Abstract
Understanding the pattern of gene expression during erythropoiesis is crucial for a synthesis of erythroid developmental biology. Here, we isolated 4 distinct populations at successive erythropoietin-dependent stages of erythropoiesis, including the terminal, pyknotic stage. The transcriptome was determined using Affymetrix arrays. First, we demonstrated the importance of using defined cell populations to identify lineage and temporally specific patterns of gene expression. Cells sorted by surface expression profile not only express significantly fewer genes than unsorted cells but also demonstrate significantly greater differences in the expression levels of particular genes between stages than unsorted cells. Second, using standard software, we identified more than 1000 transcripts not previously observed to be differentially expressed during erythroid maturation, 13 of which are highly significantly terminally regulated, including RFXAP and SMARCA4. Third, using matched filtering, we identified 12 transcripts not previously reported to be continuously up-regulated in maturing human primary erythroblasts. Finally, using transcription factor binding site analysis, we identified potential transcription factors that may regulate gene expression during terminal erythropoiesis. Our stringent lists of differentially regulated and continuously expressed transcripts containing many genes with undiscovered functions in erythroblasts are a resource for future functional studies of erythropoiesis. Our Human Erythroid Maturation database is available at https://cellline.molbiol.ox.ac.uk/eryth/index.html.
Introduction
The study of hematopoiesis has increased our understanding of stem cells and how they differentiate along the lymphoid and myeloid lineages in response to transcription factors.1-3 To date, much of the interest has focused on the very early stages in this process, providing insight into those involved in differentiating from a stem cell to a committed progenitor.2,3 Erythropoiesis is a specialized process producing a cell dedicated to the delivery of oxygen to the tissues. As erythroblasts mature, they decrease in size, synthesize more hemoglobin, and undergo membrane reorganization and chromatin condensation. Eventually, the cells expel their organelles before taking up the biconcave discoid structure.
Understanding the pattern of gene expression and identifying the specific genes expressed during erythropoiesis are crucial for a synthesis of erythroid developmental biology and defining the molecular pathology of congenital and acquired dyserythropoietic syndromes.
Previous erythroblast microarray expression studies have focused on mouse cells or knockout models4-7 or used human cell lines8 or mixed populations of human erythroid cells of various purity and maturity.9-13 We have cultured erythroblasts from peripheral blood mononucleocytes and used flow cytometric cell sorting to isolate discrete populations of colony-forming unit-erythroid (CFU-E), pro-erythroblasts (Pro-E), intermediate (Int-E), and late (Late-E) erythroblasts.
Clustering transcripts using available proprietary software has a number of limitations; in particular, the lack of knowledge of the number of clusters, the large variability of changes in expression levels of biologically relevant coregulated genes, and the availability of measurements at a very few time points for each gene (in our dataset, this number is 4). To address these issues, we have begun to explore other methods and we present analysis of these data using matched filtering methods.
Transcription factors are integral to commitment3 ; GATA1 is essential for erythroid maturation. GATA1−/− erythroblasts in vitro fail to mature beyond the Pro-E stage, suggesting that GATA1 is essential for erythroblast survival, whereas friend of GATA1 and Tal1 are also integral to erythropoiesis. A comprehensive description of the genes expressed during erythroid development has now allowed us to analyze transcription factor binding sites (TFBSs) of erythroid-specific transcripts.
To elucidate the mechanisms involved in terminal red blood cell development, we have studied global gene expression patterns in human erythroid progenitors at several stages of differentiation using Affymetrix HGU133_plus_2.0 arrays. Here we present a comprehensive reference expression database of pure populations of normal human erythroblasts, including several previously undescribed differentially regulated erythroid genes with unknown roles in erythropoiesis. Our data will facilitate studies of erythropoiesis in health and disease.
Methods
Cell culture
Human primary differentiating erythroblasts derived from peripheral blood buffy coat (National Health Service Blood and Transplant) were obtained with informed consent in accordance with the Declaration of Helsinki. All experiments were conducted with approval from the United Kingdom Home Office. Erythroblasts were cultured using a 2-phase liquid culture system as previously described.14-16 In brief, peripheral blood mononucleocytes were obtained from peripheral blood buffy coat by Ficoll-Hypaque density-gradient centrifugation and seeded at 2.5 × 106 cells/mL in minimum essential medium with 10% fetal calf serum, 1 μg/mL cyclosporine A, and 10% conditioned medium from the 5637 bladder carcinoma cell line. The cells were cultured for 6 to 7 days (phase 1), and the nonadherent cells were washed and reseeded in fresh medium with 30% fetal calf serum, 1% deionized bovine serum albumin, 1 U/mL of human recombinant erythropoietin (EPO), 10−5M β-mercaptoethanol, 10−6M dexamethasone, 0.3 mg/mL holotransferrin (ICN Biochemicals), and 10 ng/mL of human stem cell factor (phase 2). Maturation of erythroblasts was monitored by May-Grünwald-Giemsa-stained cytospin preparations using light microscopy. Cells were harvested after phase 2 culture in the presence of EPO for 4, 5 to 6, 8 to 11, or 13 to 15 days to obtain populations of CFU-E, Pro-Es, Int-Es, or Late-Es, respectively.
Cell sorting
Cells were stained and sorted using anti-CD36–phycoerythrin (BD Biosciences; used for CFU-E, Pro-Es, and Int-Es), anti-CD71–fluorescein isothiocyanate (Dako North America), anti-CD235a–allophycocyanin (BD Biosciences; used for CFU-E, Pro-Es, and Int-Es), and anti-CD235a–R-phycoerythrin (Dako North America; used for Late-Es).
RNA extraction
The purity and morphology of all sorted erythroblast populations were confirmed by cytospin. Total RNA was extracted using Trizol (Invitrogen) according to the manufacturer's instructions, appending a phenol-chloroform step. RNA integrity was evaluated using an Agilent Bioanalyzer 2100 (Agilent Technologies).
Biotinylated cRNA preparation and hybridization
For each of the 4 stages of erythroid precursors included in the study, 3 RNA pools were prepared from nonoverlapping sets of 4 different RNA preparations mixed in equal proportions. One RNA pool of 4 RNA preparations from unsorted cells harvested at each of the 4 different stages of culture was also prepared for comparison. Biotinylated fragmented cRNA was synthesized from 100 ng of pooled RNA using the 2-Cycle cDNA Synthesis and the 2-Cycle Target Labeling and Control Reagent kits (Affymetrix), and hybridized to GeneChip Human Genome U133_Plus_2.0 arrays (Affymetrix) following the manufacturer's recommendations.
Analysis of microarray data
Cell intensity calculation and scaling were performed using GeneChip Operating Software (Affymetrix) and data analyzed by the Robust Multiarray Average (RMA) method of Irizarry et al17 using the Bioconductor suite of packages in R (www.bioconductor.org). Data preprocessing and statistical analysis of differential expression in R used the Linear Models for Microarray Data (LIMMA) package18 from Bioconductor. The returned B values (log odds of differential expression) and P values were used to cluster the data.
Heat maps of RNA expression were generated using Genesis.19 DiRE,20 Pscan Version 1.0,21 and CONFAC Version 1.0 22 were used for TFBS discovery. Ingenuity Pathways Analysis (Ingenuity Systems) and DAVID23 were used for ontology mapping.
Sets of normally distributed data were compared using Student t tests in GraphPad Prism, Version 5 (GraphPad Software).
To compare our data from Affymetrix HGU133_plus_ Version 2.0 with previously published data24 from Illumina HumanWG-6 Version 2 Expression BeadChips, we cross-referenced the unique probe identifiers (PIDs) using the IlluminaV2_AffyMapping.txt and HumanWG-6 Version 2 manifest files available at www.switchtoi.com and external gene names. A total of 33 966 Illumina PIDs matched annotated Affymetrix PIDs representing 15 858 genes from the Illumina array. Of 7599 Illumina IDs present in erythroblasts,24 7150 cross-matched to 6976 Affymetrix PIDs representing 6911 genes, represented on the Affymetrix HGU133_plus_2.0 chip by 16 120 Affymetrix PIDs.
Our data are available through National Center for Biotechnology Information Gene Expression Omnibus25 using accession number GSE22552 (www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc = GSE22552).
We have also retrospectively analyzed other datasets9-12 deposited with National Center for Biotechnology Information Gene Expression Omnibus containing expression measurements in erythroblasts at more than one time point of erythropoiesis as indicated in Table 1, using RMA (where the CEL files are available) and LIMMA, for comparison with our data.
Analysis of other datasets9-12 deposited with NCBIGEO
| Study . | NCBIGEO accession no. . | CEL files deposited? . | No. of stages . | No. of replicates . | Pooled or single samples hybridized? . | EB source and culture conditions . | FACS-sorted . | % genes DE . | % genes SIG DE . |
|---|---|---|---|---|---|---|---|---|---|
| Keller et al9 (2006) | GSE4655 | No | 6 | 3 | Some pooled samples | CD34+ progenitors in a novel serum-free system | No | 12 | 3.8 |
| Singleton et al11 (2008) | GSE10584 | Yes | 3 | 4 | Single samples | CD34+ progenitors | No | 8 | 0.1 |
| Peller et al10 (2009) | GSE12407 | Yes | 3 | 2 | Single samples | Fibachs | No | 0 | 0.0 |
| Sripichai et al12 (2009) | GSE7874 | Yes | 2 | 5 | Single samples | CD34+ progenitors cultured for 7 and 14 days + EPO/−SCF | No | 47 | 0.3 |
| Current study | GSE22552 | Yes | 4 | 3 | Pooled samples | Fibachs | Yes | 43 | 3.8 |
| Study . | NCBIGEO accession no. . | CEL files deposited? . | No. of stages . | No. of replicates . | Pooled or single samples hybridized? . | EB source and culture conditions . | FACS-sorted . | % genes DE . | % genes SIG DE . |
|---|---|---|---|---|---|---|---|---|---|
| Keller et al9 (2006) | GSE4655 | No | 6 | 3 | Some pooled samples | CD34+ progenitors in a novel serum-free system | No | 12 | 3.8 |
| Singleton et al11 (2008) | GSE10584 | Yes | 3 | 4 | Single samples | CD34+ progenitors | No | 8 | 0.1 |
| Peller et al10 (2009) | GSE12407 | Yes | 3 | 2 | Single samples | Fibachs | No | 0 | 0.0 |
| Sripichai et al12 (2009) | GSE7874 | Yes | 2 | 5 | Single samples | CD34+ progenitors cultured for 7 and 14 days + EPO/−SCF | No | 47 | 0.3 |
| Current study | GSE22552 | Yes | 4 | 3 | Pooled samples | Fibachs | Yes | 43 | 3.8 |
The number of replicates and stages of erythroblasts in each study is indicated. RMA (where the CEL files are available) and LIMMA were used to determine the proportion of genes that were: differentially expressed (DE) at the lowest stringency equal to DE1 (B value of at least 2.945, representing a 95% probability that the gene was differentially expressed, and expression levels in all replicates at any stage > 100); significantly differentially expressed (SIG DE) at the highest stringency equal to DE_Sig (minimum average expression threshold of 1000, minimum fold change threshold of 10-fold, only PIDs with a difference of at least 1000 between minimum and maximum average expression were selected).
NCBIGEO indicates National Center for Biotechnology Information Gene Expression Omnibus; and FACS, fluorescence-activated cell sorter.
Web-accessible database
After RMA and LIMMA analysis, profiles were clustered by calculating the log ratios of adjacent time points, and the resulting matrix digitized by choosing a nominal threshold, “t.” Ratios were replaced by 1 if ratio was more than t, −1 if less than −t, and 0 otherwise. Profiles with the same digital pattern were assigned to the same cluster.
The output was tabulated and stored in a MySQL database, and a custom web-interface was built to query the data using perl-cgi. The Human Erythroblast Maturation database is available at https://cellline.molbiol.ox.ac.uk/eryth/index.html.
Quantitative PCR
cDNA was synthesized from 100 ng RNA taken from individual samples using the High Capacity cDNA Reverse Transcription Kit (Applied Biosystems) after DNAse treatment. Semiquantitative polymerase chain reaction (PCR) was performed using TaqMan Universal PCR Master Mix (Applied Biosystems) and intron-spanning primers and probes from the Roche Universal Probe Library (supplemental Table 1, available on the Blood Web site; see the Supplemental Materials link at the top of the online article). Novel transcripts identified by microarray were selected for quantitative PCR if an efficient Universal Probe Library assay was available. Expression data for each transcript were normalized to that for the reference gene Ribosomal Protein L13A (RPL13A)26 and then quantified in Pro-Es, Int-Es, and Late-Es relative to the expression level in CFU-E, after running dilution standards for every assay to ensure the ratio of the differences in cycle thresholds (2−ΔΔCT) was equivalent to the fold change calculated from the standard curve.
Results
Sorted erythroblast populations exhibit expected cell surface marker expression and morphology
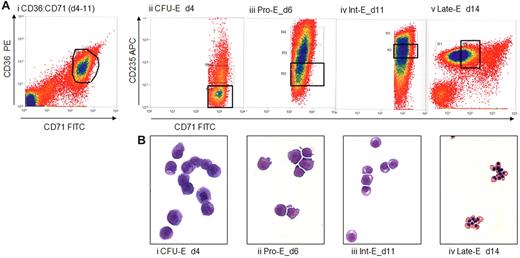
Erythroblasts were cultured from 83 Blood Transfusion Service buffy coats and sorted, by CD36, CD71, and CD235a expression and size, into enriched CFU-E, Pro-E, Int-E, and Late-E populations (Figure 1A). Typically, 5 × 105 to 107 sorted cells were obtained in the final gates, with a typical RNA yield of 2 μg from 106 cells. Postsort analysis of 2 representative populations is shown in supplemental Figure 1.
Preparation of enriched populations of CFU-E, Pro-Es, Int-Es, and Late-Es. (A) CD36, CD71 (transferrin receptor 1) and CD235a expression determined by flow cytometry of peripheral blood mononucleocyte-derived erythroblasts. Cells were stained for 15 minutes at 4°C in 2% bovine serum albumin in HBSS and washed twice. CFU-E are CD36+, CD71+, and CD235a−/low. CD235a expression increases until the intermediate erythroblast stage, when it remains high and CD71 expression begins to decrease. Stained cells were sorted on a MoFlo cell sorter (Beckman Coulter) to obtain erythroblast populations enriched for the appropriate stage of development. Our cells were tightly gated to ensure discrete populations of cells similar in maturity and lineage. Sort gates were defined by first setting a gate on the fluorescent expression profile of the population of interest, either CD71 versus CD36 (i) and/or CD71 versus CD235a (ii-v). These gates were then applied to the forward scatter versus side scatter dot plot, and a second gate was applied to exclude debris and also to define a population of cells showing similar size. The mean percentages of sorted cells collected in the total erythroid gates were 14%, 24%, 20%, and 50% for cells sorted on days 4, 5 to 6, 8 to 11, and 13 to 15, respectively. Sorted CFU-E, Pro-Es, Int-Es, and Late-Es represented an average of 28%, 29%, 37%, and 45% of erythroid cells present, respectively. (B) Cytospins of cells stained with May-Grünwald-Giemsa are shown to illustrate the morphologic changes as cultured erythroblasts mature from large CFU-E with a large, smooth nucleus at day 4, through the Pro-E and basophilic Int-E stages as the chromatin becomes more condensed, accompanied by a reduction in both cell size and the nucleus/cell size ratio, to the small Late-Es when the cytoplasm turns from blue to orange reflecting the production of hemoglobin and the nuclei become pyknotic. From all hybridized sorts, we have counted at least 250 cells per sample and find an average of 99.3% purity for our populations, with the contaminants being erythroid cells of slightly increased or decreased maturity rather than nonerythroid cells. Representative plots and cytospins viewed at 20× original magnification are shown.
Preparation of enriched populations of CFU-E, Pro-Es, Int-Es, and Late-Es. (A) CD36, CD71 (transferrin receptor 1) and CD235a expression determined by flow cytometry of peripheral blood mononucleocyte-derived erythroblasts. Cells were stained for 15 minutes at 4°C in 2% bovine serum albumin in HBSS and washed twice. CFU-E are CD36+, CD71+, and CD235a−/low. CD235a expression increases until the intermediate erythroblast stage, when it remains high and CD71 expression begins to decrease. Stained cells were sorted on a MoFlo cell sorter (Beckman Coulter) to obtain erythroblast populations enriched for the appropriate stage of development. Our cells were tightly gated to ensure discrete populations of cells similar in maturity and lineage. Sort gates were defined by first setting a gate on the fluorescent expression profile of the population of interest, either CD71 versus CD36 (i) and/or CD71 versus CD235a (ii-v). These gates were then applied to the forward scatter versus side scatter dot plot, and a second gate was applied to exclude debris and also to define a population of cells showing similar size. The mean percentages of sorted cells collected in the total erythroid gates were 14%, 24%, 20%, and 50% for cells sorted on days 4, 5 to 6, 8 to 11, and 13 to 15, respectively. Sorted CFU-E, Pro-Es, Int-Es, and Late-Es represented an average of 28%, 29%, 37%, and 45% of erythroid cells present, respectively. (B) Cytospins of cells stained with May-Grünwald-Giemsa are shown to illustrate the morphologic changes as cultured erythroblasts mature from large CFU-E with a large, smooth nucleus at day 4, through the Pro-E and basophilic Int-E stages as the chromatin becomes more condensed, accompanied by a reduction in both cell size and the nucleus/cell size ratio, to the small Late-Es when the cytoplasm turns from blue to orange reflecting the production of hemoglobin and the nuclei become pyknotic. From all hybridized sorts, we have counted at least 250 cells per sample and find an average of 99.3% purity for our populations, with the contaminants being erythroid cells of slightly increased or decreased maturity rather than nonerythroid cells. Representative plots and cytospins viewed at 20× original magnification are shown.
May-Grünwald-Giemsa-stained cytospins of cells illustrate the morphologic changes within this culture as erythroblasts mature (Figure 1B). Maturation is associated with reduction in cell size, enhanced chromatin condensation, and increased hemoglobinization. Ninety-seven sorts from a total of 54 donors gave 71 populations of suitable purity and morphology consistent with culture time. No nonerythroid cells were counted in these 71 populations, and fewer than 2% of the cells were of an earlier or later stage of maturation, as judged by inspection and counting the postsort cytospins, and consistent with observations in a previous study, which verified this method.14
Transcriptome analysis
Forty-eight RNA samples from 38 donors in 12 pools of 4 were hybridized to 12 Affymetrix HGU133_plus_2.0 arrays, 3 each of CFU-E, Pro-E, Int-E, and Late-E.
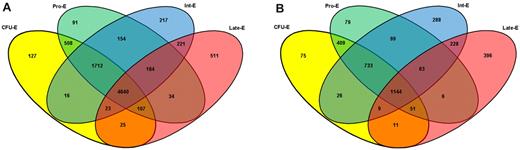
RMA analysis was first used to normalize the data.17 Expression levels of less than 100 in any replicate were considered to be below background. A total of 8570 genes were expressed at a significant level at any stage of erythroblast maturation (supplemental Table 2), whereas a further 12 094 genes represented on the HGU133_plus_2.0 arrays were absent from all stages studied. This equates to 24.5% PIDs on the array being present at any stage and is slightly more stringent than the percentage present called by the GeneChip Operating Software before normalization, which ranged from 28% to 42%. The distribution of genes present at all combinations of stages is shown in Figure 2A. A total of 8450 expressed genes were also present in previously published erythroid studies.4-13 The remaining 120 genes remained below an expression level of 500 at all stages. Figure 2B shows the distribution pattern of more highly expressed genes, including only those genes that achieved an expression value of 500 in all 3 pools at any one stage. Ingenuity Pathways Analysis was used to analyze the molecular and cellular functions enriched in these lists of genes present during erythroblast development (supplemental Figure 2A-B).
Four-way Venn diagram illustrating the overlap of genes expressed in all pools at different stages of erythroblast maturation. (A) Overlap of genes expressed at successive stages of erythroblast maturation. (B) Overlap of highly expressed transcripts at successive stages of erythroblast maturation. Expression cut-offs are selected with the caveat that expression measurements on array platforms are relative and prone to influence by physical hybridization factors. Numbers of transcripts are shown that exceeded an expression level of 100 in all 3 triplicates at each stage. A total of 12 094 genes represented on the HGU133_plus_2.0 arrays were absent from all stages studied. Numbers of transcripts that exceeded an expression level of 500 in all 3 triplicates at the combinations of stages indicated. A total of 4640 genes are common to all stages, whereas other genes show a more stage-specific expression pattern; for example, 511 are only present at the Late-E stage (red). When the expression cut-off is increased to 500, the number of transcripts meeting that criteria at all stages drops to 1144, and 396 genes exceed an expression level of 500 at the Late-E stage only.
Four-way Venn diagram illustrating the overlap of genes expressed in all pools at different stages of erythroblast maturation. (A) Overlap of genes expressed at successive stages of erythroblast maturation. (B) Overlap of highly expressed transcripts at successive stages of erythroblast maturation. Expression cut-offs are selected with the caveat that expression measurements on array platforms are relative and prone to influence by physical hybridization factors. Numbers of transcripts are shown that exceeded an expression level of 100 in all 3 triplicates at each stage. A total of 12 094 genes represented on the HGU133_plus_2.0 arrays were absent from all stages studied. Numbers of transcripts that exceeded an expression level of 500 in all 3 triplicates at the combinations of stages indicated. A total of 4640 genes are common to all stages, whereas other genes show a more stage-specific expression pattern; for example, 511 are only present at the Late-E stage (red). When the expression cut-off is increased to 500, the number of transcripts meeting that criteria at all stages drops to 1144, and 396 genes exceed an expression level of 500 at the Late-E stage only.
The GeneChip Operating Software percentage present in the hybridizations of RNA from unsorted cells ranged from 38% to 45%. We normalized hybridization files from the unsorted RNA populations together with those from the sorted populations. In the unsorted populations, 7352 genes were present in cultures at all time points, representing 75% of the total 9766 genes present at any time point. In sorted erythroblasts, 55% of the genes present at any stage were present at all stages, when normalized with unsorted cell data. This indicates that sorting the cells to eliminate nonerythroid cells and work with defined erythroblast populations considerably improves detection of differentially expressed transcripts.
Validation of erythroid gene expression by quantitative PCR
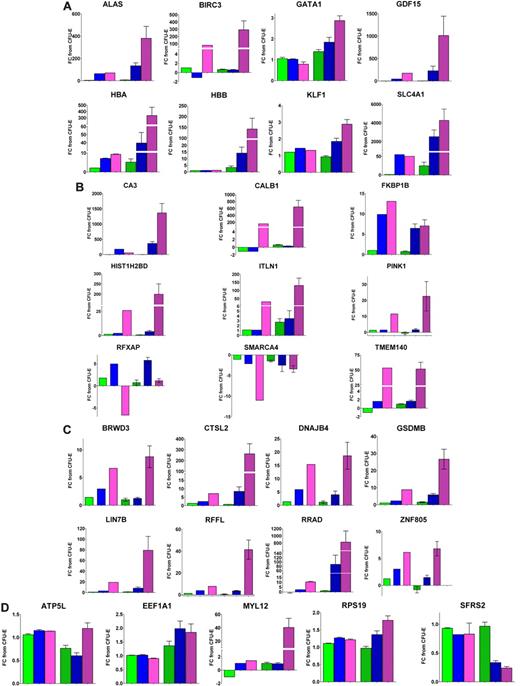
Quantitative PCR was used to confirm that array signal intensity reflected the trend in gene expression for a selection of highly differentially expressed genes and genes expressed at a consistently high level (Figure 3; supplemental Figure 3).
Quantitative PCR confirmation of transcript fold changes from microarray expression profiling. Transcripts were selected for quantitative PCR based on the availability of an efficient intron-spanning quantitative PCR assay. For each gene, the fold change is given between CFU-E and Pro-E (C-P, green bars), Int-E (C-I, blue bars), and Late-E (C-L, pink bars). Solid bars on the left-hand side of each graph represent microarray data; and hatched bars on the right, quantitative PCR data in triplicate from at least 3 biologic replicates of individual samples, including some additional samples not included in the pools on the Affymetrix arrays. (A) Previously described erythroid genes. (B) Differentially regulated transcripts from DE_Sig. (C) Differentially regulated transcripts identified by matched filter analysis (MFUp). (D) Transcripts that are continuously expressed at a high level throughout maturation (CHE).
Quantitative PCR confirmation of transcript fold changes from microarray expression profiling. Transcripts were selected for quantitative PCR based on the availability of an efficient intron-spanning quantitative PCR assay. For each gene, the fold change is given between CFU-E and Pro-E (C-P, green bars), Int-E (C-I, blue bars), and Late-E (C-L, pink bars). Solid bars on the left-hand side of each graph represent microarray data; and hatched bars on the right, quantitative PCR data in triplicate from at least 3 biologic replicates of individual samples, including some additional samples not included in the pools on the Affymetrix arrays. (A) Previously described erythroid genes. (B) Differentially regulated transcripts from DE_Sig. (C) Differentially regulated transcripts identified by matched filter analysis (MFUp). (D) Transcripts that are continuously expressed at a high level throughout maturation (CHE).
For most of the differentially expressed genes analyzed, expression ratios were confirmed by quantitative PCR of individual samples to be even greater than those observed using microarray hybridization of pooled samples, establishing our dataset as a useful resource. Many of the genes that appeared to be expressed at a consistently high level from the microarray data were shown by quantitative PCR to be more differentially expressed. Previous investigators have also reported greater fold changes detected using quantitative PCR compared with microarray for ALAS, GATA1, and KLF1 (Figure 3A) during erythropoiesis.9 Significant discrepancies included HBB, which appeared to be highly expressed at all maturation stages after microarray hybridization, but quantitative PCR confirmed differential HBB expression as expected (Figure 3A). A large difference in HBB fold changes detected by microarray compared with quantitative PCR during erythropoiesis has been previously reported,9 and it is commonly observed that quantitative PCR is a more sensitive indicator of expression fold changes; the abundance of globin mRNA is well documented to interfere with the ability of microarray hybridizations to accurately detect significant biomarkers in whole blood.27
Differentially expressed genes: LIMMA
After LIMMA analysis, pairwise comparisons of gene expression between stages were considered significant if the B value was at least 2.945 (representing a 95% probability that the gene was differentially expressed) and expression levels in all 3 replicates at any stage were more than 100. The pattern of B values for comparisons of expression of a PID between any pair of stages (CFU-E to Pro-E, CFU-E to Int-E, CFU-E to Late-E, Pro-E to Int-E, Pro-E to Late-E, and Int-E to Late-E) was used as a basis of clustering (supplemental Table 3). This lowest stringency list of 3679 differentially expressed genes (Differentially Expressed 1 [DE1]; supplemental Table 3) included a large cluster of 1180 terminally down-regulated genes, which were significantly down-regulated from CFU-E to Late-E, Pro-E to Late-E, and Int-E to Late-E but no other combinations of pairs of stages. Approximately one-third of genes in DE1 (1174) were differentially expressed in previously published erythroid studies,4-13 and a further 258 were differentially expressed in our reanalysis of existing data9,11 (Table 1; supplemental Table 3). Another 1096 genes were differentially expressed in maturing erythroblasts cultured in the absence of stem cell factor (SCF), which promotes erythroid expansion and delays differentiation,28 leaving the final one-third (1151) genes, which have not been previously observed to be differentially expressed in maturing erythroblasts.
DE1 was refined by increasing the minimum average expression threshold to 1000 and imposing a minimum fold change threshold of 10-fold between stages. Finally, PIDs were selected whose minimum and maximum average expression levels between any stage varied by at least 1000, so that the most stringent list (DE_Sig; supplemental Table 4) of significantly differentially regulated transcripts is composed of 327 genes, which have a minimum average expression level of 1000 for at least one stage, a minimum fold change of 10-fold between any pair of stages, and a minimum difference of 1000 between maximum and minimum average expression at any stage. This list of differentially regulated genes contains many erythroblast-specific genes,24 and all of these 327 genes or their homologs have been previously observed to be expressed in at least one erythroid expression dataset.4-13 A total of 168 genes in DE_Sig were previously shown to be differentially expressed in maturing erythroid cells.4-13 This is the first report of significant differential expression during erythropoiesis of the remaining 159 genes. Our reanalysis shows that 43 of these were also differentially expressed in existing datasets of maturing erythroblasts,9,11 and a further 110 were regulated in the absence of SCF.12 The remaining 6 genes were differentially expressed in our dataset only. Table 2 lists the functions of some of the genes, which have not previously been described as differentially expressed in primary erythroblasts. Quantitative PCR was used to confirm expression profiles of selected significantly differentially regulated transcripts (Figure 3B; supplemental Figure 3).
Selected genes in DE_SigUp or MFUp that have not been previously described as differentially regulated in erythroblasts
| Gene . | Symbol . | List . | Largest significant FC between erythroid stages* . | Function . | DE in a reanalyzed dataset? (reference) . | DE in a previous cell line study? (reference) . |
|---|---|---|---|---|---|---|
| Carbonic anhydrase 3 | CA3 | DE_Sig | 177.7x C-I | Muscle-specific, catalyses reversible hydration of CO2, inhibits apoptosis | 9 | |
| Calbindin 1 | CALB1 | DE_Sig | 173.1x I-L | Inhibits apoptosis in osteoblasts by binding to and inhibiting caspase 3 | 12 | 8 |
| FK506 binding protein 1B, 12.6 kDa | FKBP1B | DE_Sig | 13.1x C-L | Inhibits the mTOR proliferation pathway | 12 | 7 |
| Intalectin 1 | ITLN1 | DE_Sig | 31.3x C-L | Binds cell surface galactofuronidase | 12 | |
| Multiple EGF-like-domains 9 | MEGF9 | DE_Sig | 18.1x C-L | Signaling molecule, regulated during development | ||
| N-myc downstream regulated 1 | NDRG1 | DE_Sig | 10.5x C-L | Involved in hypoxia response, cell differentiation, and cell growth arrest | ||
| PTEN-induced putative kinase 1 | PINK1 | DE_Sig | 11.5x C-L | Implicated in mitochondrial quality control | ||
| Regulatory factor X-associated protein | RFXAP | DE_Sig | −32.9x I-L | In association with NFY and Creb; involved in MHC class II activation | 12 | |
| S100 calcium binding protein A4 | S100A4 | DE_Sig | −24.5x P-L | Overexpression promotes metastasis, activates NF-κB; functions as a Ca2+-activated molecular switch, regulates macrophage recruitment and chemotaxis | ||
| SWI/SNF related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 4 | SMARCA4 | DE_Sig | −11.0x C-L | Coactivates γ-globin gene | ||
| Transmembrane protein 140 | TMEM140 | DE_Sig | 26.6x C-L | Potential role in supporting HSC activity of a stromal cell line | 12 | |
| Zinc finger protein 451 | ZNF451 | DE_Sig | 11.5x C-L | ZNF451 is a novel PML body- and SUMO-associated transcriptional co-regulator | ||
| Histone cluster 1, H2bd | HIST1H2BD | DE_Sig & MFUp | 26.0x C-L | 9,12 | ||
| Bromodomain and WD repeat domain containing 3 | BRWD3 | MFUp | 6.7x C-L | May have a role in JAK/STAT signaling | ||
| Cathepsin L2 | CTSL2 | MFUp | 7.1x C-L | |||
| DnaJ (Hsp40) homolog, subfamily B, member 4 | DNAJB4 | MFUp | 15.3 C-L | A tumor suppressor activated by YY1 and AP1 | 8 | |
| Gasdermin B | GSDMB | MFUp | 8.8x C-L | Involved in regulation of apoptosis in the gastrointestinal epithelium | ||
| Importin 11 | IPO11 | MFUp | 12.1x C-L | Increases the intensity of pleiotrophic cytokine responses | ||
| Lin-7 homolog B (C elegans) | LIN7B | MFUp | 19.3 C-L | Involved in Ras signaling; effects include cell growth, differentiation, and survival | ||
| Nuclear paraspeckle assembly transcript 1 (non—protein-coding) | NEAT1 | MFUp | 28.7x C-L | RNA component of paraspeckles, ribonucleoprotein structures involved in regulating the expression of certain genes in differentiated cells by nuclear retention of RNA | ||
| Retinoic acid receptor responder (tazarotene induced) 3 | RARRES3 | MFUp | 5.2x C-L | Retinoid-inducible growth regulator, induces apoptosis | ||
| Rififylin | RFFL | MFUp | 7.7x C-L | Inhibits recycling from the endocytic recycling compartment | ||
| Ras-related associated with diabetes | RRAD | MFUp | 12.5x P-L | Involved in Ras signalling; effects include cell growth, differentiation, and survival | 12 | 7 |
| Lysine (K)-specific demethylase 6A | KDM6A | MFUp | 7.8x C-L | Involved in chromatin remodeling | ||
| Zinc finger protein 805 | ZNF805 | MFUp | 6.2x C-L |
| Gene . | Symbol . | List . | Largest significant FC between erythroid stages* . | Function . | DE in a reanalyzed dataset? (reference) . | DE in a previous cell line study? (reference) . |
|---|---|---|---|---|---|---|
| Carbonic anhydrase 3 | CA3 | DE_Sig | 177.7x C-I | Muscle-specific, catalyses reversible hydration of CO2, inhibits apoptosis | 9 | |
| Calbindin 1 | CALB1 | DE_Sig | 173.1x I-L | Inhibits apoptosis in osteoblasts by binding to and inhibiting caspase 3 | 12 | 8 |
| FK506 binding protein 1B, 12.6 kDa | FKBP1B | DE_Sig | 13.1x C-L | Inhibits the mTOR proliferation pathway | 12 | 7 |
| Intalectin 1 | ITLN1 | DE_Sig | 31.3x C-L | Binds cell surface galactofuronidase | 12 | |
| Multiple EGF-like-domains 9 | MEGF9 | DE_Sig | 18.1x C-L | Signaling molecule, regulated during development | ||
| N-myc downstream regulated 1 | NDRG1 | DE_Sig | 10.5x C-L | Involved in hypoxia response, cell differentiation, and cell growth arrest | ||
| PTEN-induced putative kinase 1 | PINK1 | DE_Sig | 11.5x C-L | Implicated in mitochondrial quality control | ||
| Regulatory factor X-associated protein | RFXAP | DE_Sig | −32.9x I-L | In association with NFY and Creb; involved in MHC class II activation | 12 | |
| S100 calcium binding protein A4 | S100A4 | DE_Sig | −24.5x P-L | Overexpression promotes metastasis, activates NF-κB; functions as a Ca2+-activated molecular switch, regulates macrophage recruitment and chemotaxis | ||
| SWI/SNF related, matrix-associated, actin-dependent regulator of chromatin, subfamily a, member 4 | SMARCA4 | DE_Sig | −11.0x C-L | Coactivates γ-globin gene | ||
| Transmembrane protein 140 | TMEM140 | DE_Sig | 26.6x C-L | Potential role in supporting HSC activity of a stromal cell line | 12 | |
| Zinc finger protein 451 | ZNF451 | DE_Sig | 11.5x C-L | ZNF451 is a novel PML body- and SUMO-associated transcriptional co-regulator | ||
| Histone cluster 1, H2bd | HIST1H2BD | DE_Sig & MFUp | 26.0x C-L | 9,12 | ||
| Bromodomain and WD repeat domain containing 3 | BRWD3 | MFUp | 6.7x C-L | May have a role in JAK/STAT signaling | ||
| Cathepsin L2 | CTSL2 | MFUp | 7.1x C-L | |||
| DnaJ (Hsp40) homolog, subfamily B, member 4 | DNAJB4 | MFUp | 15.3 C-L | A tumor suppressor activated by YY1 and AP1 | 8 | |
| Gasdermin B | GSDMB | MFUp | 8.8x C-L | Involved in regulation of apoptosis in the gastrointestinal epithelium | ||
| Importin 11 | IPO11 | MFUp | 12.1x C-L | Increases the intensity of pleiotrophic cytokine responses | ||
| Lin-7 homolog B (C elegans) | LIN7B | MFUp | 19.3 C-L | Involved in Ras signaling; effects include cell growth, differentiation, and survival | ||
| Nuclear paraspeckle assembly transcript 1 (non—protein-coding) | NEAT1 | MFUp | 28.7x C-L | RNA component of paraspeckles, ribonucleoprotein structures involved in regulating the expression of certain genes in differentiated cells by nuclear retention of RNA | ||
| Retinoic acid receptor responder (tazarotene induced) 3 | RARRES3 | MFUp | 5.2x C-L | Retinoid-inducible growth regulator, induces apoptosis | ||
| Rififylin | RFFL | MFUp | 7.7x C-L | Inhibits recycling from the endocytic recycling compartment | ||
| Ras-related associated with diabetes | RRAD | MFUp | 12.5x P-L | Involved in Ras signalling; effects include cell growth, differentiation, and survival | 12 | 7 |
| Lysine (K)-specific demethylase 6A | KDM6A | MFUp | 7.8x C-L | Involved in chromatin remodeling | ||
| Zinc finger protein 805 | ZNF805 | MFUp | 6.2x C-L |
The largest significant fold change between stages of our maturing erythroblasts is indicated with a brief summary of the function and whether it was differentially expressed in any of the reanalyzed datasets listed in Table 1, or any of the previous erythroid cell line studies. References for gene function are given in supplemental Table 8B.
C indicates CFU-E; P, Pro-E; I, Int-E; and L, Late-E.
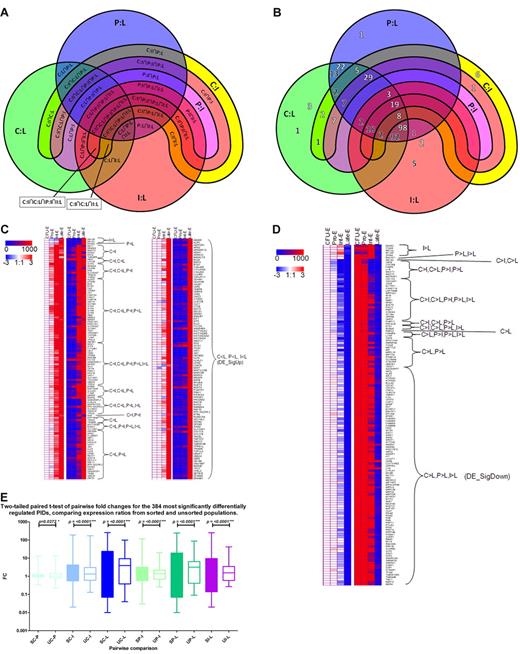
The majority of PIDs in DE_Sig, the most stringent list, fall into 2 clusters. Ninety-eight genes are terminally up-regulated (DE_SigUp), and 92 genes are terminally down-regulated (DE_SigDown; Figure 4A-D).
Expression dynamics of the most significantly regulated transcripts. (A) Key to the 5-way Venn diagram in Figure 4B of the 327 most significantly differentially expressed genes (DE_Sig) showing the overlap of fold change combinations. The 3 circles represent transcripts differentially expressed between Late-Es (L) and one of CFU-E (C, green), Pro-E (P, blue), or Int-E (I, red). The wide magenta arc represents genes differentially regulated between Pro-Es and Int-Es. The narrower yellow arc encompassing most of the perimeter of the wider arc represents genes differentially regulated between CFU-E and Int-Es. The colors merge where the shapes intersect each other. (B) Five-way Venn diagram of the 327 most significantly differentially expressed transcripts showing the overlap of fold change combinations (see panel A for key). The number of transcripts that are up-regulated during maturation are shown in white numbers with a dark shadow, whereas the number of transcripts that are down-regulated during maturation are shown in black numbers with a white shadow. Most genes are either terminally up-regulated or terminally down-regulated. For example, 98 transcripts are up-regulated between CFU-E to Late-Es, Pro-Es to Late-Es to Int-Es to Late-Es, and 92 transcripts are down-regulated between CFU-E to Late-Es, Pro-Es to Late-Es to Int-Es to Late-Es. One transcript is up-regulated between CFU-E to Int-Es and Pro-Es to Int-Es, but not between any of these stages and Late-Es. Where no number is shown, there were no transcripts in that set. (C) Heat maps of expression levels of transcripts most significantly differentially expressed (DE_Sig) up-regulated during erythroid maturation. Each column represents a different stage of erythroblast maturation, and each row represents a different PID. Red, white, and blue left-hand panels indicate log (base 2) of expression ratios between the stage indicated in the column heading and CFU-E during erythroid maturation. White represents a 1:1 ratio, so the CFU-E column is always white; red, up-regulation (log fold change of 3 indicating an 8-fold increase during maturation); and blue, down-regulation (log fold change of −3 indicating an 8-fold decrease during maturation). Red and blue right-hand panels indicate linear expression levels. Blue indicates no or low expression; and red, high expression (> 1000). Expression dynamics are indicated. (D) Heat maps of expression levels of transcripts most significantly differentially expressed (DE_Sig) down-regulated during erythroid maturation. Each column represents a different stage of erythroblast maturation, and each row represents a different PID. The red, white, and blue left-hand panel indicates log (base 2) of expression ratios between the stage indicated in the column heading and CFU-E during erythroid maturation. White represents a 1:1 ratio, so the CFU-E column is always white; red indicates up-regulation (log fold change of 3 indicating an 8-fold increase during maturation); and blue indicates down-regulation (log fold change of −3 indicating an 8-fold decrease during maturation). The red and blue right-hand panel indicates linear expression levels. Blue indicates no or low expression; and red, high expression (> 1000). Expression dynamics are indicated. (E) Significant increase in the range and magnitude of fold changes of expression of the 327 most differentially expressed transcripts (DE_Sig) from CFU-E/Pro-E (C-P), CFU-E/Int-E (C-I), CFU-E/Late-E (C-L), Pro-E/Int-E (P-I), Pro-E/Late-E (P-L), or Int-E/Late-E (I-L) in RNA from sorted erythroblasts (S) compared with that from unsorted cells (U). A 2-tailed paired t test was performed. Between each pair of stages compared, the detected fold changes were distributed over a narrower range of values in the unsorted erythroblasts compared with their sorted counterparts, indicating a dampening of the variation detected between different stages of erythroblasts in the unsorted samples.
Expression dynamics of the most significantly regulated transcripts. (A) Key to the 5-way Venn diagram in Figure 4B of the 327 most significantly differentially expressed genes (DE_Sig) showing the overlap of fold change combinations. The 3 circles represent transcripts differentially expressed between Late-Es (L) and one of CFU-E (C, green), Pro-E (P, blue), or Int-E (I, red). The wide magenta arc represents genes differentially regulated between Pro-Es and Int-Es. The narrower yellow arc encompassing most of the perimeter of the wider arc represents genes differentially regulated between CFU-E and Int-Es. The colors merge where the shapes intersect each other. (B) Five-way Venn diagram of the 327 most significantly differentially expressed transcripts showing the overlap of fold change combinations (see panel A for key). The number of transcripts that are up-regulated during maturation are shown in white numbers with a dark shadow, whereas the number of transcripts that are down-regulated during maturation are shown in black numbers with a white shadow. Most genes are either terminally up-regulated or terminally down-regulated. For example, 98 transcripts are up-regulated between CFU-E to Late-Es, Pro-Es to Late-Es to Int-Es to Late-Es, and 92 transcripts are down-regulated between CFU-E to Late-Es, Pro-Es to Late-Es to Int-Es to Late-Es. One transcript is up-regulated between CFU-E to Int-Es and Pro-Es to Int-Es, but not between any of these stages and Late-Es. Where no number is shown, there were no transcripts in that set. (C) Heat maps of expression levels of transcripts most significantly differentially expressed (DE_Sig) up-regulated during erythroid maturation. Each column represents a different stage of erythroblast maturation, and each row represents a different PID. Red, white, and blue left-hand panels indicate log (base 2) of expression ratios between the stage indicated in the column heading and CFU-E during erythroid maturation. White represents a 1:1 ratio, so the CFU-E column is always white; red, up-regulation (log fold change of 3 indicating an 8-fold increase during maturation); and blue, down-regulation (log fold change of −3 indicating an 8-fold decrease during maturation). Red and blue right-hand panels indicate linear expression levels. Blue indicates no or low expression; and red, high expression (> 1000). Expression dynamics are indicated. (D) Heat maps of expression levels of transcripts most significantly differentially expressed (DE_Sig) down-regulated during erythroid maturation. Each column represents a different stage of erythroblast maturation, and each row represents a different PID. The red, white, and blue left-hand panel indicates log (base 2) of expression ratios between the stage indicated in the column heading and CFU-E during erythroid maturation. White represents a 1:1 ratio, so the CFU-E column is always white; red indicates up-regulation (log fold change of 3 indicating an 8-fold increase during maturation); and blue indicates down-regulation (log fold change of −3 indicating an 8-fold decrease during maturation). The red and blue right-hand panel indicates linear expression levels. Blue indicates no or low expression; and red, high expression (> 1000). Expression dynamics are indicated. (E) Significant increase in the range and magnitude of fold changes of expression of the 327 most differentially expressed transcripts (DE_Sig) from CFU-E/Pro-E (C-P), CFU-E/Int-E (C-I), CFU-E/Late-E (C-L), Pro-E/Int-E (P-I), Pro-E/Late-E (P-L), or Int-E/Late-E (I-L) in RNA from sorted erythroblasts (S) compared with that from unsorted cells (U). A 2-tailed paired t test was performed. Between each pair of stages compared, the detected fold changes were distributed over a narrower range of values in the unsorted erythroblasts compared with their sorted counterparts, indicating a dampening of the variation detected between different stages of erythroblasts in the unsorted samples.
A 2-tailed t test of the most significantly differentially regulated genes comparing the microarray expression ratios observed from the well-characterized sorted cell populations with those obtained from unsorted cells shows the significant differences between all 6 pairwise stage ratios (Figure 4E) for the 327 genes. Sorting the cells permitted us to use homogeneous erythroblast populations enabling detection of a wider dynamic range of expression fold changes of transcripts throughout maturation.
Differentially expressed genes: matched filtering
Alternative ways of clustering the data using a matched filtering approach produced a list of 91 genes, which are continuously up-regulated between pairs of successive stages of erythroblasts studied (Matched Filter-derived list of up-regulated genes [MFUp]; supplemental Table 5). Essentially this process can be summarized in 3 steps: (1) the median response for each PID at each stage was calculated; (2) 3 ratios of median responses (ie, Pro-E/CFU-E, Int-E/Pro-E, and Late-E/Int-E) were calculated for each PID; (3) a list of genes was generated from the data by keeping the top 20% of the PIDs up-regulated between the CFU-E and Pro-Es, then selecting the 20% of these PIDs, which were most up-regulated between Pro-Es and Int-Es, then selecting the 20% of these PIDs, which were most up-regulated between Int-Es and Late-Es.
Fifty-two of these genes have been previously reported as differentially expressed in maturing erythroid expression studies,4-13 with a further 23 detected by our reanalysis of existing datasets, 8 of them regulated only in the absence of SCF. Matched filter analysis identified 2 regulated genes not expressed in previous erythroid expression studies4-13,24 : gasdermin B (GSDMB), which was present only in Late-E and zinc finger protein 805 (ZNF805). An additional 13 genes in MFUp were not previously differentially expressed. Table 2 lists the functions of some of the genes that have not previously been described as differentially expressed in primary erythroblasts, and Figure 3C shows quantitative PCR confirmation of expression profiles of selected genes in MFUp.
Consistently Highly Expressed transcripts
A list of genes that are Consistently Highly Expressed (CHE) throughout erythroblast maturation was generated by considering those PIDs with a minimum expression value of at least 1000 for which all B values between the possible pairwise comparisons were less than −2.945, indicating a less than 5% probability that the transcript was differentially regulated at any stage (CHE, supplemental Table 6). Any genes represented by divergent PIDs whose expression varied so much as to be included in both DE1 and CHE were removed from both lists. The final list of significant and continuously expressed transcripts contained 185 genes. Initial quantitative PCR of splicing factor arginine-serine rich (SFRS2) and myosin light chain 12B (MYL12B; Figure 3D) indicates that, although neither of these transcripts has appeared to be differentially expressed during erythropoiesis in previously published datasets, their expression may vary more throughout erythroblast maturation than is indicated by the microarray data. This emphasizes the need for verification of microarray data by alternative means. Expression of ATP synthase, H+ transporting, mitochondrial F0 complex, subunit G (ATP5L), eukaryotic translation elongation factor 1 alpha 1 (EEF1A1), and ribosomal protein S19 (RPS19) was confirmed by quantitative PCR to be at a consistently high level, changing by less than 2-fold (Figure 3D). Therefore, this set of genes (CHE) is a valid starting point for studying continuously expressed genes with significant expression levels throughout maturation. Our CHE list may be further refined by removing the 44 genes that were differentially regulated in previous studies,4-13 although 28 of these were differentially expressed only in the absence of SCF.
Ontology analysis of significant gene lists
Ingenuity Pathways Analysis and DAVID23 were used to analyze the molecular and cellular functions enriched in these gene lists. Genes affecting cell cycle were significantly represented in all 3 categories: genes controlling apoptosis and gene expression were significantly represented in terminally up-regulated genes, and genes controlling cellular assembly and organization were significantly represented in terminally down-regulated genes (supplemental Figure 4). Protein synthesis was significantly associated with transcripts in both DE_SigUp and CHE gene lists but not with DE_SigDown, in keeping with the role of the red cell as a hemoglobin factory. Similarly, genes associated with RNA posttranscriptional modification were significantly represented in the CHE and DE_SigDown gene lists but not with DE_SigUp.
TFBS analysis
TFBS enriched in the lists CHE, DE_SigUp, DE_SigDown, and MFUp were analyzed using DiRE, Pscan, and CONFAC20-22 software (Figure 5; Tables 3–4). All 3 packages query the TRANSFAC TFBS database. Pscan also queries the JASPAR database.
TFBSs significantly associated with CHE and DE_SigUP. (A) Transcription factor analysis of genes that are consistently highly expressed during erythroid maturation (CHE). Significant TFBS from Pscan output querying the JASPAR and TRANSFAC databases are shown. Green bars indicate bZIP TFBSp; red bars, ETS factor TFBS; solid bars, the TFBS was detected as significant by more than one of the 3 programs Pscan, DiRE, and CONFAC; and hashed bars, the TFBS was detected as significant by Pscan only. (B) Transcription factor analysis of genes that are significantly up-regulated during erythroid maturation (DE_SigUp). (i) Significant TFBS from Pscan output querying the JASPAR and TRANSFAC databases. (ii) Significant TFBS from DiRE output (only TFBS, which are also present in Pscan or CONFAC output or in bZIP factor family returned by Pscan search against JASPARFam database are shown). (iii) Significant TFBS from CONFAC output returned with a mean difference of the number of TFBS for each promoter between sample and control greater than 0.5 (only TFBS, which are also present in Pscan or DiRE outputs or in bZIP or ETS factor families returned by Pscan search against JASPARFam database are shown). Green bars represent bZIP TFBS; solid bars, the TFBS was detected as significant by more than one of the 3 programs Pscan, DiRE and CONFAC; and hashed bars, the TFBS was detected as significant by a single program only.
TFBSs significantly associated with CHE and DE_SigUP. (A) Transcription factor analysis of genes that are consistently highly expressed during erythroid maturation (CHE). Significant TFBS from Pscan output querying the JASPAR and TRANSFAC databases are shown. Green bars indicate bZIP TFBSp; red bars, ETS factor TFBS; solid bars, the TFBS was detected as significant by more than one of the 3 programs Pscan, DiRE, and CONFAC; and hashed bars, the TFBS was detected as significant by Pscan only. (B) Transcription factor analysis of genes that are significantly up-regulated during erythroid maturation (DE_SigUp). (i) Significant TFBS from Pscan output querying the JASPAR and TRANSFAC databases. (ii) Significant TFBS from DiRE output (only TFBS, which are also present in Pscan or CONFAC output or in bZIP factor family returned by Pscan search against JASPARFam database are shown). (iii) Significant TFBS from CONFAC output returned with a mean difference of the number of TFBS for each promoter between sample and control greater than 0.5 (only TFBS, which are also present in Pscan or DiRE outputs or in bZIP or ETS factor families returned by Pscan search against JASPARFam database are shown). Green bars represent bZIP TFBS; solid bars, the TFBS was detected as significant by more than one of the 3 programs Pscan, DiRE and CONFAC; and hashed bars, the TFBS was detected as significant by a single program only.
Two transcription factor classes significantly represented in genes in CHE and DE_SigUp
| TFBS . | Bonferroni P in CHE . | Bonferroni P in DE_SigUp . | Information content . |
|---|---|---|---|
| CREB/G-box-like bZIP | 5.85E-08 | 8.00E-03 | 10.79 |
| ETS family | 1.31E-08 | NS | 8.99 |
| TFBS . | Bonferroni P in CHE . | Bonferroni P in DE_SigUp . | Information content . |
|---|---|---|---|
| CREB/G-box-like bZIP | 5.85E-08 | 8.00E-03 | 10.79 |
| ETS family | 1.31E-08 | NS | 8.99 |
Bonferroni P value and information content returned by Pscan are given.
The CHE and DE_SigUp gene lists were both significantly enriched for binding sites of the CREB/G-box-like subclass of bZIP transcription factors (Figure 5; Table 3), and CHE was also significantly enriched for ETS class TFBS. Both classes are highly conserved and mediate diverse effects through many partners.29,30 As might be expected for genes that are switched off as cells become more specialized, fewer TFBS were found to be significantly enriched in the DE_SigDown list.
Discussion
Erythroid maturation is a gradual, continuous process, erythroblasts at different stages of maturation simultaneously coexisting in the bone marrow. CFU-E, Pro-E, Int-E, and Late-E stages represent snapshots of the maturing erythroblast, providing a tool to study gene expression during erythropoiesis. We have demonstrated that cells sorted with defined phenotypes express not only significantly fewer genes than unsorted cells but also show significantly more differences of greater magnitude in the expression levels of particular genes between stages than unsorted cells. We have also identified many differentially expressed genes whose regulation has not been previously described in maturing primary erythroblasts. Finally, using TFBS analysis, we have identified potential transcription factors that may regulate gene expression during terminal erythropoiesis.
The cRNA hybridized to the Affymetrix arrays was a pool of preparations from 4 morphologically and phenotypically homogeneous cell populations possessing a restricted range of expression of the erythroid markers CD36, CD71, and CD235a, therefore representing enriched erythroblast populations with the lowest possible level of contamination from other hematopoietic lineages.
Hybridizing pools of RNA from such rigorously defined populations clearly demonstrates significant increases in magnitude of expression ratios of differentially expressed transcripts in RNA from sorted erythroblasts compared with that from unsorted cells from the 2-stage cultures (Figure 4E). Removal of contaminating nonerythroid cells and enrichment of erythroblasts at a similar stage of maturation enables improved detection of expression ratios over a broader dynamic range. Reanalysis of other datasets from studies with a different emphasis9-12 indicates that a smaller proportion of expressed genes were differentially expressed in nonsorted erythroblasts (Table 1), with the exception of erythroblasts cultured in the absence of SCF,12 which would be expected to exhibit enhanced differentiation because SCF promotes erythroid expansion and delays differentiation.28 Furthermore, the robustness and integrity of our data are increased by the pooling strategy we used when preparing cRNA,31 as confirmed by quantitative PCR of individual samples.
Genes most highly expressed and up-regulated during erythroid maturation include ribosomal and mitochondrial genes, those involved in transcription and translation, and other known erythroid genes, such as globins and blood group antigens.
In our sorted erythroblasts, EPO-dependent transcripts previously identified using a cDNA subtraction library,32 including histones, high mobility group factors and transcription factors involved in chromatin remodeling, and DNA recombination and repair proteins, such as topoisomerases, are mostly highly expressed.
Keller et al9 cultured CD34+ cells in a serum-free culture system containing EPO and harvested the unsorted cells at 6 time points between 1 and 11 days in culture, representing differentiation from cells more immature than CFU-E to hemoglobinized cells appearing slightly less mature than our pyknotic Late-Es. They identified 1500 differentially expressed genes, approximately 20% of which are in DE1, our least stringent list of 3678 differentially regulated transcripts, whereas 93 genes are in our most stringent differentially regulated list DE_Sig, which contains 327 genes. Many of the discrepancies between these sets of differentially regulated genes may be the result of the differences in maturity of the erythroblasts studied,9 or to specific culture conditions. Our dataset includes 1151 differentially regulated genes, which were not differentially regulated during erythropoiesis in previous studies4-13 plus a further 1096, which were differentially expressed only when erythroblasts were cultured in the absence of SCF, conditions that would enhance erythroid differentiation.28 This is probably because we defined populations by expression of CD36, CD71, and CD235a, resulting in a wider dynamic range of detectable fold-change ratios (Figure 4E) and a lower proportion of transcripts apparently present at all stages of maturation. The significance of expression profiles of specific genes will become apparent as more expression data and, in particular, functional studies become available.
Some of the significantly regulated genes we have identified, such as SMARCA4, which coactivates the γ-globin gene, have known erythroid functions (Table 2; supplemental Table 8A). The erythropoietic role of others, known to have relevant functions, such as inhibition of apoptosis33,34 or proliferation35 in other cell types, must be confirmed by functional studies. Most of the significantly differentially expressed genes identified in our dataset were terminally differentially regulated (Figure 4A-D) and therefore probably beyond the scope of previous studies except for the study omitting SCF from the culture medium.12 An example of a novel erythroid expression profile identified by our study is that of the inhibitor of apoptosis protein gene baculoviral IAP repeat-containing 3 (BIRC3), which blocks caspase-induced apoptosis by direct inhibition of caspases 3 and 7.36 Previously observed to be down-regulated during the earliest stages of erythropoiesis and then remain expressed at a low level,9 BIRC3 is in DE_SigUp, significantly up-regulated between Int-Es and Late-Es, and uniquely and substantially expressed in Late-Es. This expression pattern was confirmed by quantitative PCR (Figure 3A). This difference is perhaps because the induction we observe is in homogeneous erythroblasts more mature than those in previous studies.4-13
RFXAP, significantly up-regulated in our Int-Es as confirmed by quantitative PCR (Figure 3B), was not differentially regulated during erythropoiesis in previous studies,9-11,13 although our retrospective analysis of previous data shows that it was up-regulated during maturation of erythroblasts cultured in the absence of SCF.12 The transcription factor RFXAP, in association with NF-Y and CREB, is involved in major histocompatibility complex class II activation.37 Binding sites for both CREB and NF-Y are enriched in promoters of genes in DE_Sig and also of genes expressed at a consistently high level in our maturing erythroblasts (CHE, Figure 5; Table 4). RFXAP regulation during erythroblast maturation suggests that it may have a novel role in erythropoiesis. Activating transcription factor 3 (ATF3), a CREB transcription factor, is also significantly up-regulated and was one of only 20 genes in DE_Sig, which is consistently significantly up-regulated between all pairs of stages studied except for CFU-E to Pro-E. ATF3 is involved in apoptosis, growth, commitment, proliferation, cell cycle progression, and morphology and represses the erythroid-derived 2–related factor 2 (Nrf2)–regulated stress response pathway.42 Erythropoiesis-related functions of selected transcription factors highlighted in our TFBS analysis of CHE and DE_SigUp, many of which are consistent with enriched molecular function analysis results (supplemental Figure 4), are listed in Table 4.
Selected transcription factors significantly represented in genes in CHE and DE_SigUp with examples of functions
| TF . | TFBS enriched in . | Erythroid-related functions . |
|---|---|---|
| CREB | CHE, DE_SigUp | Key mediator of myeloid differentiation and proliferation |
| AP2/ERF | CHE | Transcriptional repressor, induces erythroid differentiation in K562 and HEL cells30 |
| E2F | CHE, DE_SigUp | E2F plays critical roles in cell cycle progression by regulating the expression of genes involved in nucleotide synthesis, DNA replication, and cell cycle control as demonstrated in mouse models.38 E2F has recently been shown to form a complex with GATA1, FOG1, and retinoblastoma protein (pRb), thus stalling cell proliferation and steering erythroid precursors toward terminal differentiation.39 |
| NFY(A) (CAAT-binding protein) | CHE, DE_SigUp | Binds human β-globin promoter in K562 cells, stabilizing a complex formed with MZF1 and GATA2. Appears important in maintaining a chromatin conformation favorable to β-globin transcription.40 |
| SP1 | CHE, DE_SigUp | Ubiquitously expressed. Acts synergistically with FOXM1, GATA1,27 OCT, and AP2, which are also highlighted in DiRE analysis of DE_SigUp. |
| TATA/TBP | DE_SigUp | TBP is essential for long-range activation of β-globin transcription in response to the HS2 enhancer located 10 kb from the promoter.41 |
| TF . | TFBS enriched in . | Erythroid-related functions . |
|---|---|---|
| CREB | CHE, DE_SigUp | Key mediator of myeloid differentiation and proliferation |
| AP2/ERF | CHE | Transcriptional repressor, induces erythroid differentiation in K562 and HEL cells30 |
| E2F | CHE, DE_SigUp | E2F plays critical roles in cell cycle progression by regulating the expression of genes involved in nucleotide synthesis, DNA replication, and cell cycle control as demonstrated in mouse models.38 E2F has recently been shown to form a complex with GATA1, FOG1, and retinoblastoma protein (pRb), thus stalling cell proliferation and steering erythroid precursors toward terminal differentiation.39 |
| NFY(A) (CAAT-binding protein) | CHE, DE_SigUp | Binds human β-globin promoter in K562 cells, stabilizing a complex formed with MZF1 and GATA2. Appears important in maintaining a chromatin conformation favorable to β-globin transcription.40 |
| SP1 | CHE, DE_SigUp | Ubiquitously expressed. Acts synergistically with FOXM1, GATA1,27 OCT, and AP2, which are also highlighted in DiRE analysis of DE_SigUp. |
| TATA/TBP | DE_SigUp | TBP is essential for long-range activation of β-globin transcription in response to the HS2 enhancer located 10 kb from the promoter.41 |
The 8570 genes present in the sorted erythroid cells contain 496 genes whose expression has been reported to be specific to nonerythroid hematopoietic lineages.24 This HaemAtlas study compared erythroblasts at a stage between our Pro-E and Int-E stages43 with cells from other hematopoietic lineages, and lists lineage-specific transcripts. However, in that study, the definition of lineage-specific transcripts required at least 2-fold up-regulation in the respective lineage.24 Therefore, some erythroid transcripts may have been present at a higher level in nonerythroid lineages or absent at the specific time point studied. Erythroid expression of many of these 496 transcripts we have detected has also been reported elsewhere, including the HaemAtlas study itself4-13,32,44 (supplemental Table 7). None of the markers used to select the nonerythroid lineages24 are present in our dataset, with the exception of Integrin α2B (ITGA2B, the platelet antigen CD41), which reached an average expression level of 350 in CFU-E, and was also present in erythroblasts in the HaemAtlas study.24
Of the 127 genes present only in CFU-E with an expression cut-off of 100 (Figure 2A), only 6 exceeded a minimum expression level between the 3 CFU-E pools of 200. None reached an expression level of 500. Genes in this category are probably to be those which are more highly expressed during earlier developmental stages, being enriched for Hematologic System Development and Function (24 associated transcripts, P = 1.55 × 10−5; Figure 2C). These include Ets variant 6 (ETV6), which has a role in expanding erythroid precursors and accumulating hemoglobin,45 and hematopoietically expressed homeobox (HHEX) which regulates proliferation and differentiation of hemangioblasts and endothelial cells during ES cell differentiation.46 The mouse homologue, Hhex, was rapidly down-regulated when erythroid differentiation was induced by GATA1 in G1ER cells.7 The most significant molecular and cellular function associated with this CFU-E-specific gene list is cellular growth and proliferation (29 transcripts associated with this function, P = 1.55 × 10−5), which is entirely consistent with the role of HHEX and ETV6 in CFU-E maturation. Nineteen genes present only in CFU-E, including ETV6, have previously been reported to be down-regulated during early erythropoiesis,9 indicating that they were up-regulated at earlier stages of erythroid development.
Of the 511 genes present only in Late-Es with an expression cut-off of 100 (Figure 2A), 7 exceed a minimum expression level of 1000. Four of these genes are associated with apoptosis: tribbles homolog 3 (TRIB3), heme oxygenase 1 (HMOX1), BIRC3, and B-cell CLL/lymphoma 6 (BCL6). The molecular and cellular function Cell Death is significantly enriched in genes specifically expressed only in Late-Es (78 associated transcripts, P = 3.91 × 10−4; supplemental Figure 2A). Genes uniquely expressed in Late-Es are associated with a role in gene expression (77 associated transcripts, P = 5.09 × 10−5; supplemental Figure 2A).
The 1144 genes exceeding a minimum expression level of 500 at all stages (Figure 2B) are significantly enriched for the molecular and cellular functions “protein synthesis” (127 associated transcripts, P = 4.83 × 10−34), “RNA posttranscriptional modification” (83 associated transcripts, 5.58 × 10−17), “posttranslational modification” (84 associated transcripts, P = 1.1 × 10−8), and “protein folding” (21 associated transcripts, P = 1.1 × 10−8). This is compatible with terminal differentiation and the high level of globin synthesis required.
Of 123 genes within 60 kb of SNPs found to be associated with blood cell traits,47,48 80 genes were expressed at any stage throughout erythropoiesis, 20 were differentially expressed at the least stringent level in DE1, and 3 at the most stringent level in DE_Sig: membrane-associated ring finger (C3HC4) 8 (MARCH8), myeloblastosis oncogene (MYB), and transmembrane and coiled-coil domain family 2 (TMCC2). The CHE gene list contained sialomucin CD164, TFRC, and protein tyrosine phosphatase, nonreceptor type 11 (PTPN11).
We have begun to explore different approaches of identifying coregulated genes, for example, consistently up-regulated genes. In the broad framework of signal processing, it may be possible to design a matched filter to describe such a trend and to capture the effects of the relevant biochemical processes. In reality, however, it is much more complicated, in that not all up-regulated genes will experience equal up-regulation. Comparison of the successively up-regulated genes identified by matched filtering (MFUp) demonstrated the potential danger of missing important genes if the wrong type of analysis is used. Although the matched filtering approach identified 47 consistently up-regulated genes in our dataset, which were also identified as significantly differentially regulated when selected on the basis of their B values (DE_Sig), and 98 genes whose erythroid expression has been previously reported, several new differentially expressed genes were also identified by this approach, with the expression profiles of BRWD3, CTSL2, DNAJB4, GSDMB, LIN7B, RRAD, RFFL, and ZNF805 confirmed by quantitative PCR (Figure 3C). These 8 genes are expressed at modest levels compared with many of those already described in erythroid cells, and future functional studies will be necessary to determine their role in erythropoiesis. However, many of them have known roles in autophagy, apoptosis, and transcriptional regulation49-56 (Table 2; supplemental Table 8A), which may be relevant to the maturing erythroblast. Importantly, this approach enabled selection of genes based on their CFU-E to Pro-E expression ratios; to do this using the LIMMA approach would have required lowering the selection stringency of B and/or P values. Future studies will consider that not all up-regulated genes experience equal up-regulation, and extend the design of adaptive matched filters57 to describe a trend. Beyond the novelty of matched filtering, we intend to continue to explore such new approaches (eg, adaptive matched filtering) that have general application to microarray biologic datasets.
Microarray technology is less sensitive than quantitative PCR at quantifying expression levels, with a more restricted dynamic range. Signal intensity may be affected by location and hybridization efficiency of probes, yet arrays have the obvious advantages of large-scale analysis of gene expression. Deep sequencing will supersede microarray hybridizations as the optimum method for large-scale transcriptome analysis. However, our dataset represents a novel and valuable resource to explore gene expression changes during erythroid maturation and to compare with expression data from the diseased state.
We studied enriched erythroblast populations with the lowest possible level of contamination from other hematopoietic lineages to show that cells sorted with defined phenotypes express significantly fewer genes than unsorted cells and show significantly more differences of greater magnitude in the expression levels of particular genes between stages. We have also identified many differentially expressed genes previously unregulated in erythroid expression studies with unknown roles in erythropoiesis and highlighted potential transcription factors that may regulate gene expression during terminal erythropoiesis.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
The authors thank Chris Fisher, Jim Hughes, Abigail Lamikanra, Nick Watkins, and David Weatherall for helpful discussion; Tariq Enver for support; Helen Cattan and Emanuele Marchi for bioinformatics advice; Jackie Sloane-Stanley, Sue Butler, and Jill Brown for technical assistance; and Giulio Pavesi and Carlos Moreno for email correspondence regarding Pscan and CONFAC, respectively.
This work was supported in part by National Health Service Blood and Transplant, National Institute for Health Research Biomedical Research Center Program, and by National Institute for Health Research (Program Grants for Applied Research scheme RP-PG-0310-1004).
The Human Erythroblast Maturation (HEM) Database is available at: https://cellline.molbiol.ox.ac.uk/eryth/index.html.
The views expressed in this publication are those of the authors and not necessarily those of the National Health Service, the National Institute for Health Research or the Department of Health.
Authorship
Contribution: A.T.M.-C. and K.J.H.R. designed and performed research, analyzed data, and wrote the paper; A.A. performed research and analyzed data; S.S., N.G., and S.T. analyzed data; K.C. and C.W. performed research; S.J.M. and V.J.B. analyzed data and wrote the paper; A.K.N. analyzed data and wrote the paper; W.G.W. and D.R.H. designed research; and D.J.R. wrote the paper.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
The current affiliation of A.A. is Flow Cytometry Laboratory, Institute of Molecular Medicine, Trinity College, Dublin, Ireland.
Correspondence: Alison T. Merryweather-Clarke, Blood Research Laboratory, Nuffield Department of Clinical Laboratory Sciences, Level 4, John Radcliffe Hospital, Headington, Oxford, OX3 9DU, United Kingdom; e-mail: alison.merryweather-clarke@ndcls.ox.ac.uk.