 1 | 1.1 | 2.1 | 51.3 | 90 |
| 2 | 2.7* | 3.1 | 79 | 64 | |
| 13 | 1 | 1.1 | 1.1 | 45 | 80.8 |
| 2 | 0.88 | 6.3 | 65.1 | 30.7 | |
| 3 | 1.4 | 1.8 | 94.3 | 87.1 | |
| 4 | 1.3 | 0.4 | 95.4 | 92.7 | |
| Median (range) | 1.4 (0.88-3.5) | 2.5 (0.4-6.3) | 83.3 (45-95.4) | 73.2 (22-95) | |
| Mean | |||||
| (±SD) | 1.7 (±0.7) | 2.5 (±1.5) | 81.4 (±13.4) | 69.0 (±24.6) | |
Indicates that two leukapheresis products were pooled for this selection procedure.
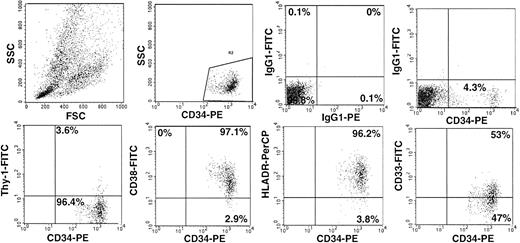
The immunomagnetic selection procedure involves chymopapain digestion to release the cells from the paramagnetic beads, and leads to cleavage of different cell-surface antigens. Therefore, we studied CD34 subset composition by three-color immunofluorescence in the LP (Fig 1). After immunomagnetic selection, two-color analysis was performed only for antigens known not to be affected by chymopapain treatment. A median of 1.4 × 1010 TNC was used for CD34 selection. Starting from a median purity of 2.5% (range, 0.4% to 6.3%) in the LP, immunomagnetic CD34 selection achieved a median final purity of 83.3% (range, 45% to 95.4%) and a median recovery of 73.2% (range, 22% to 95% (Table 2).
CD34 subset composition of PBPC. Analysis of CD34 subset composition within a representative LP is shown. A dot plot depicting the acquisition live gate is shown. Samples were run in triplicate. Cells were stained with CD34-PE, CD33-FITC, CD38-FITC, Thy-1-FITC, and HLA-DR-PerCP. The appropriate isotype-control MoAbs were used.
CD34 subset composition of PBPC. Analysis of CD34 subset composition within a representative LP is shown. A dot plot depicting the acquisition live gate is shown. Samples were run in triplicate. Cells were stained with CD34-PE, CD33-FITC, CD38-FITC, Thy-1-FITC, and HLA-DR-PerCP. The appropriate isotype-control MoAbs were used.
Analysis of CD34 subsets before enrichment disclosed a variable proportion of early immature CD34+/CD38− and CD34+/HLA-DR− populations. The amount of the Thy-1+/CD34+ cell population that has been described as highly enriched for long-term culture–initiating cells23 was found to be 8.7% (median; range, 2.6% to 38.5%). The proportion of lineage-committed CD34+/CD33+ cells was 52.2% (median; range, 19.8% to 95.3%). After the selection, CD34 subset analysis showed that the CD34+/CD38− population was recovered within the CD34+ fraction (Table 3). There might be a mild trend toward a lower number of CD34+/DR− cells after CD34 selection. Since enzymatic digestion leads to release of the cells, it cannot be ruled out that binding of the HLA-DR antibody to the HLA-DR antigen might be affected by chymopapain treatment of the cells. With regard to functional capacity, this appears to be of lesser significance, since the CD34+/DR− cell population contains fewer early hematopoietic progenitors than the CD34+/CD38− population. In addition, there is only small overlap between these two populations as determined by flow cytometry (ie, CD34+, CD38−, and DR−).24
CD34 Subsets (n = 23)
| Subset . | Before CD34 Selection . | After CD34 Selection . | ||
|---|---|---|---|---|
| . | Median (%) . | Range . | Median (%) . | Range . |
| CD34+/CD38+ | 98.7 | 91.1-99.9 | 98.2 | 90.3-99.9 |
| CD34+/CD38− | 1.3 | 0.1-8.9 | 1.8 | 0.1-9.7 |
| CD34+/DR+ | 96.2 | 83-98.9 | 99.2 | 97.7-99.6 |
| CD34+/DR− | 3.8 | 1.1-17 | 0.85 | 0.4-2.6 |
| CD34+/CD33+ | 52.2 | 19.8-95.3 | NA | |
| CD34+/33− | 47.8 | 4.7-80.2 | NA | |
| CD34+/Thy-1+ | 8.7 | 2.6-38.5 | ND | |
| Subset . | Before CD34 Selection . | After CD34 Selection . | ||
|---|---|---|---|---|
| . | Median (%) . | Range . | Median (%) . | Range . |
| CD34+/CD38+ | 98.7 | 91.1-99.9 | 98.2 | 90.3-99.9 |
| CD34+/CD38− | 1.3 | 0.1-8.9 | 1.8 | 0.1-9.7 |
| CD34+/DR+ | 96.2 | 83-98.9 | 99.2 | 97.7-99.6 |
| CD34+/DR− | 3.8 | 1.1-17 | 0.85 | 0.4-2.6 |
| CD34+/CD33+ | 52.2 | 19.8-95.3 | NA | |
| CD34+/33− | 47.8 | 4.7-80.2 | NA | |
| CD34+/Thy-1+ | 8.7 | 2.6-38.5 | ND | |
Abbreviations: ND, not determined; NA, not assessable due to chymopapain-sensitive epitope.
Autografting with immunomagnetically enriched CD34 cells.High-dose chemotherapy was administered as a tandem high-dose treatment with double-grafting. Induction therapy consisted of three cycles of epirubicin and ifosfamide. Patients responding to conventional chemotherapy proceeded to high-dose treatment. A first cycle of high-dose chemotherapy consisting of high-dose melphalan (140 mg/m2) and subsequent PBPC transplantation was given to all patients responding to conventional chemotherapy. After hematologic reconstitution, patients received an additional cycle of high-dose treatment with stem cell support according to the ICE protocol (14 g/m2 ifosfamide, 1.5 g/m2 carboplatin, and 1.5 g/m2 etoposide). This second cycle of high-dose chemotherapy was administered as soon as patients had reached a white blood cell count of more than 3,000/μL and a stable platelet count of more than 80,000/μL without support. Patients with progressive disease or major infectious complications after the first cycle of high-dose treatment did not receive a second course of high-dose chemotherapy. A median of 2.9 × 106 CD34+ cells/kg body weight were retransfused per cycle of high-dose chemotherapy. Rapid hematologic recovery and sustained engraftment could be observed in all patients transplanted. Engraftment data after high-dose melphalan and ICE therapy are shown in Table 4. Overall hematologic reconstitution after stem cell transplantation, ie, neutrophil recovery (ANC> 500/μL) and platelet recovery (>20,000/μL) occurred at day +10 and day +11, respectively (Table 4).
Hematologic Reconstitution Data
| Patient No. . | Chemotherapy . | CD34 × 106 per kg Body Weight . | Days Until ANC >500/μL . | Days Until . |
|---|---|---|---|---|
| . | . | . | . | Platelet Count >20,000/μL . |
| 2 | HD-Mel | 2.3 | 10 | 11 |
| 3 | HD-Mel | 2.7 | 11 | 11 |
| 4 | HD-Mel | 2.04 | 12 | 11 |
| ICE | 4.8 | 9 | 11 | |
| 5 | HD-Mel | 2.7 | 9 | 10 |
| ICE | 3.1 | 10 | 12 | |
| 6 | HD-Mel | 5.56 | 11 | 11 |
| 10 | ICE4-151 | 1.5 | 11 | 12 |
| ICE4-151 | 1.5 | 10 | 10 | |
| 12 | HD-Mel | 3.9 | 9 | 10 |
| 134-150 | HD-Mel | 3.15 | 10 | 10 |
| ICE | 5.19 | 10 | 11 |
| Patient No. . | Chemotherapy . | CD34 × 106 per kg Body Weight . | Days Until ANC >500/μL . | Days Until . |
|---|---|---|---|---|
| . | . | . | . | Platelet Count >20,000/μL . |
| 2 | HD-Mel | 2.3 | 10 | 11 |
| 3 | HD-Mel | 2.7 | 11 | 11 |
| 4 | HD-Mel | 2.04 | 12 | 11 |
| ICE | 4.8 | 9 | 11 | |
| 5 | HD-Mel | 2.7 | 9 | 10 |
| ICE | 3.1 | 10 | 12 | |
| 6 | HD-Mel | 5.56 | 11 | 11 |
| 10 | ICE4-151 | 1.5 | 11 | 12 |
| ICE4-151 | 1.5 | 10 | 10 | |
| 12 | HD-Mel | 3.9 | 9 | 10 |
| 134-150 | HD-Mel | 3.15 | 10 | 10 |
| ICE | 5.19 | 10 | 11 |
Abbreviation: HD, high-dose.
In addition to CD34-selected cells, unmanipulated PBPC were retransfused.
A modified ICE regimen was used: 10 g/m2 ifosfamide, 1,200 mg/m2 etoposide, and 1,200 mg/m2 carboplatin.
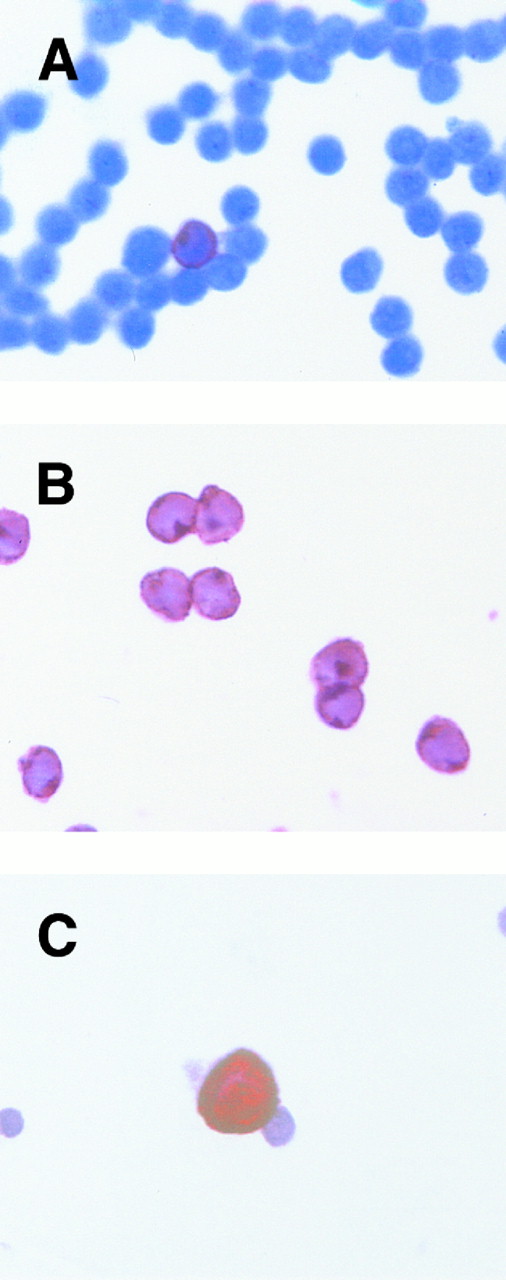
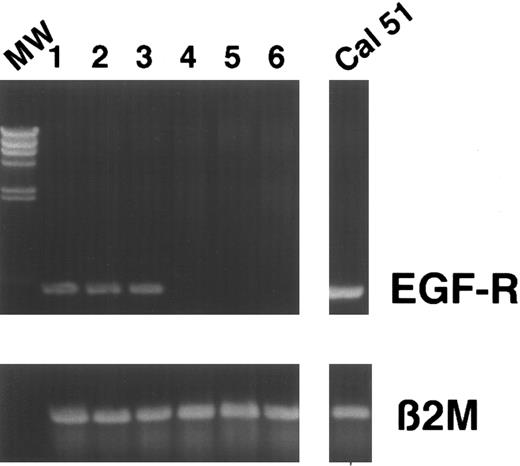
Detection of tumor cells in LP and after CD34 enrichment by RT-PCR and immunocytochemistry.Detection and quantitation of breast cancer cells within LP and after CD34 selection was performed using a standard immunocytochemical staining procedure as already described. Cells were stained with pancytokeratin monoclonal antibody A45. Using this method, the majority (17 of 21) of LP analyzed were found to contain tumor cells (Fig 2C). Despite high purities, we were still able to detect tumor cells within a substantial fraction of CD34-enriched populations (12 of 21), although tumor cell depletion could be observed ranging from 0.7 log to more than 3 log (median, 1.9). The data are summarized in Table 5. In addition, we compared these results with a RT-PCR–based detection assay. We recently established a RT-PCR method in an experimental small-scale protocol for monitoring tumor cell depletion during purging.19 We studied the expression of CK19 (as reported by Datta et al16 ) and the EGF-R as target genes. As described, RT-PCR was performed as a single amplification step for 40 cycles without reamplification using nested primers. In our hands, nested PCR for CK19 is not suitable for detection of residual tumor cells, due to false-positive results — it yields positive signals in 50% of normal buffy-coat donors despite DNAse digestion. This is probably due to the amplification of contaminating DNA and the CK19 pseudogenes, or to low-level mRNA expression in nonepithelial cells (data not shown). Using this experimental protocol, we were able to demonstrate a maximum sensitivity of one in 105 for CK19 and one in 106 for EGF-R, depending on the cell lines analyzed.19 Analysis of CK19 mRNA expression in LP or CD34-enriched cell populations showed only one positive patient sample. In contrast, results of RT-PCR analysis of EGF-R mRNA expression were comparable to the immunocytochemical detection of breast cancer cells, giving positive results in 10 of 17 LP and 6 of 17 CD34-enriched populations (Fig 3). Immunocytochemical and PCR analysis were conducted independently and matched at the end. A synopsis of results obtained by immunocytochemical staining and RT-PCR is depicted in Table 6.
Detection of breast cancer cells. CD34-selected cells were either stained with a CD3 MoAb (A), CD34 (B), or MoAb A45 anti-pancytokeratin (C). One CD3+ T cell can be detected in (A). All cells within the magnification field are CD34+ (B). Using pancytokeratin antibody A45, a breast cancer cell can be recognized in (C).
Detection of breast cancer cells. CD34-selected cells were either stained with a CD3 MoAb (A), CD34 (B), or MoAb A45 anti-pancytokeratin (C). One CD3+ T cell can be detected in (A). All cells within the magnification field are CD34+ (B). Using pancytokeratin antibody A45, a breast cancer cell can be recognized in (C).
Tumor Cell Contamination in PBPC Preparations
| Patient No. . | Cycle . | Preselection . | Postselection . | |
|---|---|---|---|---|
| . | . | Tumor Cells × 105 . | Tumor Cells × 105 . | Log Depletion Tumor Cells . |
| 1 | 0 | 0 | — | |
| 2 | 1.4 | 0 | >3 | |
| 3 | 1 | 8.2 | 0.18 | 1.7 |
| 2 | 20.8 | 0.10 | 2.3 | |
| 3 | 3.8 | 0.02 | 2.3 | |
| 4 | 1.3 | 0.03 | 1.6 | |
| 5 | 1 | 0.34 | 0 | >3 |
| 2 | 1.7 | 0.006 | 2.5 | |
| 6 | 1 | 1.56 | 0.023 | 1.8 |
| 2 | 6.2 | 0.47 | 1.1 | |
| 3 | 0.64 | 0.02 | 1.5 | |
| 7 | 2.3 | 0.05 | 1.7 | |
| 8 | 0.55 | 0.016 | 1.5 | |
| 9 | 2.5 | 0 | >3 log | |
| 10 | 0 | 0 | — | |
| 11 | 1 | 0 | 0 | — |
| 2 | 0.5 | 0.006 | 1.9 | |
| 12 | 1 | 0.42 | 0.083 | 0.7 |
| 2 | 3.4 | 0 | >3 | |
| 13 | 1 | 0 | 0 | — |
| 2 | 0.35 | 0 | >3 | |
| 3 | ND | ND | ||
| 4 | ND | ND | ||
| Patient No. . | Cycle . | Preselection . | Postselection . | |
|---|---|---|---|---|
| . | . | Tumor Cells × 105 . | Tumor Cells × 105 . | Log Depletion Tumor Cells . |
| 1 | 0 | 0 | — | |
| 2 | 1.4 | 0 | >3 | |
| 3 | 1 | 8.2 | 0.18 | 1.7 |
| 2 | 20.8 | 0.10 | 2.3 | |
| 3 | 3.8 | 0.02 | 2.3 | |
| 4 | 1.3 | 0.03 | 1.6 | |
| 5 | 1 | 0.34 | 0 | >3 |
| 2 | 1.7 | 0.006 | 2.5 | |
| 6 | 1 | 1.56 | 0.023 | 1.8 |
| 2 | 6.2 | 0.47 | 1.1 | |
| 3 | 0.64 | 0.02 | 1.5 | |
| 7 | 2.3 | 0.05 | 1.7 | |
| 8 | 0.55 | 0.016 | 1.5 | |
| 9 | 2.5 | 0 | >3 log | |
| 10 | 0 | 0 | — | |
| 11 | 1 | 0 | 0 | — |
| 2 | 0.5 | 0.006 | 1.9 | |
| 12 | 1 | 0.42 | 0.083 | 0.7 |
| 2 | 3.4 | 0 | >3 | |
| 13 | 1 | 0 | 0 | — |
| 2 | 0.35 | 0 | >3 | |
| 3 | ND | ND | ||
| 4 | ND | ND | ||
PCR before and after CD34 selection. Cells were analyzed for EGF-R expression before and after CD34 selection. Results from several patients are depicted. Lanes 1, 3, and 5, expression of EGF-R in LP. Lanes 2, 4, and 6, EGF-R expression in CD34+ fractions. Lanes 1 and 2, patient no. 6 first induction cycle. Lanes 3 and 4, patient no. 12 first analysis. Lanes 5 and 6, patient no. 12 second induction cycle. MW, molecular weight marker. Breast cancer cell line Cal 51 was used as positive control: β2-microglobulin served as a control.
PCR before and after CD34 selection. Cells were analyzed for EGF-R expression before and after CD34 selection. Results from several patients are depicted. Lanes 1, 3, and 5, expression of EGF-R in LP. Lanes 2, 4, and 6, EGF-R expression in CD34+ fractions. Lanes 1 and 2, patient no. 6 first induction cycle. Lanes 3 and 4, patient no. 12 first analysis. Lanes 5 and 6, patient no. 12 second induction cycle. MW, molecular weight marker. Breast cancer cell line Cal 51 was used as positive control: β2-microglobulin served as a control.
Detection of Tumor Cells by Immunocytochemistry and RT-PCR
| Patient No. . | Cycle . | Preselection . | Postselection . | ||||
|---|---|---|---|---|---|---|---|
| . | . | Immunocytochemistry . | CK19 RT-PCR . | EGF-R RT-PCR . | Immunocytochemistry . | CK19 RT-PCR . | EGF-R RT-PCR . |
| 1 | − | − | − | − | − | − | |
| 2 | + | − | + | − | − | − | |
| 3 | 1 | + | − | + | + | − | + |
| 2 | + | − | + | + | − | + | |
| 3 | + | ND | ND | + | ND | ND | |
| 4 | + | + | + | + | − | − | |
| 5 | 1 | + | ND | ND | − | ND | ND |
| 2 | + | − | + | + | − | + | |
| 6 | 1 | + | − | + | + | − | + |
| 2 | + | ND | ND | + | ND | ND | |
| 3 | + | ND | ND | + | ND | ND | |
| 9 | + | − | − | − | − | − | |
| 7 | + | − | − | + | − | − | |
| 8 | + | − | + | + | − | + | |
| 10 | − | − | − | − | − | − | |
| 11 | 1 | − | − | + | − | − | − |
| 2 | + | − | + | + | + | ||
| 12 | 1 | + | − | + | + | − | − |
| 2 | + | − | − | − | − | − | |
| 13 | 1 | + | − | − | − | − | − |
| 2 | − | − | − | − | − | − | |
| Patient No. . | Cycle . | Preselection . | Postselection . | ||||
|---|---|---|---|---|---|---|---|
| . | . | Immunocytochemistry . | CK19 RT-PCR . | EGF-R RT-PCR . | Immunocytochemistry . | CK19 RT-PCR . | EGF-R RT-PCR . |
| 1 | − | − | − | − | − | − | |
| 2 | + | − | + | − | − | − | |
| 3 | 1 | + | − | + | + | − | + |
| 2 | + | − | + | + | − | + | |
| 3 | + | ND | ND | + | ND | ND | |
| 4 | + | + | + | + | − | − | |
| 5 | 1 | + | ND | ND | − | ND | ND |
| 2 | + | − | + | + | − | + | |
| 6 | 1 | + | − | + | + | − | + |
| 2 | + | ND | ND | + | ND | ND | |
| 3 | + | ND | ND | + | ND | ND | |
| 9 | + | − | − | − | − | − | |
| 7 | + | − | − | + | − | − | |
| 8 | + | − | + | + | − | + | |
| 10 | − | − | − | − | − | − | |
| 11 | 1 | − | − | + | − | − | − |
| 2 | + | − | + | + | + | ||
| 12 | 1 | + | − | + | + | − | − |
| 2 | + | − | − | − | − | − | |
| 13 | 1 | + | − | − | − | − | − |
| 2 | − | − | − | − | − | − | |
Abbreviation: ND, not determined.
DISCUSSION
The aim of this study was twofold. First, we evaluated the tumor cell elimination capacity of CD34 enrichment using the immunomagnetic bead method; second, it was our aim to compare standard immunocytochemical tumor cell staining with RT-PCR in terms of the capacity for detecting residual cells within LP. Enrichment of CD34-expressing hematopoietic progenitor cells from peripheral blood or bone marrow transplantation has raised a lot of interest within the last few years. Enrichment of CD34+ cells offers a variety of novel approaches for combining transplantation with cellular therapeutic concepts, such as ex vivo expansion of hematopoietic progenitor cells,25 generation of professional antigen-presenting cells (eg, dendritic cells), gene therapy, and tumor cell purging. Different techniques have been developed for enrichment of CD34+ cells. In the meantime, a great deal of experience has been obtained with immunoaffinity selection of CD34 cells.1,2 We evaluated the immunomagnetic selection procedure with the intention to determine the capacity of this system to generate highly purified CD34 cell populations from LP of breast cancer patients, and to determine the tumor cell reduction efficiency. Using the immunomagnetic enrichment procedure, we were able to obtain a median purity of 83.3% (range, 45% to 95.4%) and a CD34 cell recovery of 73.2% (range, 22% to 95%). Phenotypic analysis of the CD34 subpopulations after enrichment showed that early CD34+/CD38−, CD34+/HLA-DR− hematopoietic stem cells are recovered in the selected fraction. Our experience with a total of 36 enrichment procedures in patients with different malignant diseases resulted in a median purity of 86.5% and a median recovery of 78.3% (submitted, data not shown). Autografting of these CD34+ cells after high-dose chemotherapy in breast cancer patients resulted in rapid and sustained engraftment. Neutrophil (ANC >500/μL) and platelet (>20,000/μL) recovery occurred at a median of day +10 and day +11, respectively. Thus, enrichment of CD34+ cells from LP using the immunomagnetic procedure results in high purities and excellent recovery rates. Transplantation of these cells leads to rapid and sustained engraftment. Our results concerning CD34 purity and yield compare favorably with other data published so far. Brugger et al26 reported a mean CD34 purity of 73% ± 24.6% and mean recovery rate of 61.4% ± 19.7% using an immunoaffinity column. Lane et al,21 using the immunomagnetic procedure, reported median purities ranging from 81% ± 11% to 77% ± 21% and recovery rates from 48% ± 12% to 51% ± 15% depending on the mobilization regimen.
Current purging strategies for tumor cell elimination include immunomagnetic tumor cell depletion,27,28 pharmacologic purging with 4-HC,29 immunotoxins,30,31 and immune-complement lysis.12 Determination of purging efficiency requires sensitive and reliable tumor cell detection methods. These methods are based on direct tumor cell staining (immunofluorescence flow cytometry and immunocytochemistry), tumor cell culture assays (determination of clonogenic growth capacity), or PCR analysis. The latter technique has been found to be of special relevance and extreme sensitivity in diseases with tumor- or even patient-specific genetic markers (eg, detection of t(14; 18) translocation in follicular lymphoma, t(9; 22) in CML, or CDR III–specific PCR in B-cell neoplasias). Due to the lack of such a tumor-specific marker in breast cancer, PCR-based tumor cell detection has been restricted to the analysis of tissue-specific transcipts. Various tissue-specific genes have been studied by RT-PCR for detection of minimal residual disease (cytokeratins 8, 18, and 19,16,32,33 breast mucin MUC-1,15 and EGF-R19 ). We evaluated the capacity of CK19- and EGF-R–based RT-PCR to monitor purging efficiency of CD34 selection in breast cancer patients in comparison to standard immunocytochemical tumor cell detection. Datta et al16 studied CK19 expression using a nested RT-PCR without DNAse digestion in 34 breast cancer patients. CK19-positive cells were only detectable in the peripheral blood or in marrow samples from stage IV patients. They were not able to detect CK19-positive cells within the LP of patients with stage IV disease. This is consistent with our data on CK19 RT-PCR. We found that CK19 RT-PCR was not sensitive using our PCR conditions (one of 17 LP analyzed). In contrast, using immunocytochemical staining, 17 of 21 (81%) LP were shown to contain cytokeratin-positive tumor cells before enrichment, compared with 10 of 17 (59%) LP using EGF-R–based RT-PCR. Using EGF-R–specific primers for RT-PCR and immunocytochemical CK19 staining, we were able to detect residual tumor cells after CD34 enrichment (12 of 21 immunocytochemically positive CD34 cell populations, and 6 of 17 using EGF-R RT-PCR). In comparison to immunocytochemical analysis of tumor cells, EGF-R–specific RT-PCR was less sensitive. This could be explained by the fact that EGF-R is expressed heterogenously in breast cancer cells.34 Moreover, mRNA expression levels of EGF-R might be low and degraded during RNA preparation. The high amount of tumor cells within our LP might be due to the fact that stem cell harvesting was performed after the first induction cycle. A similar observation has been made by Schulze et al,35 who detected tumor cells in four of six LP in high-risk patients (stage II/III >10 lymph nodes) after the first induction cycle. Another reason for the high frequency of tumor cells within the LP might be that the majority of our stage IV patients had a high tumor burden. Since tumor cells could still be detected after CD34 selection in breast cancer patients, it appears attractive to combine positive and negative purging methods for enhancing tumor cell depletion efficiency. This combined purging method might be of even greater relevance in patients with a high content of tumor cells within the autograft (eg, CLL or low-grade malignant lymphoma).
In conclusion, we have demonstrated that immunomagnetic CD34 selection results in high final purity and recovery rates. Transplantation of CD34-selected cells leads to rapid and sustained engraftment. This is accompanied by an indirect tumor cell depletion. In addition, we were able to show that tumor cells can be detected after CD34 selection, and that EGF-R–based RT-PCR appears to be a valuable novel method for detecting residual breast cancer cells within autografts.
ACKNOWLEDGMENT
We gratefully acknowledge the excellent technical assistance of Kati Hildebrandt, Kerstin Krüger, Simone Lusatis, Anke Sager, and the nurses of our transplantation unit.
Address reprint requests to Markus Y. Mapara, MD, Humboldt University, Virchow Klinikum-Robert Rössle Klinik, Department of Internal Medicine (Medical Oncology and Tumorimmunology), Lindenbergerweg 80, D-13122 Berlin, Germany.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal