The p21 is a downstream effector of p53/p73 and belongs to the CIP/KIP family of cyclin-dependent kinase inhibitors (CDKIs). It is, therefore, a potential tumor suppressor gene and probably plays an important role in tumor development. Moreover, reduced expression of p21 has been reported to have prognostic value in several human malignancies. In contrast with other CDKIs, mutational inactivation of p21 is infrequent, but gene inactivation by an alternative mechanism seems to be the general pathway. In this study, we analyzed the methylation status of the p21 promoter region using semiquantitative polymerase chain reaction in 124 patients with acute lymphoblastic leukemia (ALL). We observed p21 hypermethylation in bone marrow cells from 41% (51 of 124) of ALL patients. Hypermethylation within promoter strongly correlated with decreased p21 messenger RNA expression in tumoral cells. Clinical, molecular, and laboratory features and complete remission rate did not differ significantly between hypermethylated and normally methylated patients. Estimated disease-free survival (DFS) and overall survival at 7 and 9 years, respectively, were 59% and 65% for healthy patients and 6% and 8% for hypermethylated patients (P = .00001 andP = .006). Multivariate analysis of potential prognostic factors demonstrated that p21 methylation status was an independent prognostic factor in predicting DFS (P = .0001). Our results indicate that the p21 gene is subject to methylation regulation at the transcription level in ALL and seems to be an important factor in predicting the clinical outcome of these patients.
Introduction
Mitogen-dependent progression through the first gap phase (G1) and initiation of DNA synthesis (S phase) during the mammalian cell division cycle are cooperatively regulated by several classes of cyclin-dependent kinases (CDKs), the activities of which are, in turn, constrained by CDK inhibitors (CDKIs). CDKIs that govern these events have been assigned to 1 of 2 families on the basis of their structure and CDK target. The first class includes the INK4 proteins, so named for their ability to specifically inhibit the catalytic subunits of CDK4 and CDK6. This family includes p16INK4a, p15INK4b, p18INK4c, and p19INK4d. The INK4 proteins can be contrasted with more broadly acting inhibitors of the CIP/KIP family, whose actions affect the activities of cyclin D–, E– and A–dependent kinases. The latter class includes p21CIP1/WAF1/SDI1 (p21), p27KIP1, and p57KIP2.1 Increased expression of CDKIs has been recognized as a general mechanism for cell cycle arrest, and CDKIs have also been considered to be implicated in tumorigenesis as possible tumor-suppressor genes.2 3
The p21 is a strong candidate for participation in tumor progression. Expression of p21 is induced by wild-type p53 in the presence of DNA damage, leading to apoptosis or cell cycle arrest at the G1checkpoint.4 The p21 is also involved in replicative senescence and terminal differentiation and proliferation in nonhemopoietic and hemopoietic cells.5 The potential role of p21 in oncogenesis has been postulated on the basis of its transcriptional control by p53. In addition, overexpression of p21 may suppress tumor growth in different experimental models.6 7
One characteristic of cancer cells is a failure to undergo growth arrest and differentiation. Acute lymphoblastic leukemia (ALL) is the most prevalent type of cancer, as well as the most common form of leukemia in children.8 Leukemic cells from the majority of patients with ALL express on their surface a variety of protein antigens that are found at discrete stages of maturation on normal B- or T-lymphocyte precursors. Thus, leukemic clones from ALL patients are thought to originate from any lymphoid cell blocked at an early stage of development.9 Human immature leukemic cell lines of both myeloid and lymphoid lineages frequently show lack of p21 expression. Induction of apoptosis in this setting is accompanied by cell cycle arrest and by accumulation and stabilization of p21. These observations suggest that p21 plays an inhibitory role in the proliferation of ALL cells and that the suppression of p21 expression may be advantageous for leukemic growth.10-12
Changes in DNA methylation patterns are known to occur during tumorigenesis.13 CpG islands near promoters and the 5′ regulatory region are usually unmethylated in normal somatic cells. In contrast, widespread methylation of CpG islands occurs on autosomal genes and leads to the silencing of the gene during oncogenic transformation. In hematologic malignancies, aberrant DNA hypermethylation is thought to play a role in leukemogenesis. For example, during the progression of chronic myeloid leukemia, the ABL1 promoter of the BCR-ABL fusion gene becomes significantly hypermethylated.14 Also, aberrant hypermethylation of the p15INK4b tumor-suppressor gene is associated with its inactivation in at least 50% of patients with acute leukemia.15 Furthermore, hypermethylation of p15INK4b is concomitant with disease progression in myelodysplastic syndromes.16 Previous studies have shown that, although decreased p21 expression is often detected in cancer cells and correlated with disease progression, including lymphoid malignancies,17,18 mutation and allelic loss of the p21 gene are rarely detected.19,20 In light of our recent finding that altered DNA methylation in ALL is associated with decreased expression of some KIP family proteins (ie, p57KIP2),21 we hypothesized that the abnormal methylation in the p21 promoter region might play a role in the down-regulation of this gene in ALL. We now show that hypermethylation of the p21 gene is a frequent event in ALL and correlates with decreased p21 expression. These findings are associated with an unfavorable clinical outcome.
Materials and methods
Patients
We studied 124 consecutive patients (85 male; 39 female) who were diagnosed with de novo ALL between January 1990 and December 2000. The median age at diagnosis in the study population as a whole was 14 years (range, 0.5-82 years). Of these patients, 68 were children (median age, 5 years; range, 0.5-14 years), and 56 presented adult ALL (median age, 29 years; range, 15-82 years). Informed consent was obtained from the patient or the patient's guardians. Diagnosis was established according to standard morphologic and cytochemical criteria. Patients were studied at the time of initial diagnosis; were risk-stratified according to the therapeutic protocol used, which was always based on recognized prognostic features (including cytogenetics); and were entered in ALL protocols of the “Programa para el estudio y tratamiento de las hemopatias malignas” (PETHEMA) Spanish study group. For statistical analyses, children were also grouped according to the National Cancer Institute (NCI) risk-classification criteria.22 Mononuclear cells, obtained from bone marrow samples by centrifugation on Ficoll-Hypaque density gradient, were used for immunophenotypic and DNA/RNA analyses. The specific PETHEMA ALL treatment protocols in which these patients entered included ALL-89 (between 1990 and 1994; n = 51) and ALL-93 (between 1994 and 2000; n = 73). The design and results of these studies have been previously reported.23-26 Fifty patients relapsed. Thirty patients received stem cell transplantation (10 autologous, 20 allogeneic) in the first (n = 14) or second complete remission (CR) (n = 16). There are 73 patients currently alive. Clinical characteristics of the patients are listed in Table1.
Clinical characteristics and outcome of 124 patients according to p21 gene methylation status
| Characteristic . | Hypomethylated, % (n = 73) . | Hypermethylated, % (n = 51) . | P . |
|---|---|---|---|
| Age | NS | ||
| Younger than 15 y | 56 | 53 | |
| Older than 15 y | 44 | 47 | |
| Sex, M/F | 70/30 | 67/33 | NS |
| WBC | NS | ||
| Below 50 × 109/L | 80 | 75 | |
| Above 50 × 109/L | 20 | 25 | |
| FAB classification | NS | ||
| L1 | 55 | 56 | |
| L2 | 37 | 40 | |
| L3 | 8 | 4 | |
| Blast lineage | NS | ||
| B | 84 | 80 | |
| T | 16 | 20 | |
| NCI risk groups | NS | ||
| Standard | 68 | 63 | |
| Poor | 32 | 37 | |
| PETHEMA risk groups | NS | ||
| Standard | 47 | 40 | |
| Poor | 53 | 60 | |
| Treatment | NS | ||
| PETHEMA 89 | 42 | 40 | |
| PETHEMA 93 | 58 | 60 | |
| BMT | 22 | 27 | NS |
| Best response | NS | ||
| CR | 89 | 90 | |
| Cytogenetic/molecular abnormalities | NS | ||
| BCR-ABL | 14 | 25 | |
| t(1;19) | 4 | 2 | |
| 11q23 | 4 | 2 | |
| c-Myc | 5 | 8 | |
| 7q35-14q11 | 3 | 8 | |
| Hyperdiploidy | 9 | 7 | |
| TEL-AML1 | 9 | 6 | |
| Normal | 42 | 39 | |
| Others | 5 | 3 | |
| NT | 5 | 0 | |
| Relapse | 26 | 61 | < .0001 |
| Death | 30 | 57 | .005 |
| Characteristic . | Hypomethylated, % (n = 73) . | Hypermethylated, % (n = 51) . | P . |
|---|---|---|---|
| Age | NS | ||
| Younger than 15 y | 56 | 53 | |
| Older than 15 y | 44 | 47 | |
| Sex, M/F | 70/30 | 67/33 | NS |
| WBC | NS | ||
| Below 50 × 109/L | 80 | 75 | |
| Above 50 × 109/L | 20 | 25 | |
| FAB classification | NS | ||
| L1 | 55 | 56 | |
| L2 | 37 | 40 | |
| L3 | 8 | 4 | |
| Blast lineage | NS | ||
| B | 84 | 80 | |
| T | 16 | 20 | |
| NCI risk groups | NS | ||
| Standard | 68 | 63 | |
| Poor | 32 | 37 | |
| PETHEMA risk groups | NS | ||
| Standard | 47 | 40 | |
| Poor | 53 | 60 | |
| Treatment | NS | ||
| PETHEMA 89 | 42 | 40 | |
| PETHEMA 93 | 58 | 60 | |
| BMT | 22 | 27 | NS |
| Best response | NS | ||
| CR | 89 | 90 | |
| Cytogenetic/molecular abnormalities | NS | ||
| BCR-ABL | 14 | 25 | |
| t(1;19) | 4 | 2 | |
| 11q23 | 4 | 2 | |
| c-Myc | 5 | 8 | |
| 7q35-14q11 | 3 | 8 | |
| Hyperdiploidy | 9 | 7 | |
| TEL-AML1 | 9 | 6 | |
| Normal | 42 | 39 | |
| Others | 5 | 3 | |
| NT | 5 | 0 | |
| Relapse | 26 | 61 | < .0001 |
| Death | 30 | 57 | .005 |
WBC indicates white blood count; FAB, French-American-British; NCI, National Cancer Institute; PETHEMA, Programa para el estudio y tratamiento de las hemopatias malignas; BMT, bone marrow transplantation; CR, complete remission.
Immunophenotyping
Cell surface antigens were detected by direct immunofluorescence technique in flow cytometric analysis by means of a panel of monoclonal antibodies labeled with fluorescein isothiocyanate or phycoerythrin reactive with lymphoid and myeloid antigens (CD1, CD2, CD3, CD4, CD5, CD7, CD8, CD10, CD19, CD22, CD13, CD14, CD33, CD34, and HLA-DR) (Becton Dickinson, San Jose, CA). When myeloid antigens were expressed in at least 20% of cells, dual-fluorescence analysis was performed by means of phycoerythrin-conjugated monoclonal antibodies against lymphoid antigens to confirm the coexpression of both antigens on the same cells. Data acquisition and analysis were performed by means of a Coulter Profile II (Hialeah, FL). Analysis was restricted to the leukemic population by gating on forward and side light-scatter parameters. For intracytoplasmatic detection of myeloperoxidase, CD22, CD3, and deoxynucleotidyl transferase (TdT), an immunocytochemical staining was performed by means of the alkaline phosphatase–antialkaline phosphatase technique and specific monoclonal antibodies MPO-7 (Dako, Glostrup, Denmark), CD22, CD3 (Becton Dickinson), and TdT (Immunotech, Marseilles, France).
Cytogenetics
For cytogenetic analyses, cells from bone marrow were cultured in McCoy medium supplemented with 20% fetal calf serum and antibiotics for 48 hours. Chromosome preparations were stained with 5% Giemsa solution according to standard procedures. A minimum of 25 metaphases were analyzed per sample. This allowed the detection of a minor clone at the 5% level. Successful cytogenetic analysis was performed in 120 patients (Table 1).
Nucleic acid isolation
High–molecular weight DNA and total RNA were prepared from mononuclear marrow cells at the time of diagnosis by means of conventional methods. As a control, marrow DNAs/RNAs were obtained from 30 healthy bone marrow donors (5 children and 25 adults).
Restriction digest and polymerase chain reaction procedure to detect p21 gene methylation
Polymerase chain reaction (PCR) analysis relying on the inability of HpaII to cut methylated sequences was used to analyze the p21 gene. The sites examined were 2 HpaII sites in the promoter of p21 (Figure 1A). Then, 5 μg genomic DNA was completely digested in 50 μL of total volume with 50 U HpaII, as recommended by the manufacturer (Promega, Madison, WI), and precipitated with ammonium acetate and ethanol. The pellet obtained was washed twice with 70% ethanol and dissolved in 50 μL H2O. Next, 5 μL (approximately 0.5 μg) of this solution was withdrawn and pipetted into 45 μL PCR mixture: 200 μM each deoxynucleoside 5′-triphosphate; 10 mM Tris-HCl (pH 8.3); 1.5 mM MgCl2; 50 mM KCl; 0.001% (wt/vol) gelatin; 5 UPfu DNA polymerase (Stratagene, La Jolla, CA); 50 pmol each primer p21-sense (5′-GCCTGCTGGAACTCGGCCAGGCTCAGCCTGC-3′) and p21-antisense (5′-GAGGCGACCCGCGCTCGGCCCAGCGCGCCG-3′). To control for intact DNA, a 75-bp fragment of the human β-actin gene without restriction sites of HpaII was coamplified as an internal standard (forward, 5′-CCGAGCGCGGCTACAGCTTCA-3′; reverse, 5′-CGTAGCACAGCTTCTCCTTAATGTC-3′).
Aberrant p21 methylation in acute lymphoblastic leukemia.
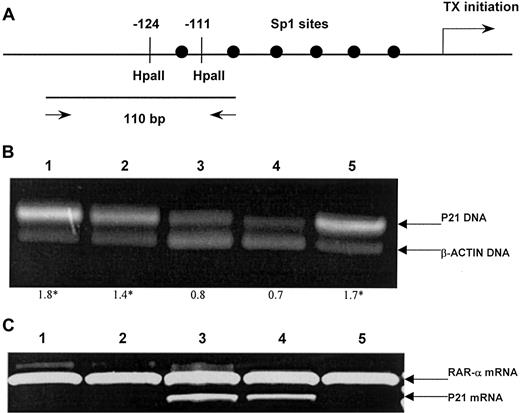
(A) A schematic map of the p21 promoter for restriction sites and the position of a primer set. Arrow indicates the location of the transcription initiation site of the p21 gene. Six Sp1-binding sites are located in a GC-rich region within 180 base pairs (bp) upstream of the transcription initiation site of the p21 gene. The −124 and −111 indicate the recognition sites of the restriction enzymeHpaII. Two arrows indicate a primer set for PCR and the size of PCR product, 110bp. (B) Methylation status of the p21 promoter region was detected by means of PCR of the HpaII-digested DNA. The upper band represents the 110-bp amplification product of the p21 gene, and the lower band, 75 bp of the β-actin gene. The optical density curves of both fragments were used to quantify the methylation level of p21 gene. The mean p21/β-actin ratio in normal bone marrow and the SE were used to assess p21 methylation state. Ratios above 3 SEs were considered hypermethylated. Lanes 1-5 show ALL samples. The p21–to–β-actin ratios are shown below the gel, with hypermethylated status indicated by asterisks. (C) Expression of p21 messenger RNA (mRNA) assessed by reverse-transcriptase (RT) PCR by means of primers for p21 and RARα (as control). Lanes 1, 2, and 5: hypermethylated ALL samples showing lack of p21 expression. Lanes 3, 4: normally methylated samples.
Aberrant p21 methylation in acute lymphoblastic leukemia.
(A) A schematic map of the p21 promoter for restriction sites and the position of a primer set. Arrow indicates the location of the transcription initiation site of the p21 gene. Six Sp1-binding sites are located in a GC-rich region within 180 base pairs (bp) upstream of the transcription initiation site of the p21 gene. The −124 and −111 indicate the recognition sites of the restriction enzymeHpaII. Two arrows indicate a primer set for PCR and the size of PCR product, 110bp. (B) Methylation status of the p21 promoter region was detected by means of PCR of the HpaII-digested DNA. The upper band represents the 110-bp amplification product of the p21 gene, and the lower band, 75 bp of the β-actin gene. The optical density curves of both fragments were used to quantify the methylation level of p21 gene. The mean p21/β-actin ratio in normal bone marrow and the SE were used to assess p21 methylation state. Ratios above 3 SEs were considered hypermethylated. Lanes 1-5 show ALL samples. The p21–to–β-actin ratios are shown below the gel, with hypermethylated status indicated by asterisks. (C) Expression of p21 messenger RNA (mRNA) assessed by reverse-transcriptase (RT) PCR by means of primers for p21 and RARα (as control). Lanes 1, 2, and 5: hypermethylated ALL samples showing lack of p21 expression. Lanes 3, 4: normally methylated samples.
The PCR was performed for 35 cycles under the following conditions: 98°C, 1 minute; 68°C, 1 minute; 74°C, 1 minute. Then, 20 μL PCR reaction was electrophoresed on a 3% agarose gel stained with ethidium bromide and photographed (Figure 1B). For semiquantitative analysis, gels were scanned by means of a Bio-Print gel scanner and the appropriate software as provided by the manufacturer (Vilber-Lourmat, Marne-La-Vallee, France). The the p21–to–β-actin ratio, the area under the curve of the p21 gene product divided by the area under the curve of the β-actin gene product, gives an indication of the methylation status of the respective sample. As previously described for normal tissues,27 analyses of bone marrow samples from 30 healthy marrow donors showed a partial methylation pattern at this CpG region. Therefore, the mean p21–to–β-actin ratio and the SE in the normal bone marrow were used to assess methylation status of the p21 promoter. Normalized values above 3 SEs were considered to be hypermethylated.
Reverse-transcription PCR analysis for p21 expression
Total RNA was isolated from 2 × 106 marrow cells by means of Ultraspec (Biotecx Laboratories, Houston, TX). Reverse transcription was performed on 1 μg total RNA, after heating at 70°C for 5 minutes, with random hexamers as reaction primer. The reaction was carried out at 42°C for 45 minutes in the presence of 12 U Avian Myeloblastosis Virus reverse transcriptase (Boehringer-Mannhein, Germany). Complementary DNA was amplified by means of a primer set that was specific for the p21 gene (sense, 5′-CAGGGGACAGCAGAGGAAGA-3′; antisense, 5′-GGGCGGCCAGGGTATGTAC-3′). The PCR reaction was performed as follows: 94°C for 7 minutes, 40 cycles of 94°C for 1 minute, 58°C for 1 minute, 72°C for 1 minute, and incubation at 72°C for 10 minutes. PCR products were resolved on 1.5% agarose gels. Amplification of RARα gene transcripts was performed to assess RNA integrity.28 A Bio-Print gel scanner was used to quantify the intensity of the p21 and the RARα bands, and the ratio was determined for each reaction. The mean p21-to-RARα ratio and the SE were used to assess levels of p21 in the tumors. Normalized values below 3 SEs were considered to represent decreased expression (Figure 1C).
Other molecular analyses
Standard Southern blot method was employed to detect immunoglobulin heavy-chain gene rearrangement and T-cell receptor β rearrangement by means of the commercially available JH DNA probe and CTβ probe (Oncogene Science, New York, NY). The 11q23 abnormalities were studied with the B859 probe kindly provided by Dr G. Cimino.29 The p210BCR-ABL and p190BCR-ABL transcripts were detected by means of RT-PCR according to the method used by Saglio et al.30
Statistical analysis
P values for comparisons of continuous variables between groups of patients were 2-tailed and based on the Wilcoxon rank sum test. P values for dichotomous variables were based on the Fisher exact test. The remaining P values were based on the Pearson chi-square test. Overall survival (OS) was measured from the day of diagnosis until death from any cause and was censored only for patients known to be alive at last contact. DFS was measured from the day that CR was established until either relapse or death without relapse, and it was censored only for patients who were alive without evidence of relapse at the last follow-up. Distributions of OS and DFS curves were estimated by the method of Kaplan and Meier, with 95% confidence intervals calculated by means of Greenwood's formula. Comparisons of OS or DFS between groups were based on the log-rank test. Comparisons adjusted for significant prognostic factors were based on Cox regression models and hazard regression models. All relapse and survival data were updated on April 30, 2001, and all follow-up data were censored at that point.
Results
Methylation state of the p21 promoter region
We first examined 30 normal bone marrow and 124 ALL samples for methylation alteration in a CpG-rich region within 180 bp upstream of the transcription initiation site of the p21 gene. The presence of methylation in the promoter of the p21 gene was tested by digestion with a methylation-sensitive restriction enzyme and a subsequent PCR reaction. The CpG-rich region in the 6 consecutive Sp1-binding sites of the p21 promoter (Figure 1A) was digested with methylation-sensitiveHpaII. Resulting DNA was subjected to a PCR reaction. Only undigested DNA will provide 110 bp of the PCR amplification product across the Sp1-binding sites (Figure 1B). Healthy individuals showed a weak band after PCR, indicating that this CpG was at least partially methylated. This result is not surprising because an incomplete methylation pattern has been previously reported in normal tissues (ie, lung, liver, and muscle).27 To better determine the degree of methylation of the p21 promoter, we established a semiquantitative analysis comparing the intensity of the methylation fragment relative to that obtained by amplifying a housekeeping gene (β-actin). The p21–to–β-actin ratios in normal bone marrow specimens ranged from 0.4 to 0.8 (mean, 0.7). Normalized values above 3 SEs were considered to be hypermethylated (ratio exceeding 1). With these principles, hypermethylation of the p21 promoter region was detected in 51 (41%) ALL patient samples.
Correlation between hypermethylation of the p21 promoter and the p21 mRNA expression in ALL
To determine whether the aberrant CpG methylation in the p21 promoter region correlates with decreased p21 expression in ALL, we analyzed levels of the p21 mRNA transcripts in the same set of tumor and normal marrow samples using a semiquantitative RT-PCR method. Levels of the p21 mRNA were normalized to an internal standard RARα, and decreased expression was defined as below 3 SEs of the mean expression level in the normal marrow specimens. All 51 hypermethylated ALL samples (100%) showed decreased p21 expression; in contrast, decreased p21 expression was detected in only 4 of the 73 ALL samples with normal methylation pattern (5%). These results indicated that CpG hypermethylation within the p21 promoter region strongly correlated with decreased constitutive expression of p21 in ALL cells.
Clinical presentation and p21 methylation
The p21 hypermethylation was detected at diagnosis in 41% (51 of 124) of patients with adult or childhood ALL of all the French-American-British subtypes. As shown in Table 1, aberrant p21 methylation was found more frequently in adults (24 of 56, 43%) than in children (27 of 68, 39%). However, this difference did not reach statistical significance. Clinical and laboratory characteristics did not differ significantly between hypermethylated and normal patients. Poor-risk cytogenetics or molecular events, risk groups according to both NCI and PETHEMA classifications, good-risk features (hyperdiploidy and TEL-AML1 fusion), type of PETHEMA protocol administered, and number of patients who received stem cell transplantation were similarly distributed between the 2 p21-methylation groups. Separate analysis of adult and childhood ALL patients gave the same results as the global series.
Clinical outcome and p21 methylation
Table 1 details the relapse history, CR rates, and mortality both for patients who exhibited p21 promoter hypermethylation and for patients with a normal p21 gene. CR rates of patients with the normal and the hypermethylated p21 gene were 89% (65 of 73) and 90% (46 of 51), respectively, accounting for 89% of the overall CR rate. This suggests that hypermethylation of the p21 promoter region did not correlate with response to remission induction therapy. However, hypermethylated patients had a higher relapse rate (61% versus 26%,P < .0001) and mortality rate (57% versus 30%,P = .005) than normally methylated patients. Similar results were obtained in the separate analyses of children (relapse rate, 48% versus 22%, P = .03; mortality rate, 37% versus 17%, P = .06) and adults (relapse rate, 85% versus 36%, P = .001; mortality rate, 79% versus 47%,P = .02).
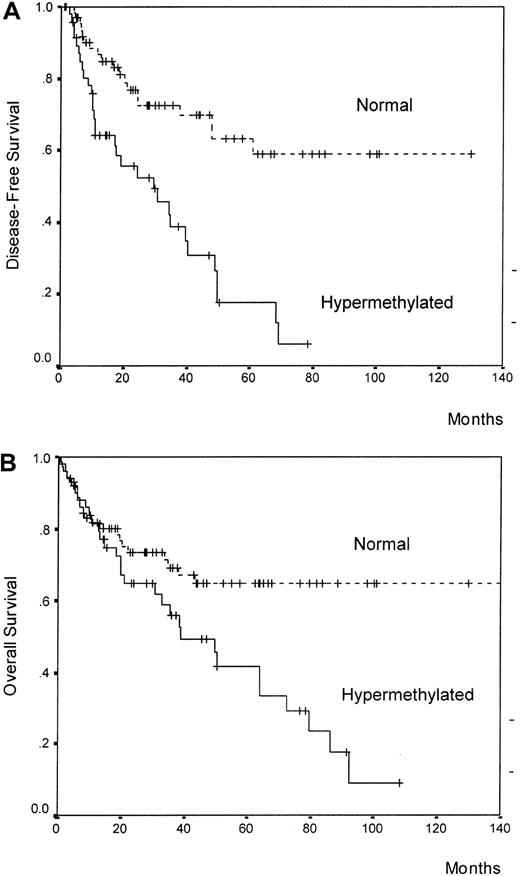
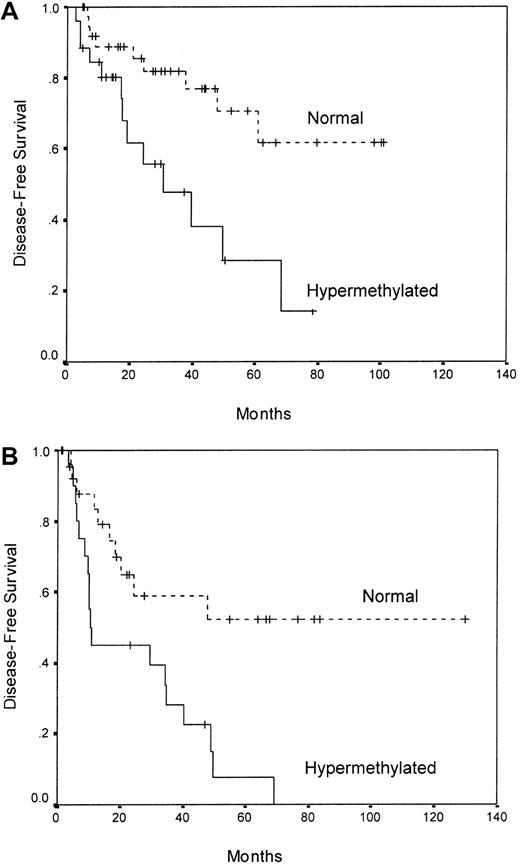
We analyzed the DFS among patients who achieved CR according to p21 methylation. Estimated DFS rates at 7 years were 59% and 6% for normal and methylated patients, respectively (P = .00001) (Figure 2A). Among normally methylated children, the 7-year DFS was 62%, contrasting with 14% for hypermethylated children (P = .006) (Figure3A). Among adult ALL patients, the 7-years DFS was 0% for methylated cases and 52% for normal cases (P = .003) (Figure 3B). The actuarial OS calculated for all leukemic patients was 65% and 8% at 9 years for patients with the normal and the hypermethylated p21 gene, respectively (P = .006) (Figure 2B). A trend toward significance was observed in the actuarial OS between normal and hypermethylated patients in the separate analyses of children (77% versus 31%,P = .06) and adults (50% versus 0%,P = .09).
Kaplan-Meier survivor function for all patients.
DFS (panel A) and OS (panel B) curves for all the patients enrolled in this study according to methylation status of the p21 promoter region.
Kaplan-Meier survivor function for all patients.
DFS (panel A) and OS (panel B) curves for all the patients enrolled in this study according to methylation status of the p21 promoter region.
Kaplan-Meier survivor function for ALL patients.
DFS curves for childhood (panel A) and adult (panel B) ALL patients based on p21 methylation status.
Kaplan-Meier survivor function for ALL patients.
DFS curves for childhood (panel A) and adult (panel B) ALL patients based on p21 methylation status.
A multivariate analysis of potential prognostic factors (including the type of PETHEMA protocol applied) demonstrated that hypermethylation of the p21 promoter region was an independent prognostic factor in predicting DFS in the global series as well as in both childhood and adult ALLs (Table 2).
Multivariate Cox model for disease-free survival
| Characteristic . | Univariate analysis . | Multivariate analysis . | ||
|---|---|---|---|---|
| RR* . | P . | RR* . | P . | |
| Global series | ||||
| p21 Hypermethylation | 4.1 | < .001 | 3.4 | .0001 |
| WBC above 50 000 mm3 | 3.1 | .006 | — | — |
| BCR-ABL positivity | 2.9 | .005 | — | — |
| T phenotype | 2.2 | .03 | — | — |
| Age older than 15 years | 1.8 | .03 | — | — |
| Childhood ALL | ||||
| p21 Hypermethylation | 4.0 | .007 | 3.1 | .01 |
| T phenotype | 2.5 | .04 | — | — |
| WBC above 50 000 mm3 | 2.2 | .05 | — | — |
| NCI poor risk | 2.2 | .05 | — | — |
| PETHEMA poor risk | 1.8 | .05 | — | — |
| Adult ALL | ||||
| WBC above 50 000 mm3 | 4.9 | .002 | 4.1 | .003 |
| p21 Hypermethylation | 3.2 | .003 | 3.0 | .01 |
| BCR-ABL positivity | 2.3 | .006 | — | — |
| Characteristic . | Univariate analysis . | Multivariate analysis . | ||
|---|---|---|---|---|
| RR* . | P . | RR* . | P . | |
| Global series | ||||
| p21 Hypermethylation | 4.1 | < .001 | 3.4 | .0001 |
| WBC above 50 000 mm3 | 3.1 | .006 | — | — |
| BCR-ABL positivity | 2.9 | .005 | — | — |
| T phenotype | 2.2 | .03 | — | — |
| Age older than 15 years | 1.8 | .03 | — | — |
| Childhood ALL | ||||
| p21 Hypermethylation | 4.0 | .007 | 3.1 | .01 |
| T phenotype | 2.5 | .04 | — | — |
| WBC above 50 000 mm3 | 2.2 | .05 | — | — |
| NCI poor risk | 2.2 | .05 | — | — |
| PETHEMA poor risk | 1.8 | .05 | — | — |
| Adult ALL | ||||
| WBC above 50 000 mm3 | 4.9 | .002 | 4.1 | .003 |
| p21 Hypermethylation | 3.2 | .003 | 3.0 | .01 |
| BCR-ABL positivity | 2.3 | .006 | — | — |
RR indicates relative risk; ALL, acute lymphoblastic leukemia. For other abbreviations, see Table 1.
Relative risk of treatment failure as compared with patients without the adverse feature. In multivariate analysis, relative risk was calculated after adjustment for other factors in the final model; risks are shown only for factors that achieved independent significance.
Discussion
In the present study, we demonstrate that epigenetic modification of p21 via hypermethylation represents a critical mechanism for inactivation of this gene in ALL. We found that aberrant promoter methylation of p21 occurs frequently in ALL and results in markedly diminished expression of p21. Our study indicates that the CpG islands within p21 promoter region are maintained in a partially methylated state in normal bone marrow as has been described for other normal tissues.27 In ALL cells, hypermethylation within this region reflects a gain in the degree of methylation at this CpG site. It is not known how methylation patterns at this CpG site were established for blood cells during development, nor is it clear to what degree the specific methylation patterns are involved in regulating p21 expression in normal marrow cells. Because several reports have not detected any mutations or deletions of p21,19 20 aberrant methylation may, at a minimum, be the most frequent way that expression of this gene is altered in this type of cancer.
Our results further suggest a tumor-suppressor role for p21 in this type of hematological malignancy. Loss of p21 could lead to defects in cell cycle regulation and confer a selective growth advantage for clones in ALL. Lack of p21 expression also facilitates the activity of other oncogenic pathways. For example, in the absence of p21, the onset of Ras-dependent oncogenesis is accelerated, with earlier tumor appearance and increased tumor multiplicity and aggressiveness in the transgenic/knockout mammary cancer model.31 Interestingly, although p53 abnormalities are the most common molecular lesions in human cancer,32 they are relatively less frequent in de novo ALL.33 One hypothesis is that loss of p21, which is transcriptionally activated by wild-type p53 protein, represents an alternative event contributing to abnormal cell cycle and cell-death regulation. In this sense, it has also been reported that a sizable proportion of ALL cell lines and primary tumors had negligible or limited expression of the p73 gene associated with hypermethylation of its promoter region.34 The p73 encodes for a protein that is both structurally and functionally homologous to the p53 protein. We have observed a close relationship between aberrant methylation of the p21 and that of the p73 gene (unpublished results), suggesting that silencing of the p73/p21 pathway by hypermethylation may contribute to development and/or progression of ALL.
In 4 (5%) of our ALL patients with decreased expression of p21, we were unable to detect hypermethylation in the p21 promoter region, suggesting an alternative mechanism of inactivation of the p21 gene in these patients. The transformation to inactive chromatin by histone deacetylation could be a good candidate mechanism to inactivate the p21 gene, as has been demonstrated in solid tumors.35 This issue raises an important question: could the hypermethylation of the p21 promoter region be a secondary event that is a consequence of the decreased p21 expression by another mechanism? The p21 might also negatively regulate DNA methylation, as it was shown to compete with DNA (cytosine-5)–methyltransferase for binding to proliferating cell nuclear antigen, thus antagonizing the DNA methyltransferase function.36 Therefore, abrogation of p21 expression (ie, by histone deacetylation) could favor the hypermethylation of its own promoter and the promoters of other genes, leading to a hypermethylator phenotype, as has been described in several studies of human neoplasms.37,38 Although this hypothesis is possible, it seems to be improbable because no aberrant methylation in the p21 promoter has been found in tumors with decreased p21 expression by histone deacetylation.35
Key tumor-suppressor genes altered in cancer include p53, p15, p16, and Rb. These genes can be inactivated by mutation, deletion, and hypermethylation. Each of these tumor suppressors has been reported to be altered in lymphoid malignancies. Furthermore, alterations in these genes have been shown to be associated with an unfavorable prognosis of ALL and B non-Hodgkin lymphomas.39,40 Loss of expression of p21 has been demonstrated to be associated with disease progression in patients with solid tumors, including mantle-cell lymphoma.17 18 However, the methylation status of the p21 promoter region and the prognostic value of p21 abnormalities in ALL have not been previously studied. In this study, we demonstrate that hypermethylation of the p21 gene provides important prognostic information in ALL patients. Hypermethylation of the p21 gene is a factor of poor prognosis in both childhood and adult ALL. Patients with hypermethylation of p21 had a poorer DFS than patients with a “normal” methylation pattern. Multivariate analysis confirmed that hypermethylation of the p21 gene was associated with a shorter DFS. Therefore, p21 methylation could have important clinical implications for guiding the selection of therapy and also for providing a basis for developing novel therapies, such as demethylation treatment.
In conclusion, our results indicate that the p21 gene is subject to methylation regulation at the transcription level and is a target of aberrant methylation in ALL cells. The methylation status of the p21 gene seems to be an important factor in predicting the clinical outcome of ALL.
Supported by grant 01/0662 from the Fondo de Investigaciones Sanitarias de la Seguridad Social, Spain; a grant from the Fundacion Carlos Haya, Malaga, Spain; and a grant from the Diputacion Provincial, Cordoba, Spain.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 U.S.C. section 1734.
References
Author notes
Jose Roman-Gomez, Hematology Department, Reina Sofia Hospital, Avda, Menendez Pidal s/n, 14004 Cordoba, Spain; e-mail:peperosa@teleline.es.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal