Abstract
We report an RNA targeting strategy, which selectively degrades bcr/abl mRNA in chronic myelogenous leukemia (CML) cells. A 2′,5′-tetraadenylate activator (2-5A) of RNase L was chemically linked to oligonucleotide antisense directed against either the fusion site or against the translation start sequence in bcr/abl mRNA. Selective degradation of the targeted RNA sequences was demonstrated in assays with purified RNase L and decreases of p210bcr/abl kinase activity levels were obtained in the CML cell line, K562. Furthermore, the 2-5A-antisense chimeras suppressed growth of K562, while having substantially reduced effects on the promyelocytic leukemia cell line, HL60. Findings were extended to primary CML cells isolated from bone marrow of patients. The 2-5A-antisense treatments both suppressed proliferation of the leukemia cells and selectively depleted levels of bcr/abl mRNA without affecting levels of β-actin mRNA, determined by reverse transcriptase-polymerase chain reaction (RT-PCR). The specificity of this approach was further shown with control oligonucleotides, such as chimeras containing an inactive dimeric form of 2-5A, antisense lacking 2-5A, or chimeras with altered sequences including several mismatched nucleotides. The control oligonucleotides had either reduced or no effect on CML cell growth and bcr/abl mRNA levels. These findings show that CML cell growth can be selectively suppressed by targeting bcr/abl mRNA with 2-5A-antisense for decay by RNase L and suggest that these compounds should be further explored for their potential as ex vivo purging agents of autologous hematopoietic stem cell transplants from CML patients.
ANTISENSE OLIGONUCLEOTIDES (ODNs) have the potential to function as highly selective therapeutic agents by virtue of their ability to bind with unique nucleotide sequences in mRNAs for disease-causing proteins, including those implicated in cancer. Indeed, several antisense ODNs, which are designed to prevent the synthesis of different proteins necessary for cancer cell growth and survival have been used in clinical trials.1 Chronic myelogenous leukemia (CML), a pluripotent hematopoietic stem cell disease, has been a frequent subject of antisense-based therapeutic approaches.1-4 CML cells contain the Philadelphia chromosome (Ph1) generated from reciprocal translocation of chromosomes 9q and 22q, in more than 90% of patients. The bcr/abl fusion genes (b2/a2 and b3/a2) in Ph1 encode tyrosine kinases implicated in the development of CML.5-7 The presence of unique nucleotide sequences at the fusion site of bcr/abl and the requirement of the encoded tyrosine kinase for cell survival make CML an attractive target for antisense strategies. In this regard, antisense ODNs directed at the breakpoint junction of bcr/abl mRNA or at the translation start site of bcr mRNA have been shown to selectively inhibit proliferation or survival of bcr/abl-expressing cell lines and primary cells.8,9 Furthermore, antisense ODN treatments in vivo suppressed development of CML-like disease in murine models.10-13 These encouraging results have led to clinical trials with antisense ODNs directed to bcr/abl mRNA for ex vivo treatment of bone marrow.14
Despite progress, antisense approaches for CML remain controversial because while hybridization-dependent, antisense mechanisms against bcr/abl mRNA have been reported,8,15 others observed that some ODNs clearly caused nonantisense or aptameric effects.16,17 For instance, phosphorothioate-capped ODNs had sequence-specific inhibitory effects against the kinase activity of p210bcr/abl.16 Moreover, a significant obstacle to the further development of antisense ODN therapeutics is that even when an antisense effect is established, the molecular mechanism is often unknown or speculative.
We have exploited the controllable endoribonuclease, RNase L, present in a wide range of mammalian cell types, for the purpose of enhancing the activity of antisense ODNs.18 RNase L is an endoribonuclease for single-stranded RNA that functions in the interferon-regulated, 2-5A system.19 A unique feature of RNase L is that it is converted from a silent to an active form in response to short 2′,5′-oligoadenylates, or 2-5A. To activate RNase L and direct it to the RNA target, a 2′,5′-tetraadenylate is attached through linkers to the 5′-terminus of an antisense ODN.18The resulting compounds, referred to as 2-5A-antisense, are a class of chimeric ODNs designed to activate the ubiquitous, intracellular RNase L in the proximity of targeted RNA molecules. A 5′-phosphorylated, 2′,5′-linked oligoadenylate (2-5A) of at least three adenylyl residues in length is required to activate RNase L.20 In the 2-5A–antisense approach, RNase L is converted to a highly specific endoribonuclease capable of selectively cleaving individual RNA targets. This was previously shown in cell-free systems and in human cells.18,21-23 For example, when applied to human HeLa cells, 2-5A–antisense resulted in the selective degradation of targeted mRNA for the protein kinase PKR.21 Recently, we showed that the recruitment and activation of RNase L to a respiratory syncytial virus mRNA with 2-5A–antisense resulted in a potent inhibition of virus replication in human tracheal epithelial cells.23 Here we have further developed 2-5A–antisense technology for the purpose of selectively degrading bcr/abl mRNA thereby blocking proliferation of leukemic cells. We show that 2-5A–antisense chimeras are effective in causing the decay of bcr/abl mRNA leading to the suppression of the growth of Ph1positive K562 cells and of primary cells from CML patients. This work could lead to an improved method of ex vivo purging Ph1cells from bone marrow or peripheral blood stem cell preparations intended for autologous transplantation.
MATERIAL AND METHODS
Cell culture and ODN treatments of cells.
The CML cell line, K562, containing a bcr/abl gene with a b3/a2 fusion site (bcr exon 3 to c-abl exon 2) and the promyelocytic leukemia cell line, HL60, were maintained in RPMI 1640 medium (GIBCO-BRL, Gaithersburg, MD) supplemented with 10% heat-inactivated fetal bovine serum (FBS) in a moist atmosphere with 5% CO2at 37°C. Primary cells were isolated from bone marrow of three CML patients in chronic phase by Ficoll-Hypaque density gradient sedimentation. Analysis by reverse transcriptase-coupled polymerase chain reaction (RT-PCR) showed a b3/a2 type of bcr/abl mRNA in bone marrow from all three patients. In addition, normal bone marrow was collected from two healthy individuals. Patient cells (1 × 105) were placed in 1 mL liquid suspension culture, Isocove’s modified Dulbecco’s modified medium (GIBCO-BRL) supplemented with 2.5% human AB serum, penicillin, streptomycin, 20 U/mL of interleukin-3 (Sigma, St Louis, MO), 5 ng per mL of granulocyte-macrophage colony-stimulating factor (GM-CSF) (Sigma) and 50 ng/mL of stem cell factor (SCF) (Pharmingen, San Diego, CA).24 Experiments were performed in duplicate or triplicate. ODNs were added every 12 hours to the cells (about 1 × 105) in 1 mL of media/serum. The viable cell numbers were determined by trypan blue dye exclusion assay in a hemocytometer.
Chemical synthesis of 2-5A–antisense chimeras and control ODNs.
Chimeric phosphodiester ODNs for this study were synthesized and purified using previously published procedures.25-29 The generic structure of 2-5A–antisense is; Sp5′A2′p(5′A2′p)3-[O(CH2)4Op]2-5′dN3′p(5′dN3′p)m5′dN3′p-3′pdN5′ (for specific structures see Table 1). All of the 2-5A–antisense chimeras described herein were characterized by high performance liquid chromatography (HPLC) and digestion with snake venom phosphodiesterase.27,28Antisense ODNs were complementary to either the b3/a2 or the b2/a2 types of bcr/abl fusion sites or to a sequence beginning at the translation start codon of bcr/abl mRNA (Table 1). The 5′-thiophosphorylated (2′,5′)di- or tetraadenylate was attached to the antisense ODNs through two tandem butanediol phosphate linkers. The 3′-terminal nucleotides in the antisense sequences were linked through a 3′-3′ phosphodiester bond to enhance their stability to 3′ to 5′ exonuclease activity.28 The 5′-thiophosphate moiety was present to enhance resistance to phosphatases.26 A nuclease resistant 2-5A-iso-antisense, SpA4-iso-anti-bcr, containing a reverse polarity antisense moiety was synthesized as described.29
Assay for targeted cleavage of RNA by purified RNase L.
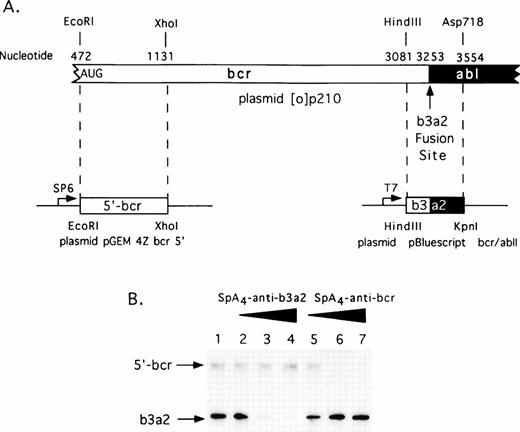
A complete coding sequence cDNA for the p210bcr/abl in vector pGEM 4Z (plasmid [0]p210) was a generous gift from Dr Owen Witte (UCLA, Los Angeles, CA). Restriction fragments from this plasmid were subcloned to produce two derivative plasmids encoding the 5′-bcr and bcr/abl RNA segments (Fig 1A). The former plasmid, pGEM 4Z bcr 5′ was constructed by digesting plasmid [0]p210 first withEcoRI followed by a double digestion with Sal I and Xho I. An EcoRI/Xho I fragment encoding the 5′bcr segment was subcloned into pGEM 4Z (predigestedEcoRI and Sal I). To produce 5′bcr RNA, plasmid pGEM 4Z bcr 5′ was linearized with Xho I and then transcribed with SP6 RNA polymerase. The second plasmid was constructed by digesting [0]p210 with HindIII and Asp718 releasing a fragment encoding the bcr/abl fusion segment. TheHindIII/Asp718 fragment was subcloned into the plasmid pBluescript II KS+ (predigested with HindIII andKpn I) to produce plasmid pBluescript bcr/ablI. To produce the bcr/abl RNA segment, plasmid pBluescript bcr/ablI was digested with Asp718 and transcribed with T7 RNA polymerase.
The RNA segments were dephosphorylated with alkaline phosphatase (Boehringer, Indianapolis, IN), then incubated with proteinase K and phenol extracted before labeling at the 5′-termini with 2 U of T4 polynucleotide kinase (U.S.B., Arlington Heights, IL) and 50 μCi of [γ-32P] adenosine triphosphate (ATP) (3,000 Ci/mmol). The RNAs were purified from 6% polyacrylamide/8 mol/L urea gels for use in the cleavage reactions. Reactions were performed in the absence or presence of ODNs (50, 100, or 200 nmol/L) with 50 nmol/L each of 5′-radiolabeled 5′-bcr RNA and 5′-radiolabeled bcr/abl RNA segment in buffer containing 25 mmol/L Tris-HCl pH 7.4, 10 mmol/L magnesium acetate, 8 mmol/L β-mercaptoethanol, and 100 mmol/L KCl. RNase L (20 ng; 12 nmol/L final concentration) was added to a final volume of 20 μL and incubations were at 37°C for 30 minutes. This RNase L preparation was made in SF21 insect cells from a human cDNA subcloned in a baculovirus vector (Clontech, Palo Alto, CA) and was purified.30 31 Reactions were terminated with 10 μL of formamide stop buffer. RNA was analyzed in 6% polyacrylamide/8 mol/L urea gels (30 × 40 × 0.04 cm). Degradation of32P-labeled RNA was monitored by analysis in a PhosphorImager (Molecular Dynamics, Sunnyvale, CA).
p210bcr/abl kinase assays for oligonucleotide-treated cells.
ODN-treated K562 cells were harvested and washed twice with ice-cold phosphate-buffered saline (PBS). Cell lysis, immunoprecipitation, and assays of p210bcr/abl kinase activity were performed as described.5 Briefly, protein samples (100 μg) of 100,000g centrifugations of cell lysates were incubated with preimmune sera or anti-abl antisera (a kind gift of O. Witte) at 4°C with gentle shaking for 16 hours. Protein A-sepharose beads (25 μL packed volume) (Pharmacia, Piscataway, NJ) were added to the protein samples and incubated for 2 hours at 4°C. The immune complexes were harvested by centrifugation at 5,000g for 10 minutes at 4°C, washed, and phosphorylation was performed in the presence of 10 μCi of [γ-32P]-ATP (NEN, Boston, MA) (5,000 Ci/mmol). Reactions were incubated at 30°C for 30 minutes. Proteins were eluted by boiling in sodium dodecyl sulfate (SDS)/gel sample buffer for 5 minutes and subjected to electrophoresis on SDS/8% polyacrylamide gels. Gels were treated with 1 mol/L potassium hydroxide (KOH) for 2 hours at 55°C, fixed, dried, and analyzed by autoradiography. The positions of the bands were compared with molecular weight standards. The relative radioactivity in the bands were determined using a PhosphorImager (Molecular Dynamics).
RT-PCR.
Cells were harvested from 1-mL cultures and the total RNA was extracted from untreated and ODN-treated cell lines or primary cells using RNazol (Tel-Test, Friendswood, TX). RNA (3 μg) was treated with RNase free DNase, extracted with phenol chloroform, and precipitated with ethanol. The mRNA was used as template for cDNA synthesis by reverse transcriptase using oligo (dT) primer. The cDNA was then diluted serially with PCR buffer and cDNA at each dilution was amplified with the use of primers to sense sequences in exon 3 of bcr (5′-GTCTCCGGGGCTCTATGGGT-3′) and antisense sequences in exon 2 of c-abl (5′-CACAGGCCCATGGTACCAGG-3′) or to the β-actin coding sense sequence, 5′-GCTGTGCTATCCCTGTACG-3′ and antisense 5′TGCCTCAGGGCAGCGGAA-3′, respectively. Denaturation was at 94°C for 1 minute, annealing was at 48°C for 1 minute, and elongation was at 72°C for 1.5 minutes. PCR products in primary cells were obtained after 30 cycles for bcr/abl and β-actin cDNA from patient 1 and after 40 and 30 cycles, respectively, for bcr/abl and β-actin cDNA from patient 2. The size of the DNA products for bcr/abl and β-actin PCR products are 385 and 368 bp, respectively. The PCR products were analyzed by agarose gel (1.2%) followed by ethidium bromide staining.
RESULTS
Selection of target sites in bcr/abl mRNA and design of ODNs.
Two potential binding sites for ODNs in the b3/a2 form of bcr/abl were selected for this study. The b3/a2 fusion site was chosen because it offered a means of selectively targeting bcr/abl mRNA present only in the malignant cells. The fusion site was in the center of these ODNs, such that there would be 18 continuous complementary nucleotides to bcr/abl mRNA. Therefore, bcr mRNA and c-abl mRNA contain only 9 nucleotides each of sequence complementary to the ODN. 2-5A–antisense against the translational start site sequence was also synthesized because of prior success in targeting this region of bcr/abl mRNA with antisense.8,32 Both ends of the chimeric ODNs were chemically modified to enhance their stability against enzymatic degradation. A 5′-thiophosphate on the 2-5A moiety provides resistance to phosphatase activity without impairing activation of RNase L.26 At the opposite termini in the DNA or antisense sequence, the last internucleotide linkage was inverted from 5′,3′ to 3′,3′; a modification that substantially enhanced the stability of the ODNs in human serum.28 In addition, a nuclease-resistant ODN, 2-5A-iso-anti-bcr, was synthesized in which the antisense portion was 3′ to 5′ instead of 5′ to 3′.29 A panel of 2-5A–antisense and control ODNs was synthesized and purified (Materials and Methods and Table 1) (the two test ODNs,SpA4-anti-b3a2 orSpA4-anti-bcr, are indicated in text in bold print).
Selective cleavage of the target RNA sequence in vitro.
To determine if the 2-5A–antisense molecules could direct RNase L to cleave the translation start site or fusion site of bcr/abl mRNA, two regions of the bcr/abl mRNA containing these sequences were synthesized in vitro (Fig 1A). One RNA segment of 659 bases, referred to as 5′ bcr RNA, is the translational start region of bcr/abl mRNA. The other RNA segment of 473 bases, is the b3/a2 bcr/abl fusion region. Incubations with equal amounts of both RNA segments, RNase L and increasing amounts of eitherSpA4-anti-b3a2 orSpA4-anti-bcr resulted in specific degradation of the targeted, but not of the nontargeted RNA (Fig 1B). ODN concentrations of 100 nmol/L produced greater than 90% reductions in the targeted RNA molecules. In contrast, there was only minimal degradation (≤30%) of the nontargeted RNA molecules present in the same reactions. In addition, 2-5A-iso-anti-bcr, containing reverse polarity antisense, was able to effect the selective in vitro cleavage of 5′-bcr RNA by RNase L.29 These results show that these 2-5A–antisense chimeras activate RNase L and produce targeted degradation of RNA in vitro.
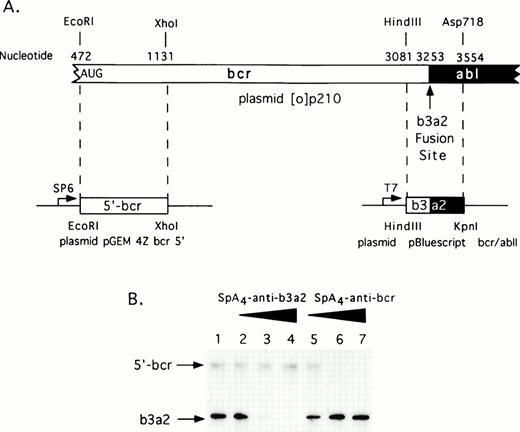
Selective targeting of sequences in bcr/abl mRNA for in vitro degradation by RNase L activated with 2-5A–antisense. (A) Construction of plasmids for in vitro synthesis of bcr/abl and 5′-bcr RNA segments (Materials and Methods). (B) The positions of the 5′-bcr and b3/a2 fusion site RNAs are indicated in the autoradiogram of the dried, SDS/polyacrylamide gel (arrows). Lane 1, absense of ODN; lanes 2 to 4: 50, 100, and 200 nmol/L of SpA4-anti-b3a2; lanes 5 to 7: 50, 100, and 200 nmol/L of SpA4-antibcr. The b3a2 RNA was labeled to a higher specific activity than the 5′ bcr segment, and thus appears darker in the autoradiogram, although equivalent amounts were included.
Selective targeting of sequences in bcr/abl mRNA for in vitro degradation by RNase L activated with 2-5A–antisense. (A) Construction of plasmids for in vitro synthesis of bcr/abl and 5′-bcr RNA segments (Materials and Methods). (B) The positions of the 5′-bcr and b3/a2 fusion site RNAs are indicated in the autoradiogram of the dried, SDS/polyacrylamide gel (arrows). Lane 1, absense of ODN; lanes 2 to 4: 50, 100, and 200 nmol/L of SpA4-anti-b3a2; lanes 5 to 7: 50, 100, and 200 nmol/L of SpA4-antibcr. The b3a2 RNA was labeled to a higher specific activity than the 5′ bcr segment, and thus appears darker in the autoradiogram, although equivalent amounts were included.
Effect of oligonucleotides on p210bcr/abl kinase activity levels.
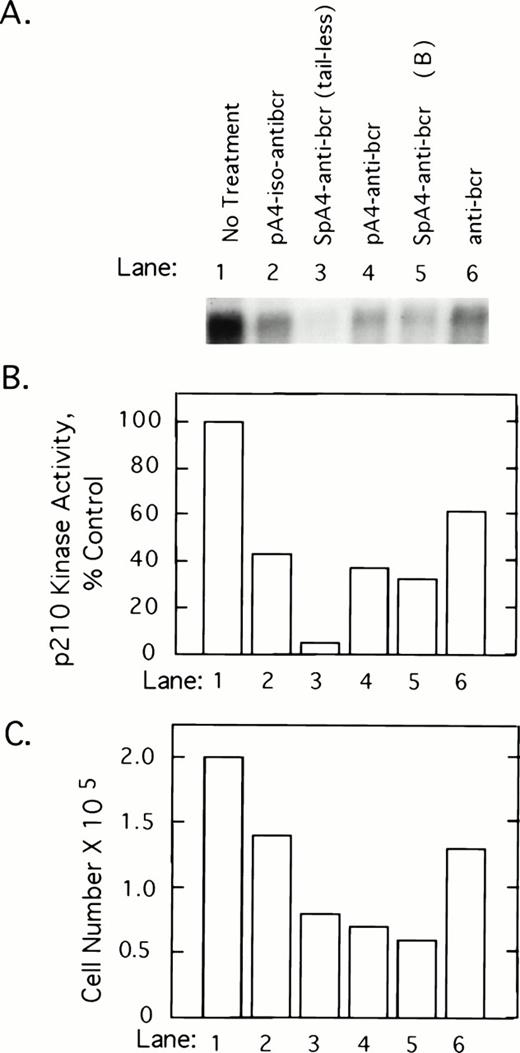
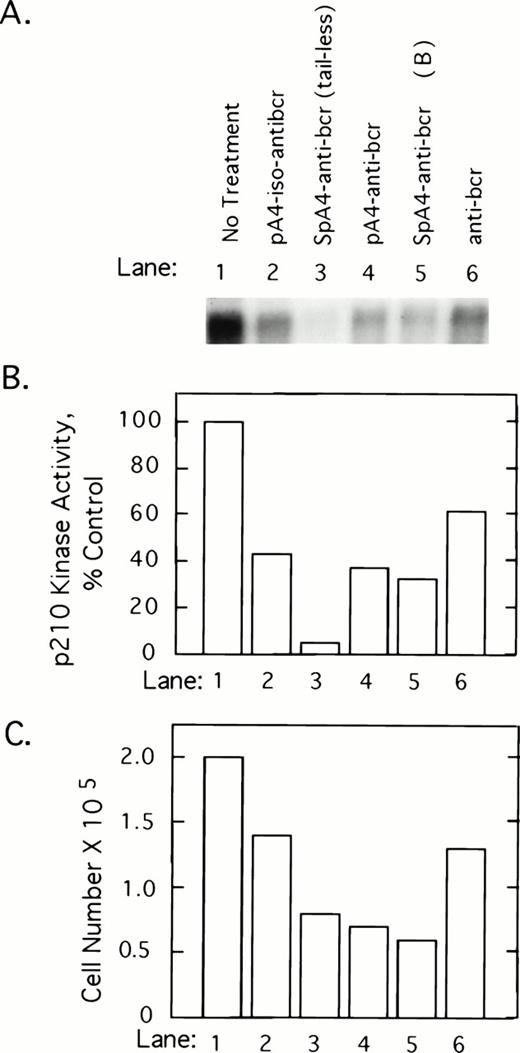
To determine whether 2-5A–antisense against the 5′-end of the bcr/abl mRNA coding sequence has an impact on p210bcr/abl, K562 cells were treated with 5 μmol/L of test and control oligonucleotides at 0, 12, 24, and 36 hours and harvested at 40 hours after the first treatment. p210bcr/abl kinase present in cell lysates was immunoprecipitated and assayed by autophosphorylation with γ-32P-labeled ATP. Results show that p210bcr/abl kinase activity levels declined in response to the different ODNs in the following order: SpA4-anti-bcr (tail-less) > SpaA4-anti-bcr (B)>pA4-anti-bcr > pA4-iso-anti-bcr > anti-bcr (Fig 2A and B). Enhanced inhibition was observed when 2-5A moieties were attached to the antisense ODNs, suggesting that the effect observed is due to RNase L-mediated degradation of the bcr/abl mRNA. The three ODNs, which produced the greatest inhibition of p210bcr/abl kinase, also were the most potent inhibitors of K562 cell growth (Fig 2C). However, SpA4-anti-bcr (tail-less) produced the largest decline in p210bcr/abl kinase activity whereas SpA4-anti-bcr (B) showed slightly greater anticellular activity (lanes 3 and 5).
2-5A–anti-bcr treatment of K562 cells leads to decreased levels of p210bcr-abl kinase activity and cell growth inhibition. (A) An autoradiogram and (B) results of phosphorImage analysis of the gel from a p210bcr/abl kinase activity assay measured from K562 cells treated with different ODNs as indicated in the figure and described in Materials and Methods. (C) Effects on K562 cell growth and survival of ODN treatments. Cells were seeded at 105 cells per well at time 0 and counted after 40 hours of ODN treatment (y-axis).
2-5A–anti-bcr treatment of K562 cells leads to decreased levels of p210bcr-abl kinase activity and cell growth inhibition. (A) An autoradiogram and (B) results of phosphorImage analysis of the gel from a p210bcr/abl kinase activity assay measured from K562 cells treated with different ODNs as indicated in the figure and described in Materials and Methods. (C) Effects on K562 cell growth and survival of ODN treatments. Cells were seeded at 105 cells per well at time 0 and counted after 40 hours of ODN treatment (y-axis).
Growth suppression of a CML cell line, K562, after treatment with 2-5A–antisense.
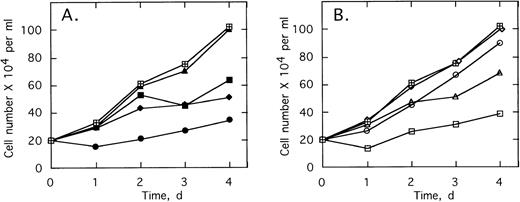
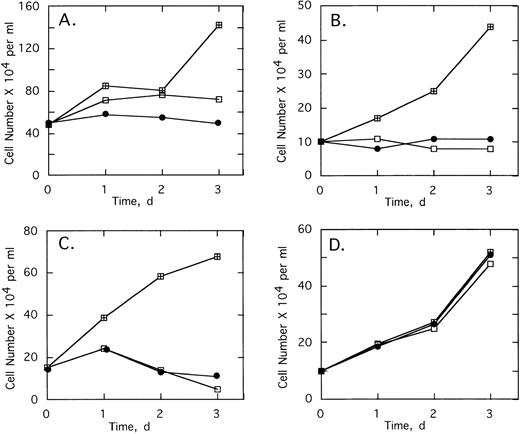
ODNs at concentrations of 2 μmol/L per treatment were added twice daily to cultures of K562 cells, a CML cell line containing the b3/a2 type of the bcr/abl oncogene. Addition of 2-5A–antisense against the translation start sequence,SpA4-anti-bcr,substantially suppressed cell proliferation (Fig3A). The cell doubling time increased from 1.7 to >4 days in the presence ofSpA4-anti-bcr. In contrast, the control ODN containing a disabled, dimeric, 2-5A moiety, SpA2-anti-bcr, had no effect on K562 cell growth (Fig 3A). Antisense lacking a 2-5A moiety, anti-bcr, as well as ODN with five mismatches, SpA4-(M5)anti-bcr, had intermediate effects on cell growth, increasing the doubling time to 3.0 and 2.4 days, respectively. 2-5A–antisense to the b3a2 fusion site in bcr/abl mRNA,SpA4-anti-b3a2, suppressed K562 cell proliferation, increasing the doubling time to 4.2 days (Fig3B). In contrast, SpA4-(M6)anti-b3a2, with six mismatches in the antisense sequence, had no effect on the cell growth rate. A 2-5A–antisense to a b2/a2 fusion site, SpA4-anti-b2a2, increased the doubling time slightly to 1.85 days, while anti-b3a2, lacking 2-5A, increased the doubling time to 2.2 days. These findings demonstrate the ability of 2-5A–antisense to suppress proliferation of CML cells in a sequence-specific manner. In addition, the enhanced anticellular activity of 2-5A–antisense chimeras, compared with ODNs lacking 2-5A or containing inactive dimeric 2-5A, provided evidence that recruitment of RNase L enhances the antisense effect.
Growth of the CML cell line, K562, is suppressed by 2-5–antisense directed against bcr-abl mRNA. Cells were without treatment (⊞) or with twice daily treatments with 2 μmol/L of (A) SpA4-anti-bcr (•), anti-bcr (⧫), SpA4-(M5)anti-bcr (▪), or SpA2-anti-bcr (▴) or in (B) of SpA4-anti-b3a2 (□), anti-b3a2 (▵), SpA4-anti-b2a2 (○), or SpA4-(M6)anti-b3a2 (◊). Cell numbers are the average obtained from duplicate wells.
Growth of the CML cell line, K562, is suppressed by 2-5–antisense directed against bcr-abl mRNA. Cells were without treatment (⊞) or with twice daily treatments with 2 μmol/L of (A) SpA4-anti-bcr (•), anti-bcr (⧫), SpA4-(M5)anti-bcr (▪), or SpA2-anti-bcr (▴) or in (B) of SpA4-anti-b3a2 (□), anti-b3a2 (▵), SpA4-anti-b2a2 (○), or SpA4-(M6)anti-b3a2 (◊). Cell numbers are the average obtained from duplicate wells.
2-5A–antisense ODNs selectively inhibit proliferation of CML cells.
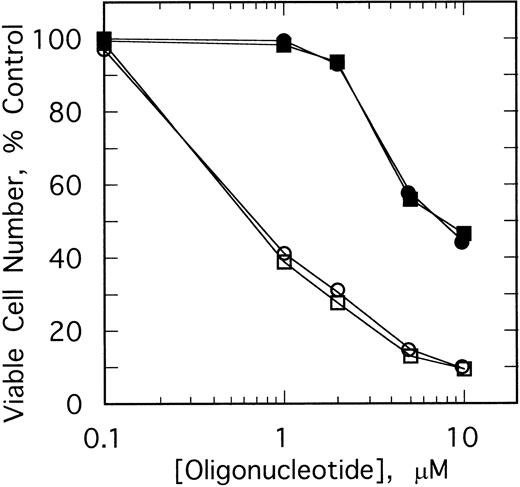
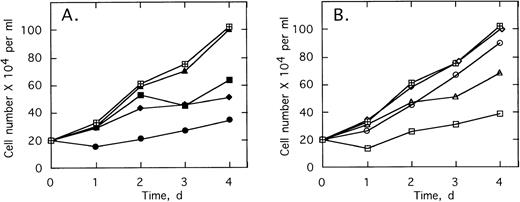
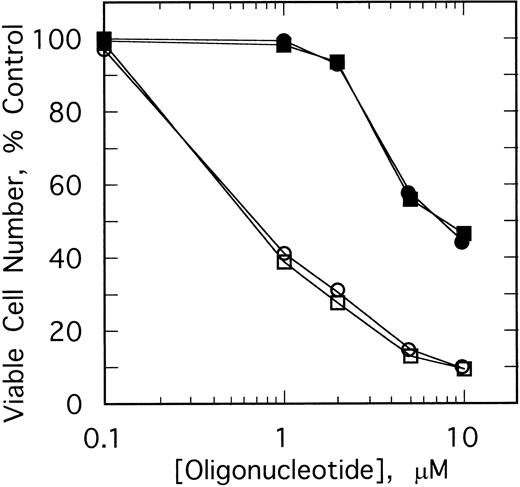
The specificity of this approach was investigated by comparing the effects of different doses of the ODNs on growth of K562 cells and HL60 cells, a promyelocytic cell line. After twice daily treatments of the K562 cells for 4 days with 1 μmol/L ofSpA4-anti-b3a2 orSpA4-anti-bcr, cell numbers were 41% to 44% those of the untreated cultures, while 10 μmol/L treatments of the K562 cells reduced the cell numbers to about 10% of the control culture (Fig 4). In contrast, 1 μmol/L ODN treatments had no effect on the HL60 cells. A small (10% to 11%) inhibition of HL60 cell growth was observed with 2 μmol/L ODN treatments, and 10 μmol/L treatments reduced the cell numbers to about 45% those of the control culture (Fig 4). The concentrations ofSpA4-anti-b3a2 orSpA4-anti-bcr required to inhibit growth by 50% were 0.7 μmol/L and 7.5 μmol/L for the K562 cells and for the HL60 cells, respectively. Therefore, there was a considerable (10-fold) specificity in inhibitory activity of the 2-5A–antisense ODNs for the K562 cells compared with the HL60 cells.
Effect of different doses of SpA4-anti-b3a2 (□,▪), SpA4-anti-bcr (○,•) on the growth of K562 cells (open symbols) and HL60 cells (closed symbols). Data points are the ratio of the viable cell numbers for the treated and untreated cultures × 100. ODNs at concentrations of 5 μmol/L were added twice daily.
Effect of different doses of SpA4-anti-b3a2 (□,▪), SpA4-anti-bcr (○,•) on the growth of K562 cells (open symbols) and HL60 cells (closed symbols). Data points are the ratio of the viable cell numbers for the treated and untreated cultures × 100. ODNs at concentrations of 5 μmol/L were added twice daily.
2-5A–antisense blocks proliferation and survival of primary CML cells.
To determine the effect of 2-5A–antisense on the growth and survival of primary leukemia cells, bone marrow aspirates from three CML patients were analyzed. Chromosome analysis was performed on 20 cells from each of the three patients’ bone marrow. In cells from patients 1 and 3, Ph1 was present in all 20 cells analyzed, whereas in patient 2, Ph1 was observed in 17 of 20 of the bone marrow cells, which were analyzed (not shown). In addition, bone marrow cells from all three patients contained the b3/a2 form of bcr/abl as determined by RT-PCR (see below). Bone marrow cells of the three patients were cultured and then treated with the ODNs. In every case, treatments of 5 μmol/L ofSpA4-anti-b3a2 orSpA4-anti-bcr caused cell proliferation to cease after 24 hours (Fig 5A through C). In contrast, bone marrow cells from two healthy individuals were resistant to the growth-suppressing effects of the ODNs (Fig 5D and data not shown).
Effects of twice daily treatments with 5 μmol/L of SpA4-anti-bcr (•) or SpA4-anti-b3a2 (□) on the growth and survival of CML cells derived from (A) patient 1, (B) patient 2, and (C) patient 3, and (D) of bone marrow from a healthy individual. No treatment: (⊞). Cell numbers are the average obtained from duplicate wells.
Effects of twice daily treatments with 5 μmol/L of SpA4-anti-bcr (•) or SpA4-anti-b3a2 (□) on the growth and survival of CML cells derived from (A) patient 1, (B) patient 2, and (C) patient 3, and (D) of bone marrow from a healthy individual. No treatment: (⊞). Cell numbers are the average obtained from duplicate wells.
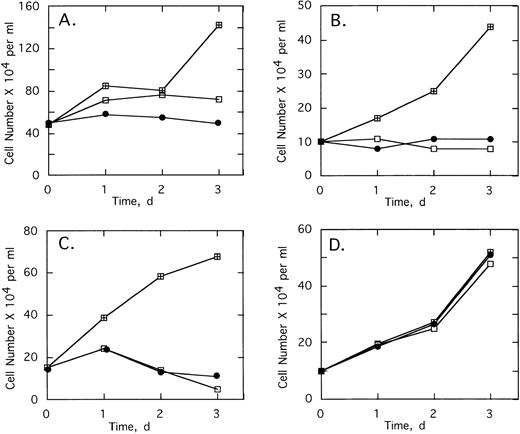
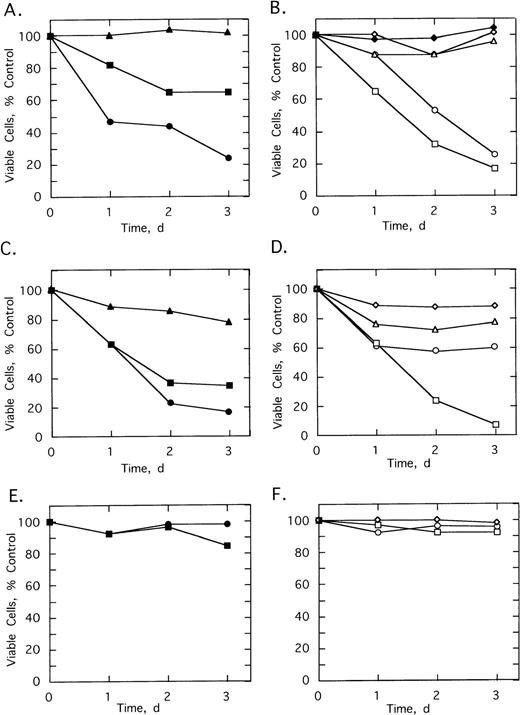
Various control ODNs were used to show both an antisense effect and involvement of RNase L in the growth suppression. Cell growth and survival was measured using cells from patients 2 and 3 cultured in the absence or presence of 5 μmol/L of ODNs added twice daily for 3 days (Fig 6). A control ODN, SpA2-anti-bcr, containing a shortened 2-5A moiety incapable of activating RNase L, had no effect on the cells from patient 2 and reduced the number of viable cells from patient 3 by only 21% (Fig 6A and C). In contrast,SpA4-anti-bcr reduced by 76% and 83% the numbers of viable cells from patients 2 and 3, respectively (Fig 6A and C). Surprisingly, the 2-5A–antisense containing five mismatches, SpA4-(M5)anti-bcr, produced a substantial, 35% and 65%, reduction in cell viability (Fig 6A and C). In contrast, the other control mismatched ODN, SpA4-(M6)anti-b3a2, had no effect on the cell count from patient 2 and produced only a 12% decrease in cell number from patient 3 (Fig 6D and B). The ODN, anti-b3a2, lacking a 2-5A moiety, reduced the cell number by ≤28% (Fig 6B and D). 2-5A–antisense directed against a b2/a2 type of bcr/abl fusion site, SpA4-anti-b2a2, produced a 74% and a 39% relative decrease in cell numbers from patients 2 and 3, respectively (Fig 6B and D). Finally,SpA4-anti-b3a2 caused a potent, 83% and 92%, anticellular effect on cells from patients 2 and 3, respectively (Fig 6B and D). In contrast, there was little or no effect of the ODNs on the survival or growth of the cells from the healthy, control individuals (Fig 6E and F and data not shown).
Effects of twice daily treatments on growth and survival of CML cells derived from (A, B) patient 2, (C, D) patient 3 (E, F) healthy control individual of SpA4-anti-bcr (•), SpA4-(M5)anti-bcr (▪), SpA2-anti-bcr (▴), SpA4-anti-b3a2 (□), SpA4-anti-b2a2 (○), anti-b3a2 (▵), SpA4-(M6)anti-b3a2 (◊) or anti-bcr (⧫). ODNs were added twice daily at a concentration of 5 μmol/L. Data points are the ratio of the viable cell numbers for the treated and untreated cultures × 100. Experiments were performed in duplicate and results averaged.
Effects of twice daily treatments on growth and survival of CML cells derived from (A, B) patient 2, (C, D) patient 3 (E, F) healthy control individual of SpA4-anti-bcr (•), SpA4-(M5)anti-bcr (▪), SpA2-anti-bcr (▴), SpA4-anti-b3a2 (□), SpA4-anti-b2a2 (○), anti-b3a2 (▵), SpA4-(M6)anti-b3a2 (◊) or anti-bcr (⧫). ODNs were added twice daily at a concentration of 5 μmol/L. Data points are the ratio of the viable cell numbers for the treated and untreated cultures × 100. Experiments were performed in duplicate and results averaged.
2-5A–antisense caused a selective decrease in the levels of the bcr/abl mRNA in the primary CML cells.
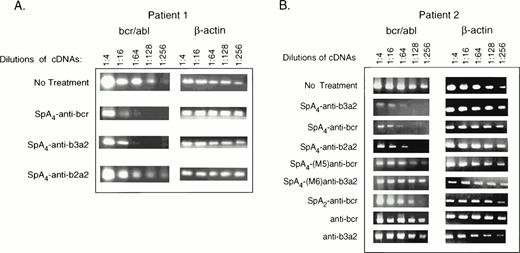
RT-PCR analysis was performed to determine if there was a specific effect of the 2-5A–antisense ODNs on amounts of the bcr/abl mRNA. Primary CML cells from patients 1 and 2 were treated twice daily for 2.5 days with ODNs before isolation of RNA (Materials and Methods). As a reference, relative levels of β-actin mRNA were also determined. After the reverse transcriptase reactions, the cDNA was serially diluted and used for amplification (Fig 7). Amounts of β-actin mRNA were not significantly affected by the ODN treatments. In contrast,SpA4-anti-bcr resulted in ≥16-fold decreases in levels of the bcr/abl mRNA in cells from patients 1 and 2, as estimated by comparison to the untreated controls (Fig 7A and B). Treatment withSpA4-anti-b3a2 also reduced levels of bcr/abl mRNA by ≥16-fold (Fig 7A and B). SpA4-anti-b2a2, directed against a b2a2 fusion sequence, was less active on the cells from patient 1, reducing levels of bcr/abl mRNA by fourfold, while reducing bcr/abl mRNA levels from patient 2 by ≥16-fold. The mismatched 2-5A–antisense ODN, SpA4-(M6)anti-b3a2 had no effect while SpA4-(M5)anti-bcr reduced bcr/abl mRNA levels by about fourfold. SpA2-anti-bcr reduced bcr/abl mRNA amounts by twofold, while anti-bcr and anti-b3a2 had no effect. Of the ODNs tested, SpA4-anti-bcr andSpA4-anti-b3a2 had the greatest effects on reducing both bcr/abl mRNA levels and CML cell numbers.
Targeted degradation of bcr-abl mRNA in CML cells as determined by RT-PCR. RT-PCR products from bcr-abl and β-actin mRNAs were separated on agarose gels and stained with ethidium bromide. ODNs (as indicated in the figure) were added twice daily to CML cells from (A) patient 1 (5 μmol/L per treatment) or (B) patient 2 (2 μmol/L per treatment) for 2.5 days before isolation of RNA. The images show the PCR products stained with ethidium bromide in 1.2% agarose gels.
Targeted degradation of bcr-abl mRNA in CML cells as determined by RT-PCR. RT-PCR products from bcr-abl and β-actin mRNAs were separated on agarose gels and stained with ethidium bromide. ODNs (as indicated in the figure) were added twice daily to CML cells from (A) patient 1 (5 μmol/L per treatment) or (B) patient 2 (2 μmol/L per treatment) for 2.5 days before isolation of RNA. The images show the PCR products stained with ethidium bromide in 1.2% agarose gels.
DISCUSSION
2-5A–antisense molecules complementary to either of two regions in the bcr/abl mRNA, the translation start site or the fusion site, caused bcr/abl mRNA to be selectively degraded by RNase L both in a cell-free system and in intact CML cells (Figs 1 and 7). Selective targeting of these specific regions in bcr/abl RNA was observed in cell-free reactions containing in vitro synthesized RNA, 2-5A–antisense and human RNase L (Fig 1). The addition of the 2-5A moiety to antisense ODNs to bcr resulted in larger declines in p210bcr/ablkinase activity levels than was obtained with the antisense sequence by itself, thus implicating RNase L in the effect (Fig 2). Proliferation of the Ph1 positive cell line K562 and of primary bone marrow CML cells was also suppressed by 2-5A–antisense treatment (Figs2 to 6). Loss of the cell survival function associated with p210bcr/abl could account for the substantial decrease in the CML cell populations treated with 2-5A–antisense species to bcr/abl.8 9 While the 2-5A–antisense ODNs directed to the bcr/abl fusion site are expected to be specific for the bcr/abl mRNA, the ODNs targeted to the translational start sequence might also cause degradation of bcr mRNA. Therefore, the anti-CML effect ofSpA4-anti-bcr could be due to a combination of targeting both the bcr and bcr/abl mRNAs. However, the absence of an effect ofSpA4-anti-bcr on normal bone marrow cells suggests that targeting bcr mRNA may have minimal effects on normal cell survival.
The advantages and specificity of the 2-5A–antisense approach was shown with a panel of control ODNs. Our results indicate enhanced inhibitory effects of attaching functional 2-5A molecules to antisense ODNs. For instance, attaching a dimeric form of 2-5A, which lacks the ability to activate RNase L, to antisense to bcr/abl had no anticellular activity on K562 cells and only minimal activity against primary CML cells (Figs 3A, 6A and C). Similarly, antisense ODNs lacking 2-5A were consistently much less active than corresponding 2-5A–antisense chimeras (Figs 2, 3, and 6). These findings strongly implicate RNase L in the mechanism of action of 2-5A–antisense. Specificity of this approach is also apparent from the decreased activities of 2-5A chimeras containing several mismatched bases in the DNA moieties. Nevertheless, the mismatched ODN, SpA4-(M5)anti-bcr retained a surprising level of activity against CML cells (Fig 6). Although this ODN has five mismatches out of a total of 18 nucleotides in the continuous 3′->5′ antisense sequence, it was able to reduce CML cell growth and bcr/abl mRNA levels (Figs 3, 6, and 7). However, the lack of an effect on the normal cells suggest an increased susceptibility of CML cells to nonspecific effects of the ODNs (Fig 6E and F). Because the CML cells used in this study had the b3/a2 form of bcr/abl, further evidence for specificity was obtained with SpA4-anti-b2a2, complementary to a b2a2 type of fusion site. This ODN showed greatly reduced effects on K562 cells, CML cells from patient 3, and on the bcr/abl mRNA from patient 1, but it had an unexplained, substantial activity against CML cells and bcr/abl mRNA from patient 2 (Figs 3, 6, and 7). Despite partial effects shown by some control ODNs, the 2-5A–antisense ODNs,SpA4-anti-b3a2 orSpA4-anti-bcr, had the greatest level of activities against CML cell growth and survival in every experiment.
The efficacy and specificity of the 2-5A–antisense approach was apparent at the molecular level. Bcr/abl mRNA was depleted by treatment of CML cells with 2-5A–antisense molecules complementary to either of two regions in the bcr/abl mRNA (Fig 7), while levels of β-actin mRNA were unaffected. The selective loss of bcr/abl mRNA and reductions in p210bcr/abl kinase activity levels in the 2-5A–antisense treated cells are also consistent with a mode of action involving RNase L (Figs 2 and 7).
These results suggest that the 2-5A–antisense strategy may eventually provide an effective method for purging hematopoietic stem cell cultures of CML cells or perhaps even of controlling CML in vivo. Further improvements in this technology might include facilitating uptake of the ODNs into the CML cells. Because other gene products, such as c-myb or c-myc13,33 play roles in leukemia, the combination of 2-5A–antisense targeted against different oncogene mRNAs or the combination with other purging reagents like cyclophosphamide derivatives might also be expected to enhance the therapeutic efficacy of 2-5A–antisense ODNs.12
ACKNOWLEDGMENT
We thank Roland Mertelsmann (University of Freiburg) for discussions and encouragement, Gerald A. Hoeltge (Cleveland Clinic) for the chromosome analysis, and to Owen N. Witte (UCLA) for the gift of plasmid [0]p210 and anti-abl antisera.
A.M. and C.F.W. were equal contributors to this study.
Supported in part by United States Public Health Service Grant No. 1 PO1 CA 62220 awarded by the Department of Health and Human Services, National Cancer Institute and by funds from Atlantic Pharmaceuticals, Inc (to R.I.I.S.) by a Cooperative Research and Development Agreement between the National Institutes of Health and Atlantic Pharmaceuticals, Inc, Raleigh, NC (to P.F.T.) and a grant from the Deutsche Forschungsgemeinschaft (to C.F.W.).
The publication costs of this article were defrayed in part by page charge payment. This article must therefore be hereby marked “advertisement” in accordance with 18 U.S.C. section 1734 solely to indicate this fact.
REFERENCES
Author notes
Address reprint requests to Robert Silverman, PhD, Department of Cancer Biology, NB40, The Lerner Research Institute, Cleveland Clinic Foundation, 9500 Euclid Ave, Cleveland, OH 44195; e-mailsilverr@cesmtp.ccf.org.