Key Points
HCAR2 is identified as a receptor for heme.
Heme triggers HCAR2 expression in mice with SCD and intravascular hemolysis, which is regulated by HO-1.
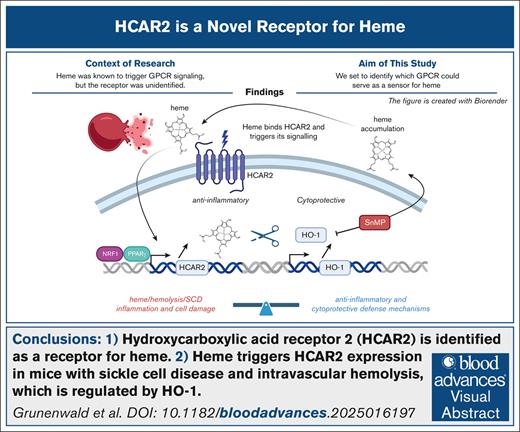
Visual Abstract
Extracellular heme, released during intravascular hemolysis in sickle cell disease (SCD) and hemolytic anemia, acts as a proinflammatory danger signal, requiring robust defense mechanisms. Previous studies identified G protein–coupled receptor (GPCR) signaling triggered by heme, but the specific receptor remained unknown. Transcriptomic analysis of bulk RNA sequencing of liver tissues from SCD and hemolytic mice (injection of phenylhydrazine) revealed GPCR signaling as a commonly enriched pathway. Unbiased screening of 241 GPCRs identified Hydroxycarboxylic Acid Receptor 2 (HCAR2/GPR109A), an anti-inflammatory receptor for niacin, as a novel heme sensor. Heme binding to human HCAR2 was validated using a functional reporter cell assay and direct interaction analyses via surface plasmon resonance and absorbance spectroscopy. In vivo, HCAR2 was upregulated in the liver of SCD and hemolytic mice, paralleling the expression of the heme-degrading enzyme heme oxygenase-1 (HO-1). HO-1 inhibition or heme injection further increased HCAR2 expression, indicating that heme acts as both a ligand and an inducer of HCAR2. These findings identify HCAR2 as a novel heme receptor and reveal a heme–HCAR2–HO-1 negative feedback loop involved in tissue protection in hemolytic diseases.
Introduction
Heme is an essential cofactor of aerobic metabolism, present in various hemoproteins, including hemoglobin, and is confined intracellularly under physiological conditions.1 However, it can be released during tissue damage, and notably intravascular hemolysis, occurring in sickle cell disease (SCD) and hemolytic anemias. Extracellular cell-free heme acts as an alarmin2,3 by inducing proinflammatory pathways, oxidative stress, complement cascade activation, and immune cells chemotaxis and activation.4,5 The level of subsequent damage depends on the balance between the deleterious effects of heme and the physiological defense mechanisms activated by heme release, which is still not well understood.
During heme overload, most cells in the body upregulate the cytoprotective heme degrading enzyme heme oxygenase-1 (HO-1) to counteract the oxidative stress and the inflammation induced upon hemolytic event.4 HO-1 is the dominant defense mechanism against heme,6 but whether other protective pathways are also triggered during intravascular hemolysis remains poorly understood.
The proinflammatory signaling of heme on monocytes, macrophages, and endothelial cells (ECs) is partly mediated via Toll-like receptor 4 (TLR4)/myeloid differentiation factor 2 complex.7-13 TLR4 inhibition or deficiency partially reduces vascular inflammatory response and acute liver injury in response to cell-free heme in models of intravascular hemolysis7 and SCD.8,14 Heme is a promiscuous molecule and can interact with a large set of proteins.15 Besides TLR4 engagement, other heme receptors including Receptor for Advanced Glycation product,16 the platelets receptors C-type lectin–like receptor 2, and glycoprotein VI (GPVI)17,18 have been reported. Heme also exerts a chemotactic role on neutrophils through an unidentified G protein–coupled receptor (GPCR).19,20 GPCRs constitute a family of >800 receptors characterized by 7 transmembrane domains linked at its intracellular domain to 3 G proteins subunits Gα, Gβ, and Gγ.21 The diversity of GPCRs,22 coupled with their signaling redundancy, makes it challenging to identify specific GPCRs involved in a given biological process.
Here, we identified the GPCR signaling as a common pathway in mice with SCD or intravascular hemolysis and explored the potency of GPCR to signal upon heme sensing. Using an unbiased screening approach, along with functional and binding assays, we identified the β-hydroxybutyrate and niacin receptor hydroxycarboxylic acid receptor 2 (HCAR2/GPR109A) as a new GPCR involved in heme sensing. Moreover, heme regulated the expression of this cytoprotective and anti-inflammatory receptor in an HO-1–dependent manner. These findings establish HCAR2 as a receptor for heme and uncover a heme–HCAR2–HO-1 negative feedback loop that contributes to tissue protection under conditions of heme release.
Materials and methods
Animal experimentation
Animal experimentation was conducted in accordance with the guidelines for the care and use of laboratory animals and approved by the French Ministry of Agriculture for phenylhydrazine hydrochloride (PHZ)–induced intravascular hemolysis (Autorisation de Projet utilisant des Animaux a des Fins Scientifiques number 26678 – 2020072013247945 v6) and SCD (Autorisation de Projet utilisant des Animaux a des Fins Scientifiques number 2148 – 2019091015099240v1). At the study end point, livers were collected and snap-frozen for subsequent RNA expression analysis.
PHZ (114715; Sigma-Aldrich) was freshly prepared at a concentration of 25 mg/mL in sterile phosphate-buffered saline immediately (PBS) before use. Mice received an intraperitoneal injection of PHZ at 900 μmol/kg (0.125 mg/g body weight) or PBS on day 0 and were euthanized by cervical decerebration 24 hours after injection (day 1) without perfusion to visualize vaso-occlusion if indeed present, following previously described protocols.23
SCD Townes HbAA and HbSS mice (aged 7-12 weeks) were used in the experiments, as previously described.24 The frozen livers and transcriptomic data from this study were used here in accordance with the reduce, reuse and recycle (3R) principle.
Hemin (ferriprotoporphyrin IX; Frontiers Scientific) was dissolved at 10 mM in 50-mM NaOH and 145-mM NaCl under sterile conditions. The pH of the solution was adjusted to pH 7.8 using phosphoric acid, then further diluted in sterile PBS (pH 7.4) immediately before use. Freshly prepared heme solution was injected IV into the retroorbital sinus of HbAA and HbSS mice at 24 μmol/kg, and mice were euthanized 4 hours after injection, as previously described.8,24,25 Sn(IV) mesoporphyrin IX dichloride (SnMP; Frontier Scientific) was prepared similarly to heme and diluted to 40 μmol/kg in sterile PBS. SnMP or PBS was administered intraperitoneally at consistent time points on days 0, 1, and 2. On day 2, heme or PBS was injected IV into the retroorbital sinus at 24 μmol/kg, 1 hour after the final SnMP injection. All mice were euthanized 4 hours after injection of heme without perfusion to visualize vaso-occlusion if indeed present. The experiment followed the protocol previously described,24 and frozen livers and transcriptomic data from this study were reused here in accordance with the 3R reuse principle.
Transcriptomic analyses: bulk RNAseq
The RNA sequencing (RNAseq) was performed in the GenomIC platform Cochin Institute INSERM U1016. Fastq files were then aligned using STAR algorithm (version 2.5.2b), and quality control of the alignment was realized with Picard tools (version 2.8.1). Reads were then counted using Featurecount (version Rsubread 1.24.1), and the statistical analyses on the read counts were performed with the DESeq2 package version 1.14.1 to determine the proportion of differentially expressed genes. Data visualization was performed using ggplot2 and ComplexHeatmap R packages. Data used in this study come from our SCD mouse transcriptomic atlas24 and intravascular hemolysis transcriptomic atlas (J. Peliconi, N.S. Merle, J. Lavergne, C. Brochier, F. Guinot, T. David, L.T. Roumenina, and A. Grunenwald, unpublished data, June 2025) and are available with following details: liver from PBS-injected C57Bl/6 mice, 10.5281/zenodo.14845241; liver from PHZ-injected C57Bl/6 mice, 10.5281/zenodo.14845299; liver from HbAA mice, 10.5281/zenodo.10963640; and liver from HbSS mice, 10.5281/zenodo.10962971.
Pathway analysis was performed using gene set enrichment analysis GSEA (v4.3.3) on the matrix of normalized counts from RNAseq data. GSEA was run using the following parameters: “gene_set” permutation; “weighted” enrichment statistic; Signal2Noise ranking metric; and gene sets with <15 genes or >500 genes were excluded. Results are analyzed based on normalized enrichment score (NES) and false discovery rate (FDR). Volcano plot of human peripheral blood mononuclear cells (PBMC) from patients with SCD undergoing vaso-occlusive crisis or acute chest syndrome (ACS) have been reanalyzed from GSE139912.
Messenger RNA expression analysis: QuantiGene
Frozen liver tissue sections (30-μm thick) were cut at –20°C using a Cryostat (Leica AS-LMD; Leica Biosystem) and homogenized in 200 μL of 1-Thioglycerol/Homogenization Solution (QuantiGene; Thermo Fisher). Mouse Hcar2 and Hmox1 target genes and housekeeping genes Actb, Gusb, Rps18, Rps3, Polr2a, and Gapdh underwent targeted hybridization and signal amplification following the manufacturer's recommendations. Streptavidin-phycoerythrin signals were detected using a Luminex photometer (Luminex Corporation). Blank well fluorescence was subtracted from the median fluorescence values, and housekeeping genes were validated based on the ratio of standard deviation to mean. Data analysis was conducted using GraphPad Prism software, normalizing mean fluorescence values to housekeeping gene expression and comparing gene expression levels with those of pooled control mice.
Heme-responsive GPCR
The high-throughput gpcrMAX and orphanMAX panels (DiscoveRx) were used to identify possible heme interactions with a selection of known GPCR or orphan GPCR targets. GPCR activity26 was monitored in reporter cell lines using the PathHunter β-arrestin assay. Briefly, cells are engineered to coexpress a GPCR and β-arrestin fused with 2 recombinant inactive β-galactosidase enzyme fragments. Upon GPCR signaling stimulation by a potential ligand, recruitment of β-arrestin allows b-galactosidase association and produces a chemiluminescence signal. The antagonist mode measures the loss of reporter activity of each GPCR during a competition assay performed with a known agonist in the presence of heme. The gpcrMAX assay evaluates 168 receptors, and the orphan assay evaluates 73 receptors. The stimulation of all these receptors in the reporter cell lines by heme was performed by the DiscoveRx profiling service.
The reporter cells were incubated with 3-μM heme in agonist or antagonist mode in the standard experimental conditions, and the top candidate was selected. The concentration of 3 μM was chosen based on manufacturer recommendation to limit interference with the assay and aligning with previous data on 2 to 5 μM of heme being bioavailable.27 The top candidate, HCAR2/GPR109A, was explored in a titration assay in agonist mode. Cells were washed with assay buffer before cyclic adenosine monophosphate detection to eliminate the effect of compound interference in signal generation.
Surface plasmon resonance (SPR)
All experiments were performed in HEPES-buffered saline with EDTA and P20 buffer (10-mM HEPES [N-2-hydroxyethylpiperazine-N′-2-ethanesulfonic acid] pH 7.4, 150-mM NaCl, 3-mM EDTA, and 0.005% Tween 20). Recombinant HCAR2 (NM-177551; Origene) and C3b (Comp Tech A114; serving as a negative control, because it does not bind heme) were immobilized on the general long chain sensor chips of Biacore 2000 equipment, following the manufacturer’s instructions. Serial dilution of 6 different concentrations of heme (starting with 39 nM) were injected as analytes with a flow rate of 10 μL/min. The association and dissociation were monitored for 5 minutes. Regeneration of the chip surface was performed by a 30-second exposure to 300-mM imidazole. The signal from the mock immobilized channel was subtracted. All measurements were performed at 25°C.
Spectrophotometry
Absorbance spectra of hemin were recorded by Cary series UV VB spectrophotometer at 414 nm (by Agilent Technologies). Human recombinant HCAR2 (NM-177551; Origene) was diluted to 727 nM in PBS. Aliquots of heme resulting in a concentration range from 0.35 to 10.2 μM were added to the optical cell–containing proteins and to a PBS-containing reference optical cell. The absorbance spectra in the wavelength range of 300 to 700 nm were recorded (300 nm/min).
Hemopexin enzyme–linked immunosorbent assay
Mouse plasmas were diluted 1:25 000 in reagent diluent. Hemopexin were then performed using the mouse hemopexin enzyme–linked immunosorbent assay kit (ab157716; Abcam), according to the manufacturer’s recommendations. Absorbance was measured at 450 nm with an Infinite 200 PRO spectrophotometer (Tecan).
Flow cytometry
Human umbilical vein ECs were cultured in supplemented endothelial growth medium-2 according to the manufacturer’s instructions (Lonza; CC-3156) in 6-well plate until reaching confluency. Media were washed in PBS 1× (Gibco; 14196-094) 2 times and cultured overnight in optimized minimum essential medium serum–free media (Gibco; 31985-062), with increasing concentration of heme (Frontiers Scientific; H651-9) ranging from 6.25 μM to 100 μM. Cells were harvested and washed with PBS 1× and further stained with live-dead (Invitrogen; L10119A) and anti-human HCAR2 primary antibody (Novus; NBP1-92180) and labeled with chicken anti-rabbit AF647 secondary antibody (Invitrogen; A21443). Staining was measured in BD Fortessa and analyzed with FlowJo v10.
Statistical analysis
Statistical analysis was performed using R (v4.4.1), and P value <.05 was considered significant for wet laboratory experiments. For bulk RNAseq, exactTest(0) under default options within edgeR (v3.34) was used. For in vitro work, paired samples were compared using a 2-tailed paired t test, unpaired samples were compared with a Mann-Whitney U test, and Kruskal-Wallis test with Dunn correction was used for multiple comparison. The correlation r value was determined using the Spearman test, and the statistical P value was computed with the corr.test() function. Statistical tests are defined in the figure legends. Mouse plasmas were compared using 1-way analysis of variance with Tukey multiple comparisons test performed on R studio.
Results
Hemolysis induces GPCR signaling in liver tissue
To evaluate the pertinence of the hypothesis of a GPCR implication in heme sensing19,20 in a context of heme release, we focused the study on the liver, the most affected organ of our SCD transcriptomic atlas.24 Gene set enrichment analysis was performed from the normalized matrix (DESeq2 normalization method) of PHZ mice and their PBS control counterparts, as well as on HbSS and HbAA mice. The analysis revealed a total of 454 pathways enriched in PHZ vs PBS and 2421 pathways enriched in HbSS vs HbAA mice. Among them, 390 pathways were commonly enriched across the 2 data sets (Figure 1A). Further analysis of these pathways revealed a significant enrichment of the GPCR ligand binding pathway in PHZ (NES, 1.82; FDR = .0195) and HbSS mice (NES, 2.13; FDR = 4.17 × 10−05; Figure 1B-E; supplemental Figure 1).
GPCR signaling is a common enriched pathway in liver of hemolytic and SCD mice. (A) Venn diagram plot showing the altered pathways determined from GSEA; altered pathway between PHZ-treated vs PBS-treated mice is in brown; and the altered pathway between HbSS vs HbAA mice is in green. The common altered pathways appear in yellow. (B-C) Bubble plot showing the absolute NES of the common altered pathways, highlighting the GPCR signature in PHZ- vs PBS-treated mice (B) and HbSS vs HbAA mice (C). The pathways enriched in PHZ and HbSS are shown in red, and the pathways enriched in their control counterpart are shown in blue. (D-E) Gene set enrichment score of the GPCR ligand binding pathway in PHZ vs PBS mice (D) and HbSS vs HbAA mice (E). FDR qval, false discovery rate q-value; NES, normalized enrichment score.
GPCR signaling is a common enriched pathway in liver of hemolytic and SCD mice. (A) Venn diagram plot showing the altered pathways determined from GSEA; altered pathway between PHZ-treated vs PBS-treated mice is in brown; and the altered pathway between HbSS vs HbAA mice is in green. The common altered pathways appear in yellow. (B-C) Bubble plot showing the absolute NES of the common altered pathways, highlighting the GPCR signature in PHZ- vs PBS-treated mice (B) and HbSS vs HbAA mice (C). The pathways enriched in PHZ and HbSS are shown in red, and the pathways enriched in their control counterpart are shown in blue. (D-E) Gene set enrichment score of the GPCR ligand binding pathway in PHZ vs PBS mice (D) and HbSS vs HbAA mice (E). FDR qval, false discovery rate q-value; NES, normalized enrichment score.
High-throughput screening of heme-sensing GPCR
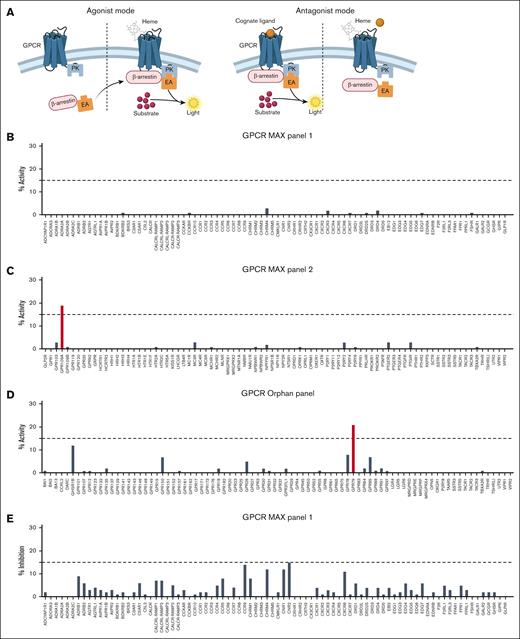
To find out whether there is a GPCR that can sense heme, we performed an unbiased, high-throughput screening of 241 GPCRs on model cells (HEK and CHO), provided by DiscoverX (Figure 2A). The gpcrMAX panel of 168 different GPCRs used in agonist mode (Figure 2B-C) revealed that most of the tested receptors do not respond to heme. Therefore, HCAR2 appeared to be a promising candidate because exposure to heme caused 19% of activity compared to vehicle control (Figure 2C). This was clearly different compared to the other receptors, which showed no or very weak positive activity. Noteworthy, GPR109B, which shares high sequence similarity with HCAR2/GPR109A, did not show any activity, suggesting a specific effect. Among the orphanMAX panel, testing 73 proteins with a GPCR structure for which functions and ligand remained unknown at the time of construction of the panel, there was a high “background” signal because most of the GPCRs showed some level of activity. Although GPR79 reached our threshold, it refers to a pseudogene and is most likely not involved in any biological mechanisms in cells (Figure 2D).
High-throughput screening for a GPCR for heme in antagonist mode. (A) Schematic overview of the PathHunter β-arrestin cell–based assay in agonist (left) and antagonist (right) mode. (B-D) Percentage of reporter activity as compared to control vehicle of 168 GPCRs tested in the GPCR Orphan and MAX and for 3-μM heme in agonist mode. (E-F) Percentage inhibition of reporter activity induced by 3-μM heme in the presence of the cognate ligand, compared to the cognate ligand with vehicle, tested in the GPCR MAX panels in antagonist mode. Figure created with BioRender.com.
High-throughput screening for a GPCR for heme in antagonist mode. (A) Schematic overview of the PathHunter β-arrestin cell–based assay in agonist (left) and antagonist (right) mode. (B-D) Percentage of reporter activity as compared to control vehicle of 168 GPCRs tested in the GPCR Orphan and MAX and for 3-μM heme in agonist mode. (E-F) Percentage inhibition of reporter activity induced by 3-μM heme in the presence of the cognate ligand, compared to the cognate ligand with vehicle, tested in the GPCR MAX panels in antagonist mode. Figure created with BioRender.com.
Because identified ligands are necessary to perform competition assay and measure competitiveness at the GPCR level, the antagonist mode could not be performed on orphan GPCRs. For GPCRs’ known ligands, we revealed that no receptor showed prominent signaling inhibition by the cognate ligand in presence of heme; ±10% activity was detected in all of them (Figure 2E-F). Interestingly, heme did not antagonize niacin-induced HCAR2 signaling, suggesting that heme and niacin could bind to 2 different epitopes (Figure 2F).
Validation of HCAR2 as a receptor for heme
Because HCAR2 was the most prominent candidate from our unbiased screening, we aimed to explore further the dose dependence of the heme-induced signaling. Moreover, we explored the molecular interactions between heme and this receptor. Titration of heme in the DiscoverX assay showed a significant and dose-dependent signal transduction, with an effective concentration 50 (EC50) of 21 μM (Figure 3A). As a comparison, the well-characterized ligand of HCAR2, niacin (nicotinic acid), showed an EC50 of 0.2 μM (Figure 3B), which is a 100× lower EC50 concentration.
Functional validation of HCAR2 as a receptor for heme measured by receptor signaling. (A) Titration of heme in the activity assay in agonist mode of HCAR2/GPR109A. (B) Titration of nicotinic acid (niacin) in the activity assay in agonist mode of HCAR2/GPR109A.
Functional validation of HCAR2 as a receptor for heme measured by receptor signaling. (A) Titration of heme in the activity assay in agonist mode of HCAR2/GPR109A. (B) Titration of nicotinic acid (niacin) in the activity assay in agonist mode of HCAR2/GPR109A.
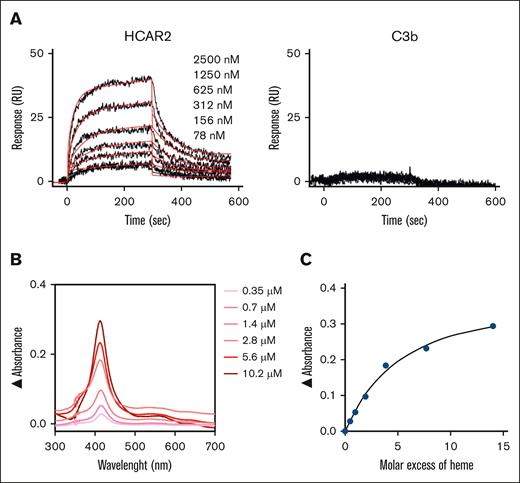
To test whether there is a direct binding of heme to recombinant HCAR2, SPR and ultraviolet-visible absorbance spectroscopy assays were applied. Indeed, heme significantly and dose dependently bound to HCAR2 immobilized on the SPR chip (Figure 4A), with an apparent affinity of 1 μM, even though correct establishment of the affinity of heme binding to proteins is difficult due to the inherent limitation of the method.
Biochemical validation of HCAR2 as a receptor for heme, illustrated by direct ligand/receptor binding. (A) Comparison by SPR of the interaction of immobilized recombinant HCAR2 compared to C3b (negative control) with different concentrations of heme (39 to 2500 nM). (B) Analysis of HCAR2-heme interaction by spectroscopy (0.35 to 10.2 μM). (C) Absorbance at the Soret peak (414 nM) as a function of the molar excess of heme over the protein. RU, response units.
Biochemical validation of HCAR2 as a receptor for heme, illustrated by direct ligand/receptor binding. (A) Comparison by SPR of the interaction of immobilized recombinant HCAR2 compared to C3b (negative control) with different concentrations of heme (39 to 2500 nM). (B) Analysis of HCAR2-heme interaction by spectroscopy (0.35 to 10.2 μM). (C) Absorbance at the Soret peak (414 nM) as a function of the molar excess of heme over the protein. RU, response units.
The binding was confirmed by ultraviolet-visible absorbance spectroscopy analysis by incubating HCAR2 with increased concentrations of free heme. These analyses revealed a heme dose-dependent increase in signal in the Soret spectral region, with a slight red shift to 414 nm, supporting that heme binds to HCAR2 (Figure 4B). The estimated stoichiometry from the fit of the absorbance at Soret peak as a function of the molar excess tends to saturation at 3.8× heme over protein excess, suggesting the presence of >1 binding site (Figure 4C).
Hcar2 correlates with Hmox1 expression under hemolytic conditions
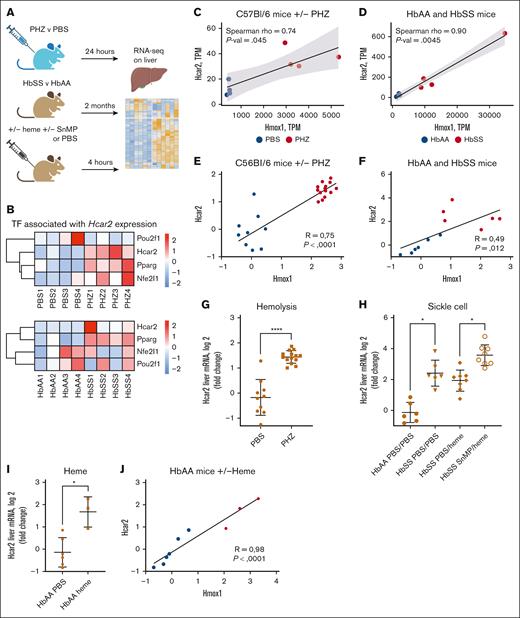
Analyzing the RNAseq data sets of the PHZ and SCD mice revealed that Hcar2 gene was upregulated in both pathological conditions. To further investigate whether heme induces the expression of its newly identified receptor and given the established link between HCAR2 and HO-1 expression in presence of niacin,28 we questioned whether HCAR2 and HO-1 expression are interrelated in hemolytic conditions. We analyzed RNAseq data sets obtained in different mouse models of heme-mediated diseases (Figure 5A), such as (1) intravascular hemolysis model by exposure to PHZ vs PBS control; (2) SCD mice (HbSS vs HbAA); and (3) SCD mice (HbSS) exposed to heme with or without a HO-1 inhibitor (SnMP). Hcar2 was among the significantly upregulated genes in the liver from the GPCR Ligand Binding gene signature (supplemental Figure 2A-B). We compared the expression levels of the 3 main transcription factors associated with Hcar2 expression, POU2F1, PPARγ (Pparg),29 and NRF1 (Nfe2l1; Uniprot). Among them, Nfe2l1 and Pparg in PHZ-treated mice (Figure 5B, upper panel) and Pparg in HbSS mice (Figure 5B, bottom panel) clustered with Hcar2 at the messenger RNA level, suggesting that they may be implicated in the expression of Hcar2 under hemolytic conditions. We further showed the correlation between the expression of Hmox1, as a proxy of heme overload, and one of Hcar2, with data obtained from RNAseq (Figure 5C-D), and further confirmed this observation by QuantiGene in an independent experiment (Figure 5E-F). Similarly, we confirmed the upregulation of Hcar2 expression upon hemolysis within kidneys, as another tissue sensitive to hemolysis (supplemental Figure 3). Figuring out the cell types expressing HCAR2 would help comprehend which cells may respond to heme under hemolysis and further express HO-1. According to single-cell RNA sequencing data from protein atlas, HCAR2 expression under basal condition seems to be restricted to Kupffer cells in the liver and to granulocytes (supplemental Figure 4A-B). We reanalyzed RNAseq data from PBMCs of patients with SCD30 during vaso-occlusive crisis (VOC) and ACS and found that HCAR2 was upregulated in both conditions compared to baseline condition (supplemental Figure 4C-D).
HCAR2 is overexpressed in hemolytic and SCD mice and its expression is regulated by heme/HO-1 axis. (A) Schematic overview of the experiment performed on mice. (B) Heat map of the normalized counts of the 3 main Hcar2 transcription factors of PHZ vs PBS (upper) and HbSS vs HbAA mice (lower). Normalized counts in each mouse are represented with row scale normalization. (C-F) Spearman correlation of Hcar2 and Hmox1 normalized gene expression levels in PHZ- and PBS-treated mice from RNAseq (C) and QuantiGene data (E) and HbSS and HbAA mice from RNAseq (D) and QuantiGene data (F). (G) QuantiGene analysis of Hcar2 gene expression levels in PHZ- and PBS-treated mice. (H) QuantiGene analysis of Hcar2 gene expression levels in HbSS and HbAA mice, either treated with PBS (left) or pretreated with SnMP, followed by an injection of 24-μM heme (right). (I-J) QuantiGene analysis of Hcar2 expression in HbAA mice injected with heme. (I) Level of expression of Hcar2. (J) Spearman correlation of Hcar2 and Hmox1 normalized gene expression in the HbAA mice injected with PBS or heme. ∗∗∗∗P < .0001; ∗P < .05; Mann-Whitney test in panels G,I. ∗P < .05; Kruskal-Wallis with Dunn test for multiple pairwise comparisons in panel H.
HCAR2 is overexpressed in hemolytic and SCD mice and its expression is regulated by heme/HO-1 axis. (A) Schematic overview of the experiment performed on mice. (B) Heat map of the normalized counts of the 3 main Hcar2 transcription factors of PHZ vs PBS (upper) and HbSS vs HbAA mice (lower). Normalized counts in each mouse are represented with row scale normalization. (C-F) Spearman correlation of Hcar2 and Hmox1 normalized gene expression levels in PHZ- and PBS-treated mice from RNAseq (C) and QuantiGene data (E) and HbSS and HbAA mice from RNAseq (D) and QuantiGene data (F). (G) QuantiGene analysis of Hcar2 gene expression levels in PHZ- and PBS-treated mice. (H) QuantiGene analysis of Hcar2 gene expression levels in HbSS and HbAA mice, either treated with PBS (left) or pretreated with SnMP, followed by an injection of 24-μM heme (right). (I-J) QuantiGene analysis of Hcar2 expression in HbAA mice injected with heme. (I) Level of expression of Hcar2. (J) Spearman correlation of Hcar2 and Hmox1 normalized gene expression in the HbAA mice injected with PBS or heme. ∗∗∗∗P < .0001; ∗P < .05; Mann-Whitney test in panels G,I. ∗P < .05; Kruskal-Wallis with Dunn test for multiple pairwise comparisons in panel H.
Furthermore, although ECs may show very low levels of expression of HCAR2, we show that ECs upregulate HCAR2 upon exposure to cell-free heme in vitro (supplemental Figure 4E).
Heme induces Hcar2 expression in mouse liver
Because Hcar2 expression was increased in PHZ- vs PBS-treated mice (Figure 5G), as well as in HbSS vs HbAA mice (Figure 5H), measured by QuantiGene, we hypothesized that heme may be responsible for this upregulation. Although heme injection did not alter Hcar2 expression in HbSS, treatment of HbSS mice with the HO-1 inhibitor SnMP before heme injection further increased Hcar2 expression, suggesting that Hmox1 controls Hcar2 levels, potentially by regulating the pool of available heme. To confirm that heme alone can upregulate Hcar2, we measured it in HbAA control mice injected with heme and indeed detected the expected upregulation (Figure 5I). Accordingly, cell-free heme was found elevated in HbSS only after pretreatment with SnMP (supplemental Figure 5A). Of note, heme levels at steady states were similar in HbSS and HbAA, differing from the 20% increase reported before but in older mice (aged 7-12 weeks here vs 16-24 weeks in that study). The reason for this discrepancy is unknown but may stem from differences in age, heme measurement assay, and animal facility conditions. We confirm previous results showing a dramatic drop of circulatory hemopexin in HbAA mice upon treatment with heme, whereas it is constitutively low in HbSS mice24 (supplemental Figure 5B). Moreover, Hcar2 expression correlated with Hmox1 in this experiment (Figure 5J).
Discussion
In this study, we identified HCAR2 (GPR109A) as a novel receptor for cell-free heme, unveiling a previously unrecognized mechanism of heme sensing. Through high-throughput GPCR screening and biophysical assays, we demonstrated that heme directly binds to HCAR2, a potent anti-inflammatory receptor for β-hydroxybutyrate and niacin. In mouse models of hemolytic conditions, including SCD and acute intravascular hemolysis, GPCR signaling emerged as a common pathway. Moreover, Hcar2 expression in the liver was regulated in a heme/HO-1–dependent manner. These findings suggest an unanticipated feedback loop in which heme triggers HCAR2 expression and further signals through HCAR2. In turn, HO-1 appears to modulate heme sensing by downregulating HCAR2, forming a negative feedback loop.
Hemolytic crises and chronic hemolysis are potent drivers of inflammation and organ injury, with heme acting as a damage-associated molecular pattern that promotes proinflammatory responses, primarily through TLR4 signaling31 and activation of the inflammasome.32 Although TLR4 activation by heme has been established in macrophages and ECs,8,31 our findings, along with previous reports, indicate that TLR4 inhibition does not completely abolish heme-induced responses.7,8 This suggests the presence of additional TLR4-independent mechanisms of heme sensing. In support of this, neutrophils have been proposed to use an unknown GPCR to mediate their response to heme.19
Transcriptomic analysis of our newly generated PHZ liver bulk RNAseq data set (manuscript in preparation) and its comparison to our recent transcriptomic atlas of the Townes SCD mice24 revealed enrichment of GPCR signaling pathways in both acute (PHZ) and chronic (SCD) hemolysis models. These results suggest the involvement of GPCRs in heme sensing alongside TLR4. To identify which is the heme sensor from the GPCR family, we conducted an unbiased, high-throughput screen of 241 GPCRs using a functional cell-signaling assay. This screen revealed a single candidate: HCAR2, known as a receptor for endogenous ligand β-hydroxybutyrate and also exogenous ligand niacin.33 The relevance of HCAR2 was confirmed through heme titration at the cellular level and by biophysical assays using SPR and absorbance spectroscopy. Notably, all known HCAR2 ligands, including niacin, and ketone bodies butyrate, β-hydroxybutyrate, and 4-hydroxynonenal are highly lipophilic, similar to heme. The mode of interaction of these ligands to HCAR2 was recently depicted and shows a complex binding mechanism.33-35 Further research is necessary to understand whether heme interacts with HCAR2 in the same binding pocket as the ketone bodies and niacin.
Proinflammatory stimuli such as lipopolysaccharide, interleukin-6, and interleukin-1β are known to upregulate HCAR2 expression in macrophages.36 Given that heme-induced TLR4 signaling, similar to lipopolysaccharide stimulation, promotes inflammatory responses, this mechanism may contribute to the upregulation of HCAR2 in hemolytic conditions. In vivo models of heme overload here similarly exhibited increased Hcar2 expression in the liver and kidney, which reproduced the effect of injection of cell-free heme or blocking of HO-1. Analysis of public data sets suggests that Kupffer cells express HCAR2 under basal condition, and we demonstrate that ECs can upregulate this receptor upon exposure to heme. These 2 cell types overexpress HO-1, particularly under hemolytic conditions.37 Proteomic analysis of neutrophils revealed no increase of HCAR2 expression in patients with SCD vs controls,38 but mature neutrophils do not overexpress HO-1 in response to heme either.39 RNAseq of monocytes40 from patients with SCD at steady states showed no significant change in HCAR2 expression, although a slight increase was observed in monocytes of a few individuals (data not shown). In contrast, our reanalysis of RNAseq from PBMCs of patients with SCD undergoing VOC or ACS30 revealed elevated HCAR2 expression compared to their respective baseline levels. Overall, the results obtained from public data sets on monocytes30,40 from patients suggest that low-grade hemolysis occurring in patients with SCD at steady state is potentially not strong enough to promote the negative feedback loop necessary to control cell-free heme present in small amount in the circulation, compared to the higher amounts when patients undergo VOC.41,42 These findings are consistent with our mouse data and support the relevance of HCAR2 in human hemolytic conditions.
Because heme is a potent alarmin, we anticipated identifying yet another proinflammatory receptor in our GPCR screen.20,43 Quite excitingly, the newly identified receptor HCAR2 is associated with strong anti-inflammatory pathways when activated by any of its known ligands, either natural or therapeutic molecules.33 Indeed, its activation inhibits macrophage chemotaxis in response to chemoattractants in vitro and protects mice from septic shock by reducing proinflammatory symptoms.36 Additionally, niacin treatment enhances endothelial protection by increasing reduced glutathione levels, a key intracellular antioxidant that scavenges reactive oxygen species.44 Even if niacin treatment did not improve endothelial function in patients with SCD in a clinical trial,45 an abstract from American Society of Hematology suggests that it could improve anemia and decrease reactive oxygen species in SCD mice.46 In addition, HCAR2 has been linked to anti-inflammatory functions in contexts of colitis and protecting colonic epithelial cells,47,48 protecting retinal EC barrier in diabetic retinopathy49-51 and brain inflammation.29,52 In the latter condition, studies have focused on HCAR2 functions on microglia and how it limits disease progression in a mouse model of Alzheimer52 and is currently tested in 2 clinical trials in Parkinson disease (NCT03808961 and NCT04677049). Because extensive body of literature supports the cytoprotective and anti-inflammatory role of HCAR2 signaling,35,53-59 we hypothesize that HCAR2 activation by heme may counteract its proinflammatory effects. This suggests that the heme–HCAR2–HO-1 axis could serve as a regulatory mechanism to balance inflammatory responses and promote cytoprotection through HO-1–dependent pathways. However, it is interesting to note that HO-1 exacerbates early brain injury during intracerebral hemorrhage.60 Although it remains speculative, HCAR2 activation on microglia by heme could work in this condition in a positive feedback loop, and further analysis of HCAR2 functions of this receptor in such context are needed.
Interestingly, treatment of HbSS mice with heme did not further induce Hmox1 or Hcar2 expression beyond levels observed in PBS-treated HbSS mice, supporting an adaptive response to chronic heme overload.24 Given that HO-1 primarily functions to degrade cell-free heme, increased HO-1 activity mitigate heme toxicity and limit Hcar2 expression. Supporting this, treatment of HbSS mice with the HO-1 inhibitor SnMP before heme injection led to a further increase in Hcar2 expression compared to PBS-pretreated HbSS mice. Hemopexin/CD91 complex is critical to regulate free heme from the circulation.61 However, upon consumption of the circulating hemopexin, the cells rely on HO-1 expression to control heme-derived stress. Therefore, HO-1 inhibition, concomitant with a lack of hemopexin, disrupts cells’ ability to degrade heme, leading to increased heme in the circulation and hence upregulation of Hcar2. These findings highlight the dynamic interplay between heme, HCAR2, and HO-1 in regulating inflammatory responses.
Given that neutrophils are known to be attracted by heme12 and that HCAR2 activation can induce neutrophil apoptosis62 and modulate T-cell responses,48 it is likely that HCAR2 signaling exerts cell type– and context-dependent effects. In our model, the early response dynamics suggest that monocytes, granulocytes, and ECs play a more prominent role than adaptive immune cells, given the acute nature of the hemolytic event. Moreover, our PathHunter approach screens 241 GPCR among the 1000 belonging to the human repertoire. Therefore, the existence of another proinflammatory GPCR-sensing heme is still possible.
In summary, we identified HCAR2 as a novel receptor for cell-free heme, unveiling a previously unrecognized heme-sensing mechanism regulated by HO-1 in hemolytic diseases. The observed upregulation of HCAR2 in intravascular hemolysis and SCD suggests its role in adapting to acute and chronic heme exposure. Our findings emphasize a negative feedback loop between heme catabolism and HCAR2 expression, offering new insights into the control of heme-induced inflammation. Further research is necessary to distinguish the specific signaling pathways activated by heme and ketone bodies or niacin through HCAR2 and to investigate their broader impact on inflammation and tissue protection in vivo. The identification of HCAR2 as a heme receptor expands our understanding of heme sensing and the delicate interplay between proinflammatory and cytoprotective responses in hemolytic diseases.
Acknowledgments
The authors are grateful for excellent technical assistance from the Centre d’Expérimentations Fonctionnelles team and the genotyping team of the Centre de Recherche des Cordeliers and for their support with animal experimentation. The authors are grateful for the next-generation sequencing analyses from the team of the Genom’IC platform, Institut Cochin, INSERM U1016.
This study was supported, in part, by Agence Nationale de la Recherche (ANR) JCJC-INFLACOMP 2015-2018 ANR-15-CE15-0001, by Fondation pour la Recherche Médicale (FRM) team label (FRM EQU202503020008), by the SVAR Complement excellence award (L.T.R.), and by France 2030 program through the Idex Université Paris Cité (InIdex GR-Ex ANR-18-IDEX-0001), and by INSERM. A.G. received a fellowship from the FRM.
Authorship
Contribution: N.S.M., A.G., and L.T.R. designed the research; A.G., J.P., A.Z., N.S.M., and J.D.D. performed the research; A.G., N.S.M., and J.D.D. analyzed data; A.G., L.T.R., and N.S.M. discussed the data and wrote the manuscript; and all authors revised and approved the manuscript.
Conflict-of-interest disclosure: L.T.R. reports research grants on intravascular hemolysis and sickle cell disease from CSL Behring and Roche. The remaining authors declare no competing financial interests.
Correspondence: Lubka T. Roumenina, Cordeliers Research Center, INSERM UMRS 1138, 15 rue de l'Ecole de Medecine; 75006 Paris, France; email: lubka.roumenina@sorbonne-universite.fr; and Nicolas S. Merle, Cordeliers Research Center, INSERM UMRS 1138, 15 rue de l'Ecole de Medecine; 75006 Paris, France; email: nicolas.merle@inserm.fr.
References
Author notes
L.T.R. and N.S.M. contributed equally to this study
All raw data from the RNA sequencing are available publicly. Accession numbers are given in “Materials and methods.”
The full-text version of this article contains a data supplement.