Abstract
Although human immunodeficiency virus (HIV)gag/pol DNA can be detected in naive T cells, whether naive T cells can be productively infected by HIV is still questionable. Given that interleukin-7 (IL-7) is a prospective therapeutic immunomodulator for the treatment of HIV, we evaluated the effect of IL-7 on promoting naive T-cell infection of laboratory-adapted (IIIB), M-tropic, and primary isolates of HIV. Initially, we determined that the 3 cell surface markers widely used to identify naive T cells (CD45RA+CD45RO−, CD45RA+CD62L+, and CD45RO−CD27+CD95low) are all equivalent in T-cell receptor excision circle content, a marker for the replicative history of a cell as well as for de novo T cells. We therefore used CD45RA+CD45RO− expression to define naive T cells in this study. We demonstrate that although untreated or IL-2–treated naive T cells are not productively infected by HIV, IL-7 pretreatment mediated the productive infection of laboratory-adapted, M-tropic, and primary isolates of HIV as determined by p24 core antigen production. This up-regulation was between 8- and 58-fold, depending on the HIV isolate used. IL-7 pretreatment of naive T cells also potently up-regulated surface expression of CXCR4 but not CCR5 and mediated the expansion of naive T cells without the acquisition of the primed CD45RO phenotype. Collectively, these data indicate that IL-7 augments naive T-cell susceptibility to HIV and that under the appropriate environmental milieu, naive T cells may be a source of HIV productive infection. This information needs to be considered in evaluating IL-7 as an immunomodulator for HIV-infected patients.
Introduction
CD4+ primed/memory T cells constitute the main target for productive human immunodeficiency virus (HIV) infection.1-3 Whether HIV productively infects naive T cells in vivo is still controversial.4-9 Naive T cells were shown to harbor replication-competent HIV,7,10 but are not an active site of HIV replication in the absence of mitogen stimulation.11 Even in studies demonstrating HIV infection of naive T cells, further analysis of integrated HIV indicated that HIV DNA did not always integrate into the genome.5,10 Infected naive T cells, nonetheless, have been identified in HIV-seropositive individuals,7,10 but it is unclear if such in vivo naive T cells were once primed cells that reverted to a naive phenotype12,13 or are truly naive T cells that are infected in vivo. In the simian immunodeficiency virus (SIV) model, SIV RNA was detected by in situ hybridization in naive T cells, suggesting that these cells may serve as an additional reservoir for HIV.14 Collectively, the consensus is that naive T cells, either in vivo or in vitro, do not support HIV productive infection mainly because these cells exist in a quiescent stage, a concept that is being challenged. The association between productive HIV replication and cell turnover is presumably due to the higher level of deoxyribonucleotides present for reverse transcription15during cell division or due to the expression, in actively replicating cells, of certain cellular factors that may allow for the stabilization of the HIV preintegration complex. However, recent data indicate that active cell division may not be required for HIV replication. In particular, ex vivo lymphoid histocultures point to HIV productive infection of naive T cells.16 These infected naive T cells were in the G0/G1A phase of the cell cycle, indicating that HIV may not require active cell replication for productive infection as was previously thought. A similar finding is also reported by Kinter et al.17 The mechanisms involved in the infection of naive CD4 T cells are unclear.
Interleukin-7 (IL-7) is a cytokine produced predominately by bone marrow and thymus stromal cells and is a critical factor in both B- and T-cell development.18,19 IL-7 treatment can also induce IL-2 receptor (CD25) expression on naive T cells, rendering them more responsive to the IL-2–mediated effects.20 In vivo IL-7 treatment was shown to successfully induce immune reconstitution following bone marrow transplantation in the mouse model, as measured by IL-7–mediated enhancement in the proliferation of immature thymocytes and improved survival after challenge with influenza virus.18 IL-7 has also been shown to increase a marker of de novo T-cell synthesis known as T-cell receptor excision circles (TRECs) in thymic explant models,21 possibly by inducing intrathymic T-cell receptor rearrangement22,23 or by enhancing the survival of early thymocyte progenitors.23-25 IL-7 can also enhance the survival of peripheral T cells by up-regulating the antiapoptosis gene, Bcl2.24 Given that IL-7 treatment can induce naive T-cell expansion without antigenic stimulation26,27 or differentiation28,29 and the recent reports that IL-7 is inversely correlated with CD4 counts in HIV+patients,30-32 we evaluated the impact of IL-7 treatment of naive T cells on their susceptibility to HIV infection by both T-cell–adapted and primary isolates of HIV.
Materials and methods
Isolation of peripheral blood mononuclear cells and naive T-cell subsets
Blood samples from healthy donors were collected with the use of EDTA Vacutainer tubes (Becton Dickinson, San Jose, CA). Peripheral blood mononuclear cells (PBMCs) were isolated by Ficoll-Hypaque density gradient centrifugation as previously described.33The 3 naive T-cell populations (CD45RA+CD45RO−, CD45RA+CD62L+, and CD45RO−CD27highCD95low) were isolated by means of multiple rounds of positive and negative immunoselection using magnetic beads, performed as described by the manufacturer (Dynal, Lake Success, NY). Briefly, to isolate CD45RA+CD45RO− cells, goat antimouse magnetic beads were preincubated with mouse antihuman CD45RO antibody (PharMingen, San Diego, CA) for 30 minutes at room temperature. Antibody-coated beads were then incubated with PBMCs for 30 minutes at 4°C. This procedure was repeated to further enhance the purity of CD45RO+ T-cell depletion. CD45RA+CD62L+ and CD45RO−CD27+CD95low populations were isolated after CD45RO depletion as described above, by means of subsequent positive-selection steps enriching for CD62L+ or CD27+. Depletion of CD95+ cells was combined with the CD45RO+ removal steps (for isolation of the CD45RO−CD27+CD95low population only). An additional step depleting CD8+ cells from PBMCs or enriching naive cells for CD4+ T cells was performed for some experiments. The purity of the isolated cells was greater than 99%, as evaluated by immunofluorescence staining with fluorochrome-conjugated antibodies and analysis by means of 3- and 4-color flow cytometry (FacsCaliber, Becton Dickinson, San Jose, CA).
Cytokine treatment of CD45RA+CD45RO−T cells
CD45RA+CD45RO− naive T cells were cultured in RPMI 1640 medium (Biowhittaker, Walkersville, MD) supplemented with 10% heat-inactivated human AB serum (Sigma, St Louis, MO); 1% penicillin/streptomycin (GIBCO-BRL, Grand Island, NY); and 2 mM l-glutamine (GIBCO-BRL). The cells were maintained at 1 × 106/mL and were propagated in the presence of 100 U/mL IL-2 (AIDS Research Reagent and Reference Program, Rockville, MD) or 1000 U/mL IL-7 (PharMingen).
Lymphocyte proliferation assay
CD45RA+CD45RO− cells were cultured in 96-well U-bottom plates at 1 × 106 cells per milliliter in 200 μL total volume. The cells were then stimulated with IL-2 (100 U/mL) or IL-7 (1000 U/mL). At 6 days after stimulation, the cells were pulsed with 1 μCi (.037 MBq)3[H]-thymidine (NEN Life Science Products, Boston, MA) for 6 hours, then lysed and harvested onto nitrocellulose paper by means of a plate harvester. The amount of radioactivity was measured with a scintillation counter.
Immunostaining and flow cytometric analysis
Isolated cell populations were stained immediately or after 6 days in culture for immunophenotypic analysis of T-cell surface markers by means of CD3-conjugated fluorescein isothiocyanate (FITC); CD4-conjugated phycoerythrin (PE), allophycocyanin (APC), or FITC; and CD8-conjugated peridinin chlorophyll protein (PerCP) antibodies. Cells were also stained for naive phenotypic markers for CD45RA-FITC:CD45RO-PE:CD4-PerCP; CD45RA-FITC:CD45RO-PE:CD8-PerCP:CD4-APC; CD45RA-FITC:CD62L-PE:CD4-PerCP; CD27-FITC:CD95-PE:CD45RO-PerCP). CXCR4 and CCR5 cell surface staining were performed by means of the following panel for CD4-FITC: CXCR4-PE:CD8-PerCP; CD4-FITC:CCR5-PE:CD8-PerCP. IL-7 receptor (IL-7R) staining was performed by means of CD45RA-FITC:IL-7R-PE:CD4-PerCP:CD45RO-APC, and CD45RA-FITC:IL-7R-PE:CD8-PerCp:CD45RO-APC antibodies, and the number of IL-7Rs per cell were quantified by means of quantibright beads. All values were normalized to isotope values. In some experiments, intracellular Ki-67 staining was performed by initially fixing the lymphocytes with fluorescence-activated cell sorter (FACS) lyse solution (Becton Dickinson). The fixed cells were then permeabilized by incubation with FACS permeabilization buffer (Becton Dickinson, San Jose, CA), followed by intracellular staining with Ki-67–FITC antibody and cell surface staining with CD45RO-PE and CD4- or CD8-PerCP. All monoclonal antibodies were purchased from Becton Dickinson (San Jose, CA) or PharMingen. Three- and 4-color flow cytometric analysis was performed by means of a FacsCaliber instrument and CELLQuest Software (Becton Dickinson, San Jose, CA).
Quantitative polymerase chain reaction–enzyme linked immunosorbent assay for the measurement of TRECs and HIV-1 gag/pol DNA content
To evaluate enrichment of naive T-cell populations (CD45RA+CD45RO−, CD45RA+CD62L+, CD45RO−CD27+CD95low) for TRECs, total DNA from these 3 isolated populations was extracted by means of DNAzol reagent (GIBCO BRL), as described by the manufacturer. The DNA was amplified for coding joint TRECs, by means of a previously described quantitative polymerase chain reaction–enzyme linked immunosorbent assay (PCR-ELISA) approach.34 Briefly, 1 and 0.5 μg genomic DNA was amplified in the presence of 200 μM digoxigenin–uridine (UTP) labeling mix; 2 μM each forward (5′-CTAATAATAAGATCCTCAAGGGTCGAGACTGTC-3′) and reverse (5′-CCTGTTTGTTAAGGCACATTAGAATCTCTCACTG-3′) primer; 1 × PCR buffer; 2.5 mM MgCl2; and 0.02 U/μL Taq polymerase (Boehringer Mannheim, Indianapolis, IN). The PCR protocol consisted of DNA denaturation at 95°C for 5 minutes, followed by 25 cycles of amplification at 90°C for 30 seconds, 60°C for 30 seconds, 72°C for 30 seconds, and a final extension step at 72°C for 7 minutes. Cloned coding joint TRECs at various copies (500 000, 50 000, 5000 and 0) were coamplified with each PCR to serve as standards for the assay. Precautions were taken to avoid PCR contamination. Quantitation of the coding joint TRECs was performed by means of an ELISA, following the manufacturer's instructions with few modifications (Boehringer Mannheim). Briefly, 10 μL each of the amplified product was denatured in a 96-well plate with 20 μL denaturation solution at room temperature for 10 minutes. Subsequently, 220 μL hybridization solution containing 7.5 pmol/mL biotin-labeled internal probe (5′-TCTGTGTCTAGCACGTAGCC-3′) (Fisher-Genosys, Woodlands, TX) was added. This mix was then transferred to a streptavidin-coated plate and incubated at 55°C for 3 hours. After extensive washing with an ELISA wash buffer, 200 μL antidigoxigenin peroxidase solution (1:100) was added to each well. The plate was then incubated at 37°C for 30 minutes with gentle shaking, washed, and exposed to 2,2-Azino-di- [3-ethylbenzthiaxoline sulfonate] substrate solution for 30 minutes at 37°C in the dark. Absorbance was read at 405 and 490 nm, and a standard curve of optic density (OD) values versus copies of input standard plasmid was generated by means of a software program (SoftMax Pro, Molecular Devices, Sunnyvale, CA). For each PCR-ELISA reaction, the standards were amplified in duplicates, and the mean OD was used to quantitate the number of copies per microgram of coding joint TREC in each of the unknown samples.
HIV-1 gag/pol DNA content was measured with a modified PCR-ELISA approach.34 Specifically, genomic DNA was extracted by means of GenomicPrep DNA isolation kit as described by the manufacturer (Amersham Pharmacia Biotech, Piscataway, NJ). The PCR mix consisted of 200 μM digoxigenin-UTP labeling mix; 1 × PCR buffer; 2.5 mM MgCl2; 2 μM each forward (SK38) (5′-ATAATCCACCTATCCCAGTAGGAGAAAT -3′) and reverse (SK39) (5′-TTTGGTCCTTGTGTTATGTCCAGAATGC-3′) primer (Fisher-Genosys), which recognize conserved sequences in HIV gag/polgenome; 1 or 0.5 μg genomic DNA; and 0.02 U/μL Taqpolymerase (Boehringer Mannheim). PCR cycling consisted of DNA denaturation at 95°C for 5 minutes, followed by 30 cycles of amplification at 90°C for 30 seconds, 60°C for 30 seconds, 72°C for 30 seconds, and a final extension step at 72°C for 7 minutes. Specified amounts of HIV-1 gag/pol DNA were generated from the 8E5 cell line, which consists of one proviral copy per cell. The HIV gag/pol standards were used at 25 000, 10 000, 6000, 2000, 1000, 500, and 0 copies and were coamplified with each PCR run. Subsequently, hybridization solution containing 7.5 pmol/mL biotin-labeled internal probe (SK19) (5′-ATCCTGGGATTAAATAAAATAGTAAGAATGTATAGCCCTAC-3′) (Fisher-Genosys) was used for ELISA quantitation, as described.34
HIV-1 infection
Phytohemagglutinin (PHA)–stimulated PBMCs or purified CD45RA+ CD45RO− naive T cells were infected with HIV IIIB, Bal, and primary isolates (302 056, 302 073, 302 142, 302 144, National Institutes of Health, AIDS Research and Reference Reagent program, Bethesda, MD) at 2 ng HIV p24 per 1 × 106 cells for 2 hours at 37°C. Subsequently, HIV-exposed cultures or mock-infected cultures were washed twice to remove unbound virus and cultured in the presence of IL-2 or IL-7. HIV primary isolates were determined to use CCR5 and not CXCR4 by genetically engineered cell line (ghost cell) analysis.35 HIV infection was performed either at day 0 of culture or 6 days after IL-2 or IL-7 treatment. HIV-1 infection was monitored by quantitation of p24 antigen in culture supernatants, by means of an HIV-1 p24 ELISA (AIDS Vaccine Program, Frederick, MD), or by measurement of HIV gag/pol DNA content with the PCR-ELISA approach described earlier.
Results
CD45RA+CD45RO−, CD45RA+CD62L+, and CD45RO−CD27+CD95lowCD4+ T cells are equivalent in TREC content
In contrast to primed/memory T cells, naive T cells have not yet encountered antigen, lack expression of cellular activation markers, and have a high response to mitogens (PHA, concanavalin A). To date, 3 phenotypes have been extensively cited in the literature to represent naive T cells, CD45RA+CD45RO−, CD45RA+CD62L+, and CD45RO−CD27+CD95low, yet there is no clear understanding of which phenotype best represents naive T cells. Prior to studies to evaluate the impact of IL-7 on naive T-cell susceptibility to HIV infection, we conducted comparative analysis of these 3 phenotypes to evaluate which phenotype is enriched for recent thymic emigrants; to this end, we used the TREC assay, which measures a byproduct of T-cell receptor rearrangements to generate episomal DNA deletion circles known as TRECs. These TRECs are diluted with each round of cell division and serve as markers of de novo T-cell synthesis, when cell turnover is normalized, or as markers of the replicative history of the cells (reviewed in Steffens et al36). We hypothesized that the phenotype that is best enriched in TRECs will be closely associated with the naive phenotype. CD45RA+CD45RO− were isolated by means of multiple immunoselection steps to purify these populations (Figure1A). The cells were evaluated for TREC DNA content by means of a quantitative PCR-ELISA approach.34 TREC content was equivalent in these 3 phenotypes (P = .67; Figure 1B). Furthermore, flow cytometric studies indicated that enrichment for the CD45RA+CD45RO− phenotype simultaneously enriched for the CD45RA+CD62L+ and CD45RO−CD27highCD95low populations (Figure 1C; P = .09). These comparative data provided a rationale for the use of CD45RA+CD45RO− phenotype in all subsequent studies, given that the 3 populations are equivalent in TREC content and that CD45RA+CD45RO−cells were the simplest to purify.
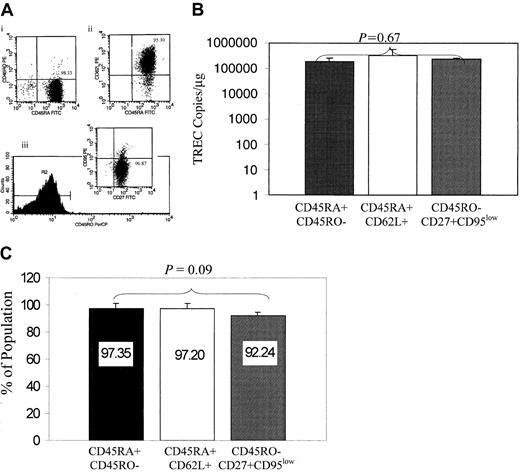
Evaluation of naive phenotypes for TREC content.
(A) The 3 naive phenotypes (CD45RA+ CD45RO−, CD45RA+CD62L+, and CD45RO−CD27+ CD95low) were isolated by sequential immunoselection, and the purity of the cultures is indicated in the upper right quadrant of the representative flow charts shown for CD45RA+CD45RO− (panel Ai), CD45RA+CD62L+ (panel Aii), and CD45RO−CD27+CD95low (panel Aiii). (B) TREC levels are equivalent in the CD4+ naive T-cell phenotypes. DNA from CD45RA+CD45RO−, CD45RA+ CD62L+, and CD45RO−CD27+CD95low populations was isolated, and the level of coding joint TRECs was quantified by PCR-ELISA. P = .67, obtained by comparing TRECs from the 3 populations; this is shown in the figure, which was based on a mixed linear model. Data are based on 5 different donors. (C) Isolation of the CD45RA+CD45RO− population enriches simultaneously for CD45RA+CD62L+ and CD45RO−CD27+CD95low naive subsets. CD45RA+CD45RO− T cells were isolated and evaluated by flow cytometry for the expression of CD62L, CD45RO, CD27, and CD95. The bar chart indicates that 97% of the CD45RA+CD45RO− cells are also CD45RA+CD62L+ and 92% are CD45RO−CD27+CD95low.
Evaluation of naive phenotypes for TREC content.
(A) The 3 naive phenotypes (CD45RA+ CD45RO−, CD45RA+CD62L+, and CD45RO−CD27+ CD95low) were isolated by sequential immunoselection, and the purity of the cultures is indicated in the upper right quadrant of the representative flow charts shown for CD45RA+CD45RO− (panel Ai), CD45RA+CD62L+ (panel Aii), and CD45RO−CD27+CD95low (panel Aiii). (B) TREC levels are equivalent in the CD4+ naive T-cell phenotypes. DNA from CD45RA+CD45RO−, CD45RA+ CD62L+, and CD45RO−CD27+CD95low populations was isolated, and the level of coding joint TRECs was quantified by PCR-ELISA. P = .67, obtained by comparing TRECs from the 3 populations; this is shown in the figure, which was based on a mixed linear model. Data are based on 5 different donors. (C) Isolation of the CD45RA+CD45RO− population enriches simultaneously for CD45RA+CD62L+ and CD45RO−CD27+CD95low naive subsets. CD45RA+CD45RO− T cells were isolated and evaluated by flow cytometry for the expression of CD62L, CD45RO, CD27, and CD95. The bar chart indicates that 97% of the CD45RA+CD45RO− cells are also CD45RA+CD62L+ and 92% are CD45RO−CD27+CD95low.
Impact of IL-7 treatment on naive cell proliferation and maintenance of the naive phenotype
IL-7 has been reported to stimulate the proliferation of naive T cells without acquiring the primed (CD45RO) phenotype.26 We have also confirmed this finding in our IL-7–treated naive T-cell cultures. Specifically, highly purified adult naive (CD45RA+CD45RO−) T cells were treated with IL-2 (100 U/mL), IL-7 (1000 U/mL), or PHA (4 μg/mL) or left untreated, and CD45RA+CD45RO− expression was evaluated by flow cytometry on days 0, 3, and 6. IL-7 treatment, like IL-2–treated cells, did not significantly alter the expression of the naive CD45RA+CD45RO− phenotype, while PHA, as expected, decreased the level of unprimed T cells (CD45RA+CD45RO−) by 53% after 6 days in culture (Figure 2A). To confirm that CD45RA+CD45RO− naive cells were proliferating in response to treatment with IL-7 without the acquisition of the primed (CD45RO) phenotype, we evaluated cellular proliferation using both a lymphocyte proliferation assay (LPA) and intracellular staining for Ki67, a nuclear antigen associated with cells in G1, S, M, or G2 phases of cell cycle but not in G0. At 6 days after IL-7 treatment, the naive T cells exhibited an approximately 7-fold increase in 3[H]-thymidine incorporation over unstimulated controls (Figure 2B). While the LPA response to PHA was diverse, with some donors exhibiting a higher level of thymidine incorporation than others, IL-7–treated naive T cells were more uniform in their response to IL-7–mediated cell proliferation. Intracellular Ki67 staining of IL-7–treated naive T cells, in contrast to IL-2–treated cultures, demonstrated a greater degree of proliferation of naive T cells, without the acquisition of the primed (CD45RO) phenotype (Figure 3). However, only a small subset of IL-7–treated naive T cells were proliferating. Specifically, 8% of the total lymphocytes treated with IL-7 were Ki67+ and CD45RO− (Figure 3Ai), and none of these proliferating (Ki67+) cells had switched to CD45RO+ phenotype after 6 days in culture (Figure 3Aiv-Avi). Of these IL-7–treated naive T cells, the majority of the proliferating cells were CD8+ cells (Figure 3Aiii) rather than CD4+ T cells (Figure 3Aii), albeit CD4+cells were also proliferating in response to IL-7 treatment. In contrast, all PHA-treated cells switched to the CD45RO phenotype (Figure 3Aiv-Aix), but the majority were Ki67− (Figure 3Avii-Aix), which may be due to the fact that the expression of Ki67 is degraded after the cells have completed the cell cycle and that by 6 days of PHA stimulation, the majority of the cells have already divided.
IL-7 treatment induces proliferation of CD45RA+CD45RO− naive cells without CD45RO up-regulation.
(A) CD45RA+CD45RO− T cells were isolated and left untreated or treated with IL-7 (1000 U/mL), IL-2 (100 U/mL), or PHA (4 μg/mL). PHA cultures were also treated with IL-2 (20 U/mL). The cells were analyzed by flow cytometry at day 0, 2, 4, and 6 for CD45RA+CD45RO− expression. Values shown represent at least 6 donors. (B) CD45RA+CD45RO− T cells were isolated and left untreated or were treated with IL-7 (1000 U/mL) or PHA (4 μg/mL). On day 6, the cells were pulsed with 1 μCi3[H]thymidine and cultured for an additional 6 hours prior to analysis by means of a scintillation counter. Values are the median of 5 different donors run in triplicate.
IL-7 treatment induces proliferation of CD45RA+CD45RO− naive cells without CD45RO up-regulation.
(A) CD45RA+CD45RO− T cells were isolated and left untreated or treated with IL-7 (1000 U/mL), IL-2 (100 U/mL), or PHA (4 μg/mL). PHA cultures were also treated with IL-2 (20 U/mL). The cells were analyzed by flow cytometry at day 0, 2, 4, and 6 for CD45RA+CD45RO− expression. Values shown represent at least 6 donors. (B) CD45RA+CD45RO− T cells were isolated and left untreated or were treated with IL-7 (1000 U/mL) or PHA (4 μg/mL). On day 6, the cells were pulsed with 1 μCi3[H]thymidine and cultured for an additional 6 hours prior to analysis by means of a scintillation counter. Values are the median of 5 different donors run in triplicate.
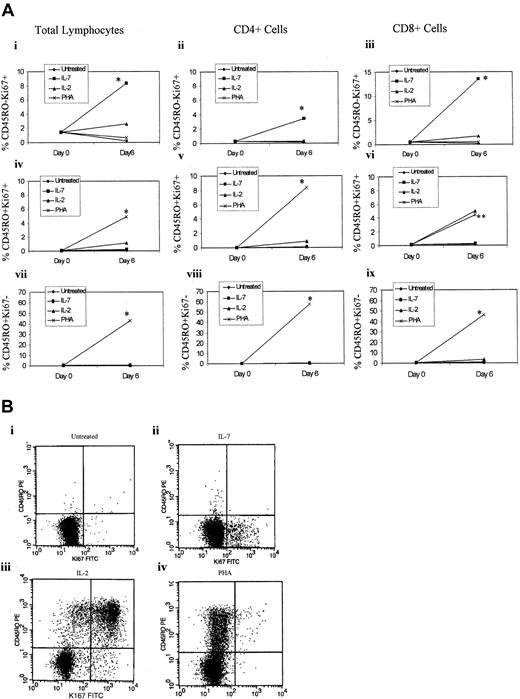
Impact of IL-7 on CD45RO and Ki67 expression.
(A) CD45RA+CD45RO− T cells were isolated and left untreated or treated with IL-7 (1000 U/mL), IL-2 (100 U/mL), or PHA (4 μg/mL). The cells were stained for CD45RO, CD4, CD8, and intracellular Ki67 and analyzed by flow cytometry by gating on total lymphocytes (panels Ai, Aiv, Avii), on CD4+ T cells (panels Aii, Av, Aviii), or on CD8+ T cells (panels Aiii, Avi, Aix). The top panel represents CD45RO−Ki67+cells; the middle panel represents CD45RO+Ki67+cells; and the bottom panel represents CD45RO+Ki67− cells. Cell surface and intracellular staining was performed on day 0 and day 6 after treatment. All values represent the median of 5 different donors treated in triplicates. *P < .05 (significant) of IL-7 (panels Ai-Aiii) or PHA (panels Aiv-Aix) treatments over the other culture conditions, which were based on the Kruskal Wallis test. **P < .05 (significant) PHA induction of percentage of expression of CD8+CD45RO+Ki67+cells over untreated and IL-7–treated cultures but not IL-2–treated naive T cells. (B) A representative Ki67 staining of naive T cells untreated (panel Bi), treated with IL-7 (panel Bii), treated with IL-2 (panel Biii), or treated with PHA (panel Biv).
Impact of IL-7 on CD45RO and Ki67 expression.
(A) CD45RA+CD45RO− T cells were isolated and left untreated or treated with IL-7 (1000 U/mL), IL-2 (100 U/mL), or PHA (4 μg/mL). The cells were stained for CD45RO, CD4, CD8, and intracellular Ki67 and analyzed by flow cytometry by gating on total lymphocytes (panels Ai, Aiv, Avii), on CD4+ T cells (panels Aii, Av, Aviii), or on CD8+ T cells (panels Aiii, Avi, Aix). The top panel represents CD45RO−Ki67+cells; the middle panel represents CD45RO+Ki67+cells; and the bottom panel represents CD45RO+Ki67− cells. Cell surface and intracellular staining was performed on day 0 and day 6 after treatment. All values represent the median of 5 different donors treated in triplicates. *P < .05 (significant) of IL-7 (panels Ai-Aiii) or PHA (panels Aiv-Aix) treatments over the other culture conditions, which were based on the Kruskal Wallis test. **P < .05 (significant) PHA induction of percentage of expression of CD8+CD45RO+Ki67+cells over untreated and IL-7–treated cultures but not IL-2–treated naive T cells. (B) A representative Ki67 staining of naive T cells untreated (panel Bi), treated with IL-7 (panel Bii), treated with IL-2 (panel Biii), or treated with PHA (panel Biv).
IL-7 promotes both T-tropic and M-tropic HIV replication in naive T cells
We evaluated the susceptibility of IL-2– and IL-7–treated naive T cells to HIV infection by T-tropic (IIIB) and primary isolates (strain 302 144 and 302 142). Receptor usage of the primary isolates was confirmed by ghost cell analysis. CD45RA+CD45RO− cells were then isolated and exposed to DNase-treated T-tropic or primary HIV strains; after extensive washing, the cells were propagated in the presence of IL-2 or IL-7. At 6 days after infection, HIV p24 was measured by means of a standard ELISA. Both untreated, IL-2–treated, and IL-7–treated cultures were not productively infected, as indicated by undetectable p24 (data not shown). To investigate whether HIV may be rescued from naive T cells after cytokine treatment, CD45RA+CD45RO− naive cells were infected as described above, cultured for 3 days with or without cytokine (IL-7 or IL-2) stimulation, and then cocultured with PHA-stimulated PBMCs depleted of CD8+ T cells. HIV productive infection still could not be rescued from naive T cells. This finding is in support of a study by Roederer et al,11 which demonstrated that HIV may not be rescued from naive T cells even after stimulation (anti-CD3/anti-CD28).
Given that IL-7 induces the proliferation of some of the naive cells, which may be a prerequisite for productive HIV infection, we evaluated the impact of IL-7 pretreatment of naive T cells to induce their susceptibility to productive HIV infection. CD45RA+ CD45RO− cells were pretreated with IL-7 or IL-2 for 6 days prior to HIV infection. T-tropic (IIIB), M-tropic (Bal), and primary (302 056, 302 073, 302 142, 302 144) HIV strains potently replicated in IL-7–pretreated naive T-cell cultures as determined by p24 analysis of culture supernatants (Figure4A). Specifically, 913 pg, 1191 pg, 7152 pg, 1072 pg, 4848 pg, and 2148 pg median p24 values were measured from HIV-1 IIIB, Bal, 302 056, 302 073, 302 142, and 302 144, respectively, after infection of IL-7–pretreated naive T cells (Figure4A). This level of IL-7–mediated induction of HIV replication is equivalent to an 8- to 58-fold increase, depending on the HIV isolate used.
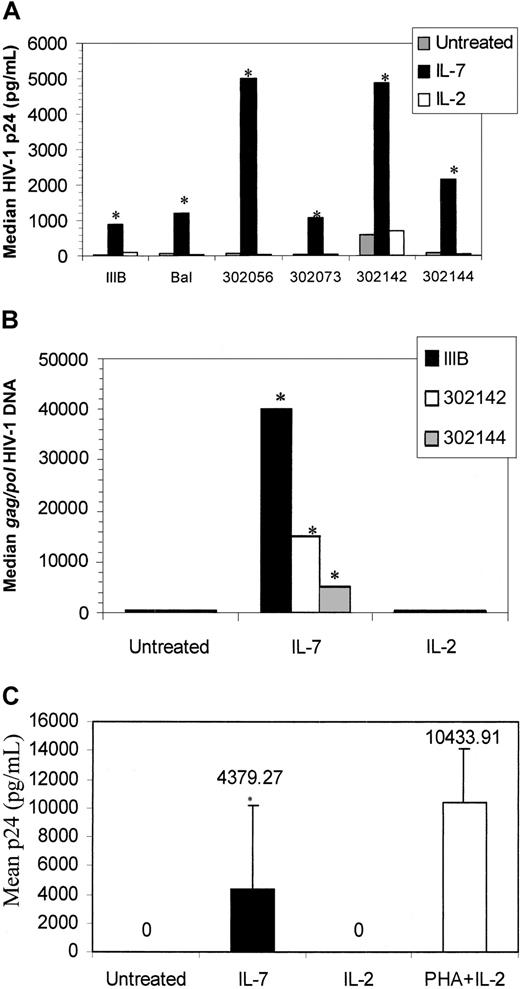
HIV infection of IL-7–treated naive T cells.
(A) CD45RA+CD45RO− T cells were isolated and pretreated with IL-2, IL-7, or were left untreated. At 6 days after cytokine treatment, the cells were infected with IIIB, Bal, 302 056, 302 073, 302 142, or 302 144. HIV-1 p24 levels were measured 6 days after infection. Values are the median p24 (pg/mL) measurement from 6 donors. Two donors were not responsive to the IL-7–mediated infection and are not included in this figure panel. (B) HIV gag/pol DNA was measured on day 6 after IIIB, 302 142, and 302 144 infection by means of a quantitative PCR-ELISA. (C) Supernatants from IL-7–, IL-2–, or PHA–treated/HIV IIIB–infected cultures shown in panel A were added to PHA-stimulated PBMCs, and p24 levels were measured on day 6. *P < .01 (significant) of IL-7–treated cells over untreated cultures, by means of the Wilcoxon rank test.
HIV infection of IL-7–treated naive T cells.
(A) CD45RA+CD45RO− T cells were isolated and pretreated with IL-2, IL-7, or were left untreated. At 6 days after cytokine treatment, the cells were infected with IIIB, Bal, 302 056, 302 073, 302 142, or 302 144. HIV-1 p24 levels were measured 6 days after infection. Values are the median p24 (pg/mL) measurement from 6 donors. Two donors were not responsive to the IL-7–mediated infection and are not included in this figure panel. (B) HIV gag/pol DNA was measured on day 6 after IIIB, 302 142, and 302 144 infection by means of a quantitative PCR-ELISA. (C) Supernatants from IL-7–, IL-2–, or PHA–treated/HIV IIIB–infected cultures shown in panel A were added to PHA-stimulated PBMCs, and p24 levels were measured on day 6. *P < .01 (significant) of IL-7–treated cells over untreated cultures, by means of the Wilcoxon rank test.
We also evaluated HIV gag/pol DNA content from representative IL-7–pretreated and infected cultures (Figure 4B). HIVgag/pol DNA content of HIV IIIB from IL-7–pretreated, infected cells (41 000 copies per microgram) was considerably more than that of primary isolates (16 000 and 4000 copies per microgram), yet the p24 values from primary isolate–infected cultures (5000 and 2500 pg/mL) was much higher than the T-tropic–infected cultures (1000 pg/mL). These data suggest that most of the DNA in IL-7–treated T-tropic–infected cultures may be unintegrated and thus account for the lower p24 measured, whereas the majority of primary isolates may be integrated. IL-2 pretreatment, on the other hand, did not promote HIV replication (Figure 4A) and did not increase HIV entry since HIV gag/pol content remained relatively the same with or without IL-2 pretreatment (Figure4B). Finally, the virions released from IL-7–pretreated naive T cells were also infectious as the supernatants from IL-7–pretreated/HIV IIIB–infected cells productively infected PHA-stimulated PBMCs (Figure 4C).
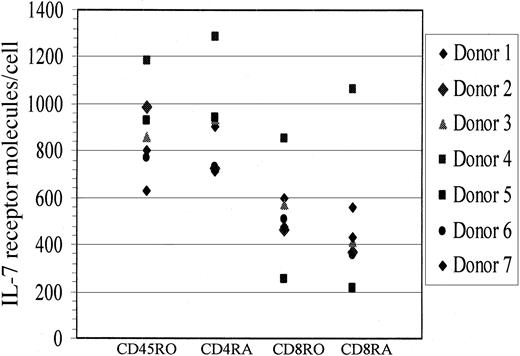
It is important to note that while the ability of IL-7 to mediate HIV productive infection of naive T cells was observed in 4 healthy donors, 2 healthy donors did not respond to this IL-7 effect. To examine whether IL-7R levels differed between donors that responded to IL-7 (n = 4) and those that did not (n = 2), we measured IL-7R levels from all donors and determined that IL-7R levels were comparable in the 2 groups as evaluated on CD4+ and CD8+naive T cells (Figure 5), and no correlation was found between nonresponders and IL-7R levels. Specifically, median IL-7R levels on CD4+ naive T cells from responders was at 723 molecules per cell versus 838.5 molecules per cell on nonresponders. These data suggest that the block to IL-7 responsiveness in a few donors is downstream of IL-7R expression and may be due to the lack of another factor (or factors) that contributed to their lack of permissiveness to IL-7–mediated HIV replication in naive T cells.
IL-7R levels on naive and memory T cells.
IL-7R levels on CD4 and CD8 naive (CD45RA) and memory (CD45RO) cells were quantified by means of quantibright beads from both healthy donors that responded to the IL-7–mediated effect (n = 4) and those that did not (n = 2).
IL-7R levels on naive and memory T cells.
IL-7R levels on CD4 and CD8 naive (CD45RA) and memory (CD45RO) cells were quantified by means of quantibright beads from both healthy donors that responded to the IL-7–mediated effect (n = 4) and those that did not (n = 2).
Impact of IL-7 treatment on CXCR4 and CCR5 expression on CD45RA+CD45RO− cells
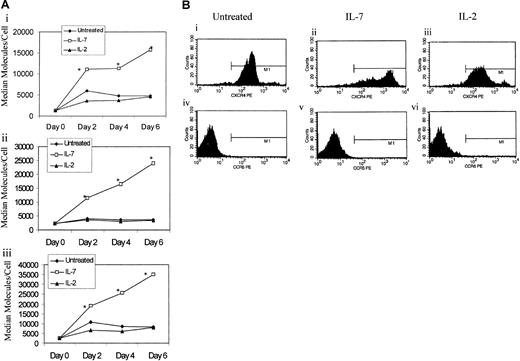
To determine whether enhanced susceptibility of IL-7–pretreated naive cells was due to altered chemokine coreceptor levels (CXCR4 and CCR5), CD45RA+CD45RO− naive T cells were cultured in the presence or absence of IL-7 or IL-2 stimulation, and the kinetics of CXCR4 and CCR5 expression was monitored for 6 days in culture. IL-7 mediated the up-regulation of CXCR4, which was evident at day 2 after stimulation and gradually increased thereafter (Figure 6). In IL-7–treated naive T cells, the level of up-regulation of CXCR4 was greater on CD8+ T cells than CD4+ T cells (Figure 6Aii-Aiii). IL-2, on the other hand, did not seem to significantly alter CXCR4 expression on naive T cells, as the level of CXCR4 on IL-2–treated naive T cells was similar to untreated cultures (Figure 6Ai-Aiii). The level of CCR5 expression on untreated naive T cells was below the detection limits of flow cytometry and was not augmented by IL-7 or IL-2 treatments (Figure 6Biv-Bvi). This observed impact of IL-7 treatment on CXCR4 and CCR5 expression on naive T cells was consistent in all donors examined, regardless of their infection outcome.
IL-7 mediates the up-regulation of CXCR4.
(A) CD45RA+CD45RO− cells were stimulated with IL-2 or IL-7 or left untreated. A portion of the cells were analyzed for expression of CXCR4 on total lymphocytes (panel Ai), CD4+ naive T cells (pane Aii), and CD8+ naive T cells (panel Aiii). Flow cytometric analysis was performed by means of quantibright beads to calculate the number of molecules per cell, which are shown on the y-axis. Data represent median values from at least 6 donors (panel A). *Significant P values (P < .001) significant of IL-7 cultures over untreated or IL-2–stimulated cultures as measured by the Kruskal Wallis test. (B) A flow cytometric representation of the data from panel A. Top panel corresponds to CXCR4 expression while bottom panel corresponds to CCR5 expression of untreated, IL-7–treated, or IL-2–treated naive T cells.
IL-7 mediates the up-regulation of CXCR4.
(A) CD45RA+CD45RO− cells were stimulated with IL-2 or IL-7 or left untreated. A portion of the cells were analyzed for expression of CXCR4 on total lymphocytes (panel Ai), CD4+ naive T cells (pane Aii), and CD8+ naive T cells (panel Aiii). Flow cytometric analysis was performed by means of quantibright beads to calculate the number of molecules per cell, which are shown on the y-axis. Data represent median values from at least 6 donors (panel A). *Significant P values (P < .001) significant of IL-7 cultures over untreated or IL-2–stimulated cultures as measured by the Kruskal Wallis test. (B) A flow cytometric representation of the data from panel A. Top panel corresponds to CXCR4 expression while bottom panel corresponds to CCR5 expression of untreated, IL-7–treated, or IL-2–treated naive T cells.
Discussion
In advanced stage of HIV disease, it is still unclear which cell type or types contribute to the propagation of HIV when CD4+ T cells are at their lowest numbers and the viral load is at its highest values. Naive T cells are not classically associated with potent virion release. At best, HIV may latently infect naive T cells, but a secondary signal such as mitogen stimulation is required to induce HIV replication in naive T cells. We evaluated the susceptibility of naive T cells to HIV infection. Initially, we validated the phenotype of naive T cells using the TREC assay. Three phenotypes (CD45RA+CD45RO−, CD45RA+CD62L+, and CD45RO−CD27+ CD95low) are associated with naive T cells. The isoforms of the CD45 (common leukocyte antigen) molecule traditionally used to distinguish naive (CD45RA+) from memory (CD45RO+) T-cell populations are problematic owing to the existence of revertants37 and double-positive populations.38-40 Other commonly used molecules, such as CD62L, may be shed from the cell surface. To determine which phenotype is best representative of naive T cells, we hypothesized that this phenotype will also be enriched in TREC content. To the contrary, our analysis indicated that the CD45RA+CD45RO−, CD45RA+CD62L+, and CD45RO−CD27+ CD95low populations are indistinguishable by TREC analysis. Flow cytometric evaluation also demonstrated that enrichment for the CD45RA+CD45RO− population subsequently enriched for the other 2 populations, suggesting that the CD45RA+CD45RO− population may be inclusive of the CD62L+CD27+CD95low phenotypes. As a result of this finding, all subsequent studies used naive cells represented by CD45RA+CD45RO− cell surface markers.
Others have reported that IL-7 mediates the expansion of CD4+ T cells without T-cell receptor engagement or the acquisition of the primed (CD45RO) phenotype.26-28,42 We validated this observation using flow cytometric analysis, LPA assay, and Ki67 staining. We have shown that IL-7, unlike IL-2 or PHA treatment of naive T cells, induces the expansion of the naive T cells while maintaining their naive phenotype (CD45RA+CD45RO−). Our data also indicate that it is the CD8+ T cells that are expanding more than the CD4+ T cells. Because IL-7 mediates the expansion of naive T cells, we evaluated the impact of IL-7 on mediating HIV productive infection of naive T cells. We demonstrate that pretreatment of naive T cells by IL-7, and not IL-2, is capable of priming these cells for the productive infection of HIV in a donor-dependent manner. Among the IL-7 responders, the productive infection occurs only when the cells are first exposed to IL-7 prior to virus infection, suggesting that IL-7 may be required to induce cellular division, which may be critical for HIV replication in naive T cells or induce cellular factors that are required for HIV replication. IL-7 may also promote HIV replication at the level of enhanced viral transcription. IL-7 induces DNA-binding activity of nuclear factor of activated T cells (NFAT) and activator protein 1 (AP-1).43Both NFAT- and AP-1–binding sites are present within the HIV long terminal repeat (LTR) (reviewed in Al-Harthi and Roebuck44). IL-7 itself does not activate nuclear factor (NF)–κB but acts in synergy with tumor necrosis factor (TNF)–α to induce NF-κB,45 which is a potent positive regulator of HIV transcription. Therefore, IL-7 may directly activate HIV transcription by promoting NFAT and/or AP-1 binding to their cognitive sites on the HIV LTR or in synergy with TNFα to induce NF-κB.
Although IL-7 up-regulated CXCR4, we believe that the IL-7–mediated productive infection of HIV may not be strictly at the level of enhanced entry. Alternative/additional mechanism(s) may be involved for the following reasons: (1) IL-7 did not up-regulate CCR5, yet M-tropic HIV replicated well in IL-7–pretreated CD45RA+CD45RO− T cells. This finding indicates that enhancement of HIV replication, at least for M-tropic HIV, in IL-7–treated cells may not be at the level of virus entry although up-regulation by other chemokine receptors, such as CCR3 or CCR2, may not be ruled out. IL-7–treated cells were, however, negative for CCR1 and CCR2 (data not shown). (2) For the reported enhanced T-tropic virus replication, untreated and IL-2–treated cells expressed considerable levels of CXCR4, yet did not replicate HIV IIIB well in comparison with potent HIV replication following IL-7 pretreatment of the naive T cells. (3) A recent report has demonstrated that IL-7 may induce virus replication in naturally infected PBMCs and that this level of induction may not be accounted for by the enhanced rate of cellular proliferation.46 In our studies, we also believe that IL-7 induction of HIV replication in naive T cells may not be due to enhanced cell proliferation since the percentage of cell turnover in response to IL-7 was only approximately 8% to 5%. Therefore, we believe that IL-7 may exert its effect through mechanisms that are not solely dependent on cell turnover. Finally, given that IL-7 and IL-2 share the common γ chain but that only IL-7 induced productive HIV replication in naive T cells, it is suggested that IL-7–mediated induction of HIV may rely on other factors in addition to the signaling that occurs through the common γ chain.
Recent studies have suggested that IL-7 may contribute to the restoration of homeostasis following T-cell depletion,30-32 as IL-7 induces the proliferation of immature thymocytes, protects developing cells from apoptosis by up-regulation of bcl-2,24 and can act as a mobilizer of T-cell precursors.42 IL-7 has also been shown to play an important role in the restoration of lymphopenia after bone marrow transplantation47 and other T-cell–depleted conditions.48 Interestingly, recent reports have correlated increased serum IL-7 levels with HIV-mediated T-cell depletion and increased viral load,30-32associating augmented levels of IL-7 with both viral replication and T-cell reconstitution. Collectively, these studies point to an important role for IL-7 in the response to and progression of HIV disease. Our finding that IL-7 can induce productive HIV infection of naive T cells in some donors suggests that it may also play a role in T-cell depletion and contribute to the observed inverse relationship between viral load and CD4 counts.30-32 A high level of provirus in CD4+CD45RA+ cells from neonates is associated with rapid disease progression in infants.9IL-7 may play an in vivo role in promoting the susceptibility of naive T cells from adults or neonates to HIV productive infection. Studies examining the mechanism(s) of refractoriness to the IL-7–mediated effect, despite normal levels of IL-7R in some donors, will be critical in identifying key cellular or genetic factors that may play a role in the outcome of the response. Future use of IL-7 as an immune modulator to promote immune reconstitution of HIV-infected patients should take into consideration that IL-7 can promote the susceptibility of naive T cells from some donors to HIV productive infection.
We thank Dr Richardson Fleuridor for assistance with statistical analysis.
C.M.S. and E.Z.M. are PhD candidates at Rush University (Chicago, IL), and this work is submitted in partial fulfillment of the requirement for the PhD.
Supported by American Foundation for AIDS Research (amFAR 02634-26-RGI and 02682-28-RGI) and Elizabeth Glaser Pediatric AIDS Foundation (PG-51112).
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 U.S.C. section 1734.
References
Author notes
Lena Al-Harthi, Rush–Presbyterian–St Luke's Medical Center, Department of Immunology/Microbiology, 1653 W Congress Pkwy, Rm 1577 JSC, Chicago, IL 60612; e-mail: lalharth@rush.edu.

![Fig. 2. IL-7 treatment induces proliferation of CD45RA+CD45RO− naive cells without CD45RO up-regulation. / (A) CD45RA+CD45RO− T cells were isolated and left untreated or treated with IL-7 (1000 U/mL), IL-2 (100 U/mL), or PHA (4 μg/mL). PHA cultures were also treated with IL-2 (20 U/mL). The cells were analyzed by flow cytometry at day 0, 2, 4, and 6 for CD45RA+CD45RO− expression. Values shown represent at least 6 donors. (B) CD45RA+CD45RO− T cells were isolated and left untreated or were treated with IL-7 (1000 U/mL) or PHA (4 μg/mL). On day 6, the cells were pulsed with 1 μCi3[H]thymidine and cultured for an additional 6 hours prior to analysis by means of a scintillation counter. Values are the median of 5 different donors run in triplicate.](https://ash.silverchair-cdn.com/ash/content_public/journal/blood/99/9/10.1182_blood.v99.9.3310/6/m_h80922495002.jpeg?Expires=1766026197&Signature=diMbIimDOOjODSzebpce4g1PJLrY0lVk7EZGhIoVXif4cPVfq8i5zIrRp0ro6UJsYxrsCeZrz-yEwADe-KgZCXsM97ve47j3VCdYRHuckLUlHoCikY4yeTYJzBfaNcyOxCoWYCavzLdHNSX-M3ZmQdgogvQO4KIyftTUdtSWyR627tQWLxuRpd-gVytHtLPo5Yw2klczt82iMd5ViIji8Ao05dCqUTTzOWQ8oJCQbUG55LdgpTqIyDEYh0sNn6C~PQsZ7hEVgYH-jYRVMIM1O6YSqJcAJmMIdWkxB3OvAIdyxUBpM1IFb8mqhfSl9~pHnywDt-X6aMC7MZ84uV3dow__&Key-Pair-Id=APKAIE5G5CRDK6RD3PGA)
This feature is available to Subscribers Only
Sign In or Create an Account Close Modal