Abstract
The phenotype of germinal center (GC) B cells includes the unique ability to tolerate rapid proliferation and the mutagenic actions of activation induced cytosine deaminase (AICDA). Given the importance of epigenetic patterning in determining cellular phenotypes, we examined DNA methylation and the role of DNA methyltransferases in the formation of GCs. DNA methylation profiling revealed a marked shift in DNA methylation patterning in GC B cells versus resting/naive B cells. This shift included significant differential methylation of 235 genes, with concordant inverse changes in gene expression affecting most notably genes of the NFkB and MAP kinase signaling pathways. GC B cells were predominantly hypomethylated compared with naive B cells and AICDA binding sites were highly overrepresented among hypomethylated loci. GC B cells also exhibited greater DNA methylation heterogeneity than naive B cells. Among DNA methyltransferases (DNMTs), only DNMT1 was significantly up-regulated in GC B cells. Dnmt1 hypomorphic mice displayed deficient GC formation and treatment of mice with the DNA methyltransferase inhibitor decitabine resulted in failure to form GCs after immune stimulation. Notably, the GC B cells of Dnmt1 hypomorphic animals showed evidence of increased DNA damage, suggesting dual roles for DNMT1 in DNA methylation and double strand DNA break repair.
Introduction
On T-cell dependent activation, resting/naive B cells (NBCs) can be induced to migrate into lymphoid follicles and form germinal centers (GCs).1,2 GC B cells subsequently undergo massive clonal expansion and mutagenesis mediated by activation-induced cytosine deaminase (AICDA).2 Tolerance of simultaneous proliferation and genomic instability is a hallmark of the GC B-cell phenotype and is required for development of B-cell clones able to generate high-affinity antibodies.1,2 AICDA not only induces mutations within the immunoglobulin loci but also localizes to many other sites of the genome including promoters and coding sequences of actively transcribed genes enriched in RGYW DNA motifs.3-6 AICDA-induced mutations can thus occur at many sites throughout the genome in normal GCs.3,6 In accordance with these observations, AICDA has been demonstrated to play a critical role in lymphomagenesis.7 While genetic diversity of B-cell clones within GCs is important for the emergence of cells encoding high-affinity immunoglobulins, it also provides opportunities for the emergence of malignant clones. In fact a majority of B-cell neoplasms originate from cells that have transited the GC reaction.1
Induction of the GC phenotype requires that NBCs undergo major changes in gene expression patterning, the basis of which are not fully understood. These shifts are mediated in part by transcription factors such as BCL6 and BACH28-10 and histone modifying enzymes such as EZH2.11 However, differential methylation of CpG dinucleotides is also known to control tissue specific gene expression.12,13 CpG methylation is mediated by a family of DNA methyltransferase enzymes (DNMTs).14 Of these, DNMT1 primarily mediates maintenance methylation, because of its preference for hemimethylated DNA15 ; while DNMT3A and 3B primarily mediate de novo DNA methylation. Differential methylation occurs at the earliest stages of lymphopoiesis16 and Dnmt1 hypomorphic mice accordingly display skewed hematopoietic differentiation toward the myeloid lineage,17 but the role of DNMT1 in mature B cells has not been studied in a detailed manner.
Both aberrant DNA hypermethylation and hypomethylation have been shown to occur in lymphomas derived from GC B-cells such as diffuse large B-cell lymphomas (DLBCL).18,19 Furthermore, DLBCLs with GCB (Germinal Center B-cell like) versus ABC (Activated B cell-like) gene expression signatures display distinct DNA methylation profiles,18 suggesting that cytosine methylation may contribute to the distinct phenotypes of these tumors. Very little is known regarding mechanisms of DNA demethylation, but reports have suggested that cytosine deamination mediated by AICDA followed by base excision repair might contribute to this process by replacing methylated cytosines with new, unmethylated nucleotides.20-23 To determine whether differential DNA methylation patterning occurs naturally in GC B-cells, we examined DNA methylation profiles and the potential role of DNMTs in mediating the GC B cell phenotype. The data suggest a function for cytosine methylation in mature B-cell gene expression patterning with implications for the contribution of AICDA and DNMT1 to genetic and epigenetic instability during lymphomagenesis.
Methods
B-cell fractionation
Leftover human tonsils were obtained after routine tonsillectomies, performed at New York Presbyterian Hospital. All tissue collection was approved by the Weill Cornell Medical College Institutional Review Board. Tonsils were minced on ice and mononuclear cells were isolated using Histopaque density centrifugation. All washes were performed in PBS/2% BSA/2% EDTA. All antibodies were used at 1:100 dilution in cold PBS and staining was done for 10 minutes on ice, followed by 3 washes. The B-cell populations were separated using AutoMACS system (Milteny Biotec) using “posselD” program. Naive B cells (NBCs) were separated using depletion of GC cells, T cells and plasma and memory cells (CD10, CD3, and CD27), followed by enrichment for IgD+ B cells; Germinal Center B (GCB) cells were separated by positive selection with CD77 (anti-CD10: BD Biosciences; anti-CD3: BD Biosciences; anti-CD27: BD Biosciences; anti-CD77: AbD Serotec; anti-IgD: BD Biosciences). Purity check was performed using CD38 and CD77 staining for GCB fractions, and CD38 and IgD for NBC fractions. Purity of all samples is specified in supplemental Table 1 (available on the Blood Web site; see the Supplemental Materials link at the top of the online article). Representative scattergrams are in supplemental Figure 1.
High molecular weight genomic DNA extraction
Genomic DNA was extracted from 5 × 106 NBCs and GCB using the Puregene Gentra cell kit (QIAGEN). High Molecular Weight DNA was diluted in water and the quality was assessed using 1% agarose gel to assure no shearing.
DNA methylation profiling using HELP and custom oligonucleotide arrays
HELP assays were performed as per our standard protocol.24 In brief, 1 μg of genomic DNA was digested with HpaII and MspI (NEB). The digestion products were extracted and then subjected to ligation of HpaII adapter using T4 DNA Ligase. This was followed by PCR amplification and labeling of HpaII and MspI digestion products. The PCR products were cohybridized to custom NimbleGen HELP microarrays (NimbleGen Inc). The microarray design was previously documented and represents ∼ 50 000 CpGs corresponding to 14 000 promoters.18,24 Design files are available on request. Data can be found in the National Center for Biotechnology Information's GEO: GSE 31671.
DNA methylation profile analysis
Primary data processing was performed using the published HELP pipeline with modifications.25 For details see supplemental Methods.
Luminometric methylation assay
Luminometric methylation assay (LUMA) is based on enzymatic digestion of DNA with EcoRI and isoschizomer enzymes MspI or HpaII, followed by single nucleotide extension and DNA pyrosequencing and luminometric detection of incorporated dNTP.26 For details see supplemental Methods.
Cytosine methylation mass spectrometry
DNA hydrolysis was performed as previously described.27 Quantification was done using a LC-ESI-MS/MS system (Agilent 1200 Series liquid chromatography machine in tandem with the Agilent 6410 Triple Quad Mass Spectrometer) in multiple reaction monitoring (MRM) mode as described.27 For details see supplemental Methods.
Single locus DNA methylation assays
EpiTYPER assays were performed on bisulfite-converted DNA. Bisulfite conversion was performed using EZ DNA Methylation kit from Zymo Research. EpiTYPER primers for technical validation were designed to cover the flanking HpaII sites of selected HpaII amplifiable fragments (HAF), as well as HpaII sites up to 2000 bp upstream of the downstream site and up to 2000 bp downstream, to cover all possible alternative sites of digestion. For the biologic validation of the signature genes, primers were designed to cover CpG islands associated with the respective HAFs. All primers were designed using Sequenom EpiDesigner BETA software (http://www.epidesigner.com/). Primer sequences are shown in supplemental Table 2.
Quantitative real-time PCR
Total RNA was extracted from 107 cells using RNeasy mini kit from QIAGEN, and eluted in RNAse-free water. cDNA synthesis was done using the Superscript III First Strand Kit from Invitrogen. The expression was determined using Power SYBRGreen from Applied Biosystems and a DNA Engine Opticon 2 thermocycler from Bio-Rad. Relative gene expression was calculated using ddC(t) method. Primer sequences are shown in supplemental Table 3.
Immunohistochemical studies
The antibodies used for immunohistochemistry are listed in supplemental Table 4. For details see supplemental Methods. Densitometry was performed after slides were scanned with a Zeiss Mirax Slide Scanner using objective 20×/0.8NA. Densitometry analysis was performed using the Mirax Viewer 4.12.22.1 Software (Carl Zeiss Inc).
Western blotting
Cells (1 to 5 × 106) were lysed in modified RIPA buffer (50mM Tris, 250mM NaCl, 1% Nonidet P-40, 0.5% deoxycholate, 0.1% SDS) containing a protease inhibitor cocktail. The proteins were transferred to a PVDF membrane (Bio-Rad) and blotted with an antibody specific for DNMT1 (Novus Biologicals) at 1:1000 dilution. The membrane was stripped and reblotted with anti-actin antibody (Santa Cruz Biotechnology) at 1:3000 dilution.
Germinal center assessment in mice
The Research Animal Resource Center of the Weill Cornell Medical College of Medicine approved all mouse procedures. Ten- to 12-week-old C57/Bl6 mice were immunized intraperitoneally with 0.5 mL of a 2% sheep red blood cell (SRBC) suspension in PBS (Cocalico Biologicals) and killed after 10 days. For decitabine experiments: drug or vehicle was injected intraperitoneally starting at day 3 after induction of GC by SRBC and administered daily at concentrations of 15 mg/m2 and 30 mg/m2 for 7 consecutive days after which the mice were killed (day 10). Decitabine was dissolved in sterile water no longer than 1 hour before each administration. Dnmt1 N/+ and Dnmt1 R/+ mice were a generous gift of Dr Peter Laird28 and were used for assessment of the germinal center formation, which were induced with SRBC.
Flow cytometry detection of GC B cells
Single-cell suspensions from mouse spleens were stained using the following fluorescent-labeled anti–mouse antibodies: APC- or PerCP-conjugated anti-B220 (RA3-6B2), PE-conjugated anti-CD95 (Jo2; BD Pharmingen) PNA (Vector Laboratories), biotinylated anti-GL7 (GL7; eBioscience) Biotinylated PNA and biotinylated anti-FAS and anti-GL7 were developed with either streptavidin-PerCP (BD Pharmingen) or Streptavidin-Pacific Orange (Invitrogen Molecular Probe) protein conjugates. 7-AAD was used for the exclusion of dead cells. Data were acquired on LSRII flow cytometer (BD Biosciences) and analyzed using FlowJo 7.6.4 software (TreeStar).
Results
GCB cells display a specific cytosine methylation pattern
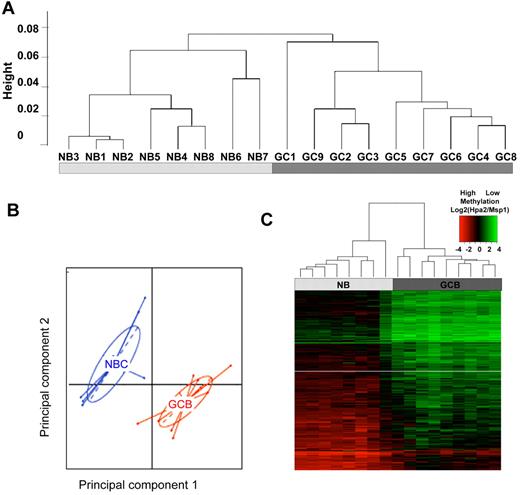
Because DNA methylation patterning is an important determinant of cellular phenotypes we reasoned that GC B cells might acquire a specific cytosine methylation profile distinct from those of NBCs. We therefore fractionated NBCs (IgD+ CD10−) and GC B cells (IgD-CD77+) from discarded human tonsillectomy specimens. Eight NBC and 9 GC B-cell preparations with > 90% purity confirmed by flow cytometry were selected for further processing (supplemental Table 1 and supplemental Figure 1). Genomic DNA was extracted and subjected to DNA methylation profiling using the HELP (HpaII Tiny Fragment Enrichment by Ligation Mediated PCR) assay with microarrays representing ∼ 50 000 CpGs distributed among 14 000 gene promoters. For technical validation of HELP accuracy we performed single locus quantitative DNA methylation assays (MassArray Epityper) on a set of 10 randomly selected variable probesets. We observed strong correlation between methylation values determined by HELP and MassArray (R2 = 0.83; supplemental Figure 2). To determine whether epigenetic patterning varies significantly between GC B cells and NBC cells we performed unsupervised hierarchical clustering (1 − Pearson correlation distance and Ward's clustering method). This procedure readily distinguished NBCs from GC B cells (Figure 1A), indicating that GC B-cells diverge considerably from the methylation profiles of NBCs. Principal component analysis (PCA) further confirmed this observation, since the first 2 principal components again separated the 2 cell types (Figure 1B).
GC B-cells feature a predominantly hypomethylated DNA methylation signature. (A) Unsupervised hierarchical clustering using the Ward method was performed on all probesets and accurately segregated NB cells from GC B cells (GCB). (B) Principal component analysis of methylation values for NBs and GCBs. The first and second principal components separate NBs from GCBs, underscoring the overall differences in methylation patterning. (C) A signature of differentially methylated genes in GC B-cells versus NB cells based on P < .01 (moderated t test with BH correction) and methylation difference of ∼ 40% was identified and included 235 genes. A heatmap representation allows visualization of the finding that the majority of differentially methylated genes are hypomethylated in GC B cells
GC B-cells feature a predominantly hypomethylated DNA methylation signature. (A) Unsupervised hierarchical clustering using the Ward method was performed on all probesets and accurately segregated NB cells from GC B cells (GCB). (B) Principal component analysis of methylation values for NBs and GCBs. The first and second principal components separate NBs from GCBs, underscoring the overall differences in methylation patterning. (C) A signature of differentially methylated genes in GC B-cells versus NB cells based on P < .01 (moderated t test with BH correction) and methylation difference of ∼ 40% was identified and included 235 genes. A heatmap representation allows visualization of the finding that the majority of differentially methylated genes are hypomethylated in GC B cells
To identify differentially methylated genes between GC B cells and NBCs, we performed a moderated 2-tailed t test and selected probes with P < .01 (after correction for multiple testing using the Benjamini-Hochberg method) and an absolute difference in methylation of > 1.5 log HpaII/MspI, which corresponds to > 40% change in methylation (as shown in supplemental Figure 2). Two hundred twenty-five probesets corresponding to 235 unique transcripts (supplemental Table 5) satisfied these stringent criteria. The majority (n = 223) of the 225 differentially methylated probesets were relatively hypomethylated in GC B cells compared with NBCs (Figure 1C). By examining NBC and GC B-cell gene expression profiles obtained by RNA-seq, we found a nonstatistically significant trend for expression of these genes to be inversely correlated.
Differential DNA methylation affects functionally relevant genes in GC B cells
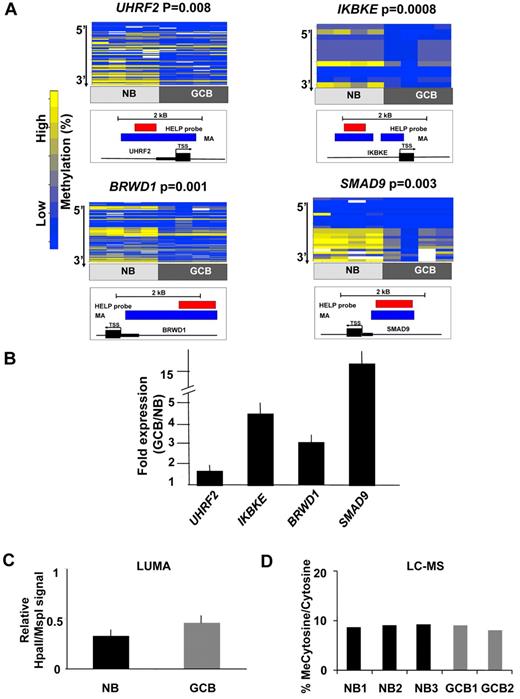
Differentially methylated genes in GC B cells were associated with specific biologic functions, with high evidence of enrichment of molecular functions associated with metabolism, synthases and synthetases (P < .05, Binomial test, supplemental Table 6) and moderate evidence of enrichment in chaperones and transporters (P < .1, Binomial test, supplemental Table 6). An Ingenuity gene pathway analysis revealed over-representation of 2 networks primarily representing components of NF-kB and MAP kinase signaling (supplemental Figure 3). To identify genes most likely to be functionally affected by DNA methylation changes, genes were ranked according to their degree of inverse correlation with gene expression (supplemental Table 7). Four inversely correlated genes were selected for single locus validation studies based on their biologic relevance and feasibility of MassArray primer design. Among these genes, UHRF2 (ubiquitin-like protein containing PHD and RING domain 2) is an E3 ubiquitin ligase that contributes to cellular proliferation and is considered to be a putative oncogene29 ; UHRF2 is a homolog of UHRF1, which is a cofactor for DNMT130 ; IKBKE (inhibitor of κ light polypeptide gene enhancer in B cells, kinase epsilon) induces NFkB signaling by phosphorylating IKB and is a known breast cancer oncogene31 ; BRWD1 (bromodomain and WD repeat domain containing 1) is a chromatin regulatory protein associated with SWI-SNF and key component of the Oct4 embryonic stem cell transcription factor network32,33 and SMAD9 (SMAD family member 9) a downstream effector of TGFbeta signaling that regulates cell migration in certain cell types.34 Four independent sets of purified NBCs and GC B cells were fractionated from human tonsils and examined by MassArray Epityping (Figure 2A). Consistent with HELP data, all 4 of these genes were hypomethylated in GC B cells compared with NBCs. Relative transcript abundance was validated by Q-PCR in 3 independent NBC and GC B-cell fractions and showed a 2-20-fold induction of these genes (Figure 2B). Collectively, these data show that differential DNA methylation in GC B cells features predominant hypomethylation of genes with possible functional impact in determining the GC B-cell phenotype.
Hypomethylation preferentially affects regulatory regions of the genome in GC B cells. (A) Four genes from the GC B-cell signature were selected for validation by MassArray. The results are represented as heatmaps in which the columns correspond to individual samples, while rows represent individual CpGs with color reflecting methylation value. P values are from moderated t test comparing methylation values from all tested CpGs between GC B cells and NBs. The location of MassArray amplicons (blue) and HELP probesets (red) relative to the transcriptional start site (TSS) of each gene (black) is illustrated below each heatmap. (B) The relative transcript abundance of the same 4 genes was measured by QPCR in 3 additional NB and GC B-cell specimens. The y-axis depicts fold expression difference in GC B-cells versus NBs calculated using ddCT method. All 4 genes are expressed at higher levels in GC B-cells, concordant with their hypomethylation. (C) LUMA assays performed on 3 NB and 2 GC B-cell specimens show a mild increase in the abundance of hypomethylated HpaII sites. The y-axis depicts relative signal of HpaII vs MspI signals. (D) Liquid chromatography-mass spectrometry was performed in 3 NB and 2 GC B-cell specimens. The y-axis depicts the percentage of methylcytosine versus total cytosines in each specimen.
Hypomethylation preferentially affects regulatory regions of the genome in GC B cells. (A) Four genes from the GC B-cell signature were selected for validation by MassArray. The results are represented as heatmaps in which the columns correspond to individual samples, while rows represent individual CpGs with color reflecting methylation value. P values are from moderated t test comparing methylation values from all tested CpGs between GC B cells and NBs. The location of MassArray amplicons (blue) and HELP probesets (red) relative to the transcriptional start site (TSS) of each gene (black) is illustrated below each heatmap. (B) The relative transcript abundance of the same 4 genes was measured by QPCR in 3 additional NB and GC B-cell specimens. The y-axis depicts fold expression difference in GC B-cells versus NBs calculated using ddCT method. All 4 genes are expressed at higher levels in GC B-cells, concordant with their hypomethylation. (C) LUMA assays performed on 3 NB and 2 GC B-cell specimens show a mild increase in the abundance of hypomethylated HpaII sites. The y-axis depicts relative signal of HpaII vs MspI signals. (D) Liquid chromatography-mass spectrometry was performed in 3 NB and 2 GC B-cell specimens. The y-axis depicts the percentage of methylcytosine versus total cytosines in each specimen.
Genomic abundance of methylcytosine in GC B cells
We noted that the GC B-cell signature featured a predominance of hypomethylated genes (Figure 1C). To determine whether this phenomenon extended beyond the 50 000 CpGs contained within HpaII sites (CCGG) detected by our microarrays we performed LUMA assays, a pyrosequencing based approach that detects the abundance of CpG methylation of the entire genomic complement of ∼ 2.3 million HpaII motifs.35 LUMA assays showed that on a genome-wide basis, HpaII sites showed a trend toward hypomethylation in GC B-cells (LUMA signal of 0.44 in GCB vs 0.33 in NB (LUMA signal is inversely correlated to the actual methylation, P = .085, t test; Figure 2C). HpaII sites are enriched in gene regulatory regions such as CpG islands.12 However, HpaII sites still represent < 10% of the genomic complement of CpG dinucleotides. To determine whether skewing toward hypomethylation extends more broadly throughout the genome we performed liquid chromatography followed by electrospray ionization and tandem mass spectrometry (LC-ESI-MS/MS).27 These studies indicated that the overall content of methylcytosine was similar between NBs and GCBs and approached 10% of all cytosines in the genome (Figure 2D). Therefore, DNA hypomethylation was mostly confined to the subset of methylcytosines located within gene regulatory regions enriched in HpaII sites.
DNA hypomethylation in GC B cells involves AICDA direct target genes
AICDA is known to associate with transcriptionally poised or actively transcribed genes.5,36 Because DNA hypomethylation in GC B cells was more associated with regulatory regions and activated genes, we wondered whether these might be associated with the activity of AICDA. First, we observed that differentially methylated genes in GC B cells have higher densities of putative AICDA recognition sites (RGYW4) in their promoters (Wilcoxon P < .001, t test P < .01). Unbiased analysis of DNA motifs using FIRE37 revealed the strong enrichment (P < .001, hypergeometric test) of a single Ebox-like motif with striking similarity to CAGGTG (supplemental Figure 4), a motif previously shown to be associated with sites of AICDA-mediated somatic hypermutation.38 AICDA binding sites and target genes were recently identified using ChIP-seq in activated mouse splenocytes.5 We found a significant overlap between differentially methylated sites and their murine orthologs biochemically proven to be AICDA binding sites (overlap greater than expected P = .056, hypergeometric test). Hypomethylated AICDA targets were more likely to be highly expressed in GC B cells versus NBCs (P < .005 Wilcoxon test). These data raise the possibility that AICDA might contribute to inducing hypomethylation in GC B-cells, in accordance with its recently identified role in DNA demethylation.21,23
DNA Methylation heterogeneity in GC B cells
Rapid proliferation and clonal expansion of GC B cells is crucial for AICDA induced mutagenesis to produce a diverse repertoire of mutated immunoglobulin loci and high affinity antibodies.1 This can lead to genetic heterogeneity,36 which is believed to play an important role in lymphomagenesis.7 By the same token clonal epigenetic diversification is hypothesized to contribute to malignant transformation.14 We wondered whether GC B cells would display increased heterogeneity in DNA methylation patterning in accompaniment to their genetic heterogeneity. Along these lines we observed that distribution of probeset methylation scores was different between GCB and NBC samples (P value < .001, Kolmogorov Smirnov Test and that the probeset scores were in fact significantly more heterogeneous among GC-B cell samples than NBCs. Accordingly, the bimodal methylation pattern usually observed in normal tissues12,25 was weaker in GC B cells compared with NBC samples (supplemental Figure 5 top panel). Altogether, we identified 1229 probesets at which GC B-cell samples have more heterogeneous methylation compared with NBC samples (see “Heterogeneity studies” in supplemental Methods for statistics). Furthermore, by comparing the distribution of per probe interquartile ranges (IQR, a measure of between-sample variation, see “Heterogeneity studies” in supplemental Methods for details), between GC B cells and NBCs, we found that GC B-cell samples display on average more between-sample methylation variation than NBC samples (P value < .001, Mann Whitney test; supplemental Figure 5 bottom panel). These differences are not accounted for by cellular heterogeneity since sample purity was verified by flow cytometry. The data suggest a scenario in which B cells may accumulate clonal epigenetic diversity, possibly induced by AICDA, and potentially contributing to lymphomagenesis, but could also reflect at least in part, heterogeneity of cell fate among GC B cells, with some being selected for terminal differentiation while others continue to proliferate and undergo somatic hypermutation.
DNMT1 is highly up-regulated in GC B cells
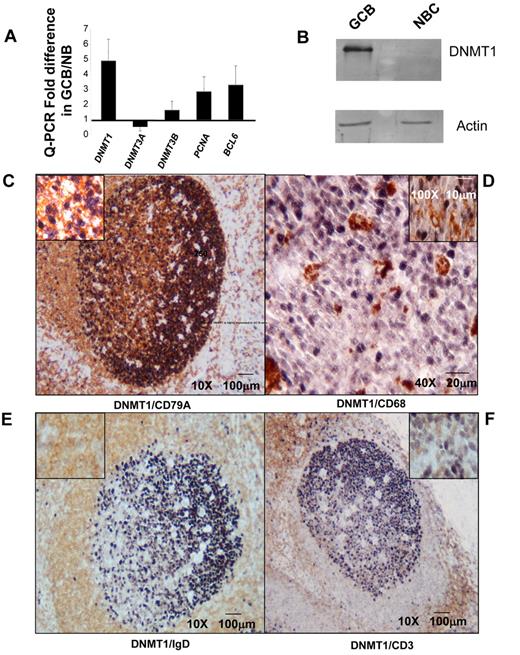
GC B cells must also be under considerable pressure to maintain DNA methylation patterning during their rapid proliferative phase, and our data show that they do indeed retain the same overall DNA methylation levels as NBCs. To determine which DNMTs are most essential to the GC B-cell phenotype we examined their relative transcriptional abundance by Q-PCR in NBCs and GC B cells. DNMT1 levels were highly up-regulated whereas DNMT3A and DNMT3B did not vary significantly or were down-regulated (Figure 3A). Up-regulation of DNMT1 was comparable in magnitude to other GC B-cell induced genes such as BCL6 and PCNA. Immunoblots confirmed the increased expression of DNMT1 in GC B cells versus NBCs (Figure 3B). To further delineate the patterns of expression of DNMTs in situ we performed immunohistochemistry for DNMT1, DNMT3A, and DNMT3B in normal reactive tonsillar tissue (Figure 3C-F and supplemental Figure 6A-C). High levels of DNMT1 were detected in GC B cells, particularly in the dark zone (centroblasts) with gradual decrease of expression in the light zone (centrocytes; Figure 3C). DNMT1 was localized to the nuclei of these B cells (Figure 3C inset). DNMT1 expression was not detected in CD68+ histiocytes, IgD+ NBCs, CD3+ T cells or CD21+ follicular dendritic cells (Figure 3C-F and supplemental Figure 6A). DNMT3A was preferentially expressed in NBCs as well as CD68+ histiocytes, CD21+ FDCs and CD3+ T cells within the germinal center (supplemental Figure 6B). DNMT3B was expressed in a subset of CD79A+ B cells within the light and dark zones of the GC, a subset of CD3+ T cells and CD21+ FDCs, but not in GC CD68+ histiocytes (supplemental Figure 6C). Altogether, DNMT1 is the most highly expressed DNA methyltransferase in GC B cells, although DNMT3B is also present (summary in supplemental Table 8).
DNMT1 is highly expressed in GC B cells. (A) Relative transcript abundance of DNMT1, DNMT3A, and DNMT3B as well as the GC B-cell specific transcripts PCNA and BCL6 were measured by QPCR. The y-axis depicts the fold difference of relative expression of each transcript in GC B cells versus NB cells. (B) Immunoblotting with anti-DNMT1 antibody was performed in GC B cells and NB cells as indicated, using actin as loading control. (C-F) Immunohistochemical double-staining in reactive human tonsillar tissue delineates expression of DNMT1 (dark purple nuclear staining), (C) in the germinal center CD79A+ (membranous brown staining) B cells and not in (D) CD68+ (membranous brown staining) tingible-body macrophages or (E) IgD+ (membranous brown staining) NB cells, or (F) CD3+ (membranous brown staining) T cells.
DNMT1 is highly expressed in GC B cells. (A) Relative transcript abundance of DNMT1, DNMT3A, and DNMT3B as well as the GC B-cell specific transcripts PCNA and BCL6 were measured by QPCR. The y-axis depicts the fold difference of relative expression of each transcript in GC B cells versus NB cells. (B) Immunoblotting with anti-DNMT1 antibody was performed in GC B cells and NB cells as indicated, using actin as loading control. (C-F) Immunohistochemical double-staining in reactive human tonsillar tissue delineates expression of DNMT1 (dark purple nuclear staining), (C) in the germinal center CD79A+ (membranous brown staining) B cells and not in (D) CD68+ (membranous brown staining) tingible-body macrophages or (E) IgD+ (membranous brown staining) NB cells, or (F) CD3+ (membranous brown staining) T cells.
Decreased expression of DNMT1 interferes with formation of GCs in mice
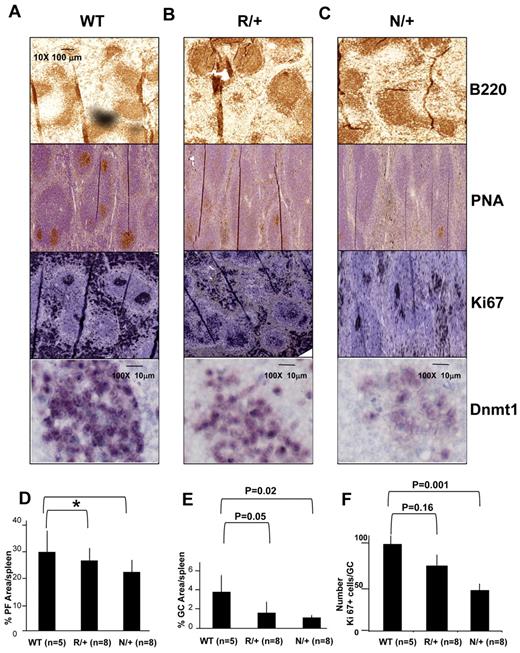
To determine the functional relevance of DNMT1 in GC B cells we examined GC formation in animals with reduced expression of Dnmt1. Dnmt1 knockout is embryonic lethal, but animals with reduced Dnmt1 are viable. We used the Dnmt1 R/+ and Dnmt1 N/+ mice, which express ∼ 60% or ∼ 30% of Dnmt1 compared with wild type, respectively.28 Dnmt1 mRNA abundance was confirmed using Q-PCR on splenic mononuclear cells (supplemental Figure 7). GCs were induced by injecting sheep red blood cells in Dnmt1 R/+ (n = 8), Dnmt1 N/+ (n = 8), and wild-type (WT) animals (n = 4) and spleens analyzed 10 days later. B220 staining indicated that the overall architecture of primary splenic lymphoid follicles was not perturbed (Figure 4A-C). However, GCs were progressively diminished in size and abundance in R/+ and N/+ animals, respectively, as demonstrated by staining for peanut agglutinin (PNA, a GC marker) and Ki67. We also demonstrated lower level of Dnmt1 protein expression in hypomorphic animals using IHC staining with anti-DNMT1 antibody (Figure 4A-C bottom panels). All sections were stained and developed under exactly similar conditions allowing semiquantitative assessment of the protein level. A more quantitative GC assessment was obtained by scanning densitometry of entire spleen sections. This analysis revealed that while the size of the primary lymphoid follicles was only slightly decreased in R/+ and N/+ animals (Figure 4D), GCs were markedly smaller, with more significant decrease in N/+ (1.2% splenic area) than in R/+ (1.6%) animals versus 4% for wild type animals (P = .02 and .05, respectively, t test; Figure 4E). We also calculated the number of Ki67+ cells within each GC to determine the number of proliferating cells. WT animals had on average 100 Ki67+ cells per GC cross-section, while R/+ animals had 80 and N/+ animals had 55 Ki67+ cells (P = .16 and .001, t test Figure 4F). To further confirm these findings we determined the relative number of splenic GC B cells by flow cytometry. Splenic mononuclear cells were isolated and stained with 7AAD, B220, FAS, and GL7. 7AAD positive cells were gated out, and the relative numbers of GL7+/FAS+ germinal center cells within B220+ B-cell gate were quantified. This procedure identified 7.11% GC B cells in WT, 2.07% GC B cells in R/+ and 1.85% GC B cells in N/+ out of all B220+ B cells (statistically significant difference between WT and R/+ at P = .003, t test; and WT and N/+ at P = .0002, t test; Figure 5A-B).
GC formation is impaired in Dnmt1 hypomorphic mice. (A-C) C57BL/6 wild type (WT; A), or Dnmt1 R/+ (B) and Dnmt1 N/+ (C) were immunized with SRBC to induce GC formation. Animals were killed on day 10 and formalin fixed paraffin embedded splenic tissue was stained with anti-B220, PNA, anti-Ki67, and anti-Dnmt1. (D) Quantitative imaging of spleen tissue based on immunohistochemistry staining from Figure 3A revealed no significant difference in the size of primary follicles (*P = .18 and .52, t test), but decreased size of the GCs (E) and the number of proliferating cells per GC (ki67) in the Dnmt1 hypomorph animals (F). The numbers of animals per condition are shown in parenthesis.
GC formation is impaired in Dnmt1 hypomorphic mice. (A-C) C57BL/6 wild type (WT; A), or Dnmt1 R/+ (B) and Dnmt1 N/+ (C) were immunized with SRBC to induce GC formation. Animals were killed on day 10 and formalin fixed paraffin embedded splenic tissue was stained with anti-B220, PNA, anti-Ki67, and anti-Dnmt1. (D) Quantitative imaging of spleen tissue based on immunohistochemistry staining from Figure 3A revealed no significant difference in the size of primary follicles (*P = .18 and .52, t test), but decreased size of the GCs (E) and the number of proliferating cells per GC (ki67) in the Dnmt1 hypomorph animals (F). The numbers of animals per condition are shown in parenthesis.
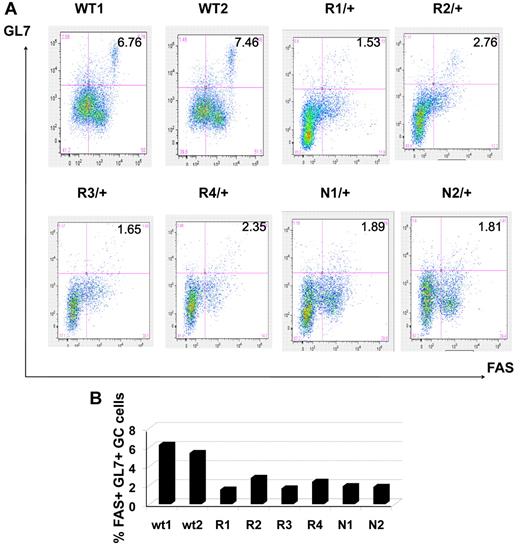
Reduction in the number of GC B cells in immunized Dnmt1 hypomorphic mice. (A) Mononuclear splenocytes were purified 10 days after immunization with SRBC from WT, Dnmt1R/+ and Dnmt1N/+ mice and stained with 7-AAD/ B220/GL7/FAS. The percentage of GL7+ FAS+ cells reflects the abundance of GC B cells in the various animals. (B) The results from panel A are represented as a histogram depicting the percentage of GC B cells for each of the mice analyzed by flow cytometry.
Reduction in the number of GC B cells in immunized Dnmt1 hypomorphic mice. (A) Mononuclear splenocytes were purified 10 days after immunization with SRBC from WT, Dnmt1R/+ and Dnmt1N/+ mice and stained with 7-AAD/ B220/GL7/FAS. The percentage of GL7+ FAS+ cells reflects the abundance of GC B cells in the various animals. (B) The results from panel A are represented as a histogram depicting the percentage of GC B cells for each of the mice analyzed by flow cytometry.
DNA methyltransferase activity is required for GC formation
To determine more globally how crucial DNMTs and DNA methylation are for GC formation, we examined the impact of the DNMT inhibitor decitabine on these structures in vivo. GC formation was induced by sheep red blood injection in 20 C57/BL6 mice, followed 3 days later by daily intraperitoneal treatments with 15 mg/m2 and 30 mg/m2 decitabine (n = 5 mice each group) of decitabine versus vehicle (n = 5 mice) or water only (n = 5 mice) for 7 days, after which the mice were killed and the spleens examined for GC formation. Splenic architecture was preserved in all animals with good delineation of the red and white pulp. Primary follicles were also intact, as highlighted by staining with B220 antibodies (Figure 6A-D). However, GCs were completely absent in spleens from both the 15 mg/m2 and 30 mg/m2 decitabine treated animals as judged by PNA immunohistochemistry (Figure 6C-D). There was also complete absence of proliferating B cells within lymphoid follicles by Ki67 staining (Figure 6) and a marked decrease in Dnmt1 positive cell foci in the spleens of decitabine treated animals with IHC staining of the spleens (supplemental Figure 8). Because DNMT1 is implicated in DNA replication39 and is required for maintaining bulk DNA methylation15 we wondered whether its suppression of GC simply reflected global suppression of cellular proliferation throughout all dividing tissues. Further examination of spleens revealed the presence of scattered Ki67 positive cells in the splenic white pulp within the peri-arteriolar lymphoid sheaths, corresponding to CD3+ T cells, and were similar to control mice (data not shown). Intestinal epithelium is also highly proliferative. A strong KI67 signal was observed in the intestinal crypts of both the control and decitabine treated animals, indicating that proliferation was preserved in this tissue, although the architecture of the crypts was slightly distorted (supplemental Figure 9). Collectively the data indicate that formation of GCs is exquisitely dependent on DNMT activity.
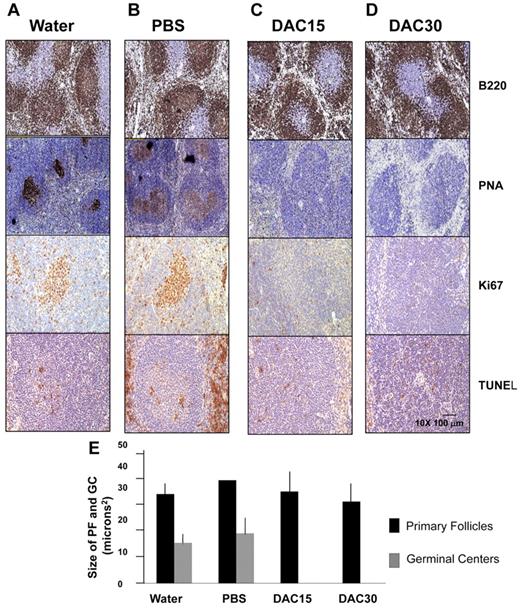
Decitabine treatment abrogates formation of GCs in mice. Twenty C57/BL6 mice were immunized with SRBCs and subjected to daily intraperitoneal injections of water (A), PBS solution (ie, vehicle, B), 15 mg/m2 decitabine (C) or 30 mg/m2 decitabine (D). Formalin fixed paraffin embedded splenic tissue recovered at day 10 was examined by immunohistochemistry for B220, PNA, Ki67 and TUNEL. (E) The histogram represents the quantitative assessment of primary follicle and GC size as assessed by immunohistochemistry staining in all animals.
Decitabine treatment abrogates formation of GCs in mice. Twenty C57/BL6 mice were immunized with SRBCs and subjected to daily intraperitoneal injections of water (A), PBS solution (ie, vehicle, B), 15 mg/m2 decitabine (C) or 30 mg/m2 decitabine (D). Formalin fixed paraffin embedded splenic tissue recovered at day 10 was examined by immunohistochemistry for B220, PNA, Ki67 and TUNEL. (E) The histogram represents the quantitative assessment of primary follicle and GC size as assessed by immunohistochemistry staining in all animals.
DNMT1 deficiency induces DNA damage in GC B cells in vivo
Ha et al reported that DNMT1 is involved in dsDNA breaks repair by interacting with PCNA and 9-1-1 complex,40 and is recruited to sites of DNA damage. Because DNMT1 is abundant within GC B cells, which are undergoing genetic recombination we considered that DNMT1 might be required for DNA damage repair in these cells. Spleen sections from WT and N/+ mice induced with sheep red blood cells and killed at day 10 were stained with antibodies against phospho-H2AX, a marker of DNA damage, and then examined by immunofluorescence microscopy. Spleen sections were then scanned with the MIRAX 1.12.22.1 software to identify clusters of positive GC B cells. This analysis revealed an 8-fold increase in H2AX positive B cells within the GCs formed in the N/+Dnmt1 animals (Average number of H2AX positive B cells is greater in N/+Dnmt1 animals vs WT, P = .0005, t test; Figure 7). Therefore Dnmt1 hypomorphic GC B cells show evidence of increased levels of DNA damage in vivo, suggesting that DNMT1 plays a role in genomic stability during affinity maturation.
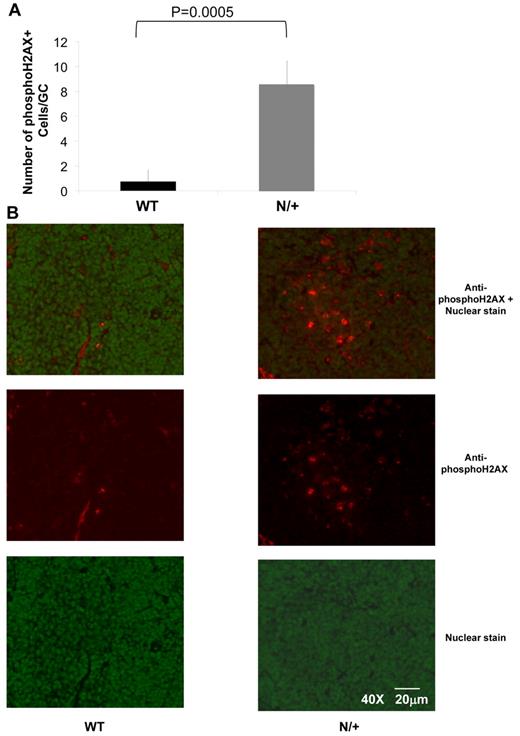
Increased H2AX phosphorylation in the GCs of Dnmt1 hypomorphic mice. (A) The number of phosphoH2AX positive cells per GC was counted in WT or Dnmt1N/+ mice at day 10 after immunization with SRBC based on immunofluorescent staining of splenic sections (B) Representative images from the immunofluorescent staining of WT and Dnmt1N/+ hypomorphic mice using a nuclear stain (green) and anti-phosphoH2AX monoclonal antibody (red), (top: overlay; middle: anti-phosphoH2AX antibody; and bottom: nuclear stain only).
Increased H2AX phosphorylation in the GCs of Dnmt1 hypomorphic mice. (A) The number of phosphoH2AX positive cells per GC was counted in WT or Dnmt1N/+ mice at day 10 after immunization with SRBC based on immunofluorescent staining of splenic sections (B) Representative images from the immunofluorescent staining of WT and Dnmt1N/+ hypomorphic mice using a nuclear stain (green) and anti-phosphoH2AX monoclonal antibody (red), (top: overlay; middle: anti-phosphoH2AX antibody; and bottom: nuclear stain only).
Discussion
In this report we examined the nature of promoter and global DNA methylation patterning occurring in GC B cells and the related question of how and whether DNMTs contribute to establishing the GC B-cell phenotype. We observe a significant number of loci undergoing differential methylation in GC B cells when compared with NBCs. Shifts in DNA methylation patterning have been observed in embryonic stem cell differentiation and during lineage commitment of hematopoietic precursors.16 Our data indicate that DNA methylation patterning continues to evolve in later stages of differentiation and can still undergo significant changes. It was notable that the predominant change in methylation in GC B cells was in the direction of hypomethylation as shown by both HELP and LUMA assays. Presumably, DNA methylation patterning changes are induced by transcription or chromatin remodeling factors that are up-regulated on T-cell dependent B-cell activation.
Expression and functional activation of AICDA defines the GC B-cell phenotype and mediates immunoglobulin affinity maturation. AICDA is also a plausible candidate to rapidly hypomethylate genes in GC B cells. AICDA can demethylate critical promoters required for the induced pluripotent state in a cell-cycle independent manner.21 DNA methylation profiling of AICDA wild type and deleted primordial germ cells indicated that loss of AICDA was associated with relative hypermethylation compared with wild type cells.23 Moreover the cytosine deaminase Apobec1 (a family member of AICDA), was recently shown to mediate site-specific demethylation in cooperation with the TET1 hydroxymethyltransferase in the central nervous system.41
A role for AICDA in GC B-cell hypomethylation is suggested by several lines of evidence presented herein: (1) hypomethylated genes were enriched for RGYW somatic hypermutation hotspot sequences, (2) hypomethylated genes were enriched for the somatic hypermutation associated CAGGTG E-box like motifs, (3) hypomethylated genes were enriched for bona fide AICDA target genes as shown by ChIP-sequencing, and (4) hypomethylated AICDA targets were also significantly more likely to be highly expressed in GC B cells versus NBCs. The overall correlation with hypomethylation and gene expression was weak, consistent with the observation that AICDA does not necessarily only target highly active genes, but also those that are poised for expression.5 Moreover, hypomethylation of genes may not necessarily be associated with hyperactivation of genes. For example, we observed that in GC B cells, histone 3 lysine 27 trimethylation is inversely correlated with DNA methylation and is associated with repression.11 Along these lines we also observe that the BCL6 transcriptional repressor associates preferentially with hypomethylated promoters (P < .0001, Wilcoxon test, based on known BCL6 targets8 ). It is not entirely clear how AICDA is recruited to gene regulatory regions, although recruitment by the bHLH transcription factor TCF3E2A is implicated in this process and enrichment in E-box binding sites in GC B-cell hypomethylated genes is consistent with involvement by this factor.38,42 AICDA was also shown to be recruited to stalled RNA polymerase II in association with replication protein A (RPA5 ). Collectively, the data suggest that AICDA contributes to epigenetic programming of the GC B cell by either directly or indirectly mediating CpG demethylation, a notion that warrants further investigation.
It is notable that DNA methylation patterning was more heterogeneous in GC B cells than NBCs. This observation is not related to the purity of cell fractions since these samples were verified by flow cytometry. Variance in DNA methylation patterning in GC B cells may be a consequence of their biologic and phenotypic heterogeneity, since CD77+ selection captures GC B cells, but does not assure isolation of cells at exactly same stage of affinity maturation or commitment to terminal differentiation. In addition, we hypothesize that variance of methylation patterning may reflect the massive clonal expansion and rapid proliferative rate of GC B cells, which may reduce the fidelity of maintenance methylation by DNMT1 or lead to the stochastic acquisition of de novo methylation. Heterogeneity may also go hand in hand with the variability with which AICDA mediates cytosine deamination throughout the genome. Genetic variability is a desired and inherent property of AICDA-mediated affinity maturation and allows diverse clones with different mutations in their immunoglobulin loci to arise, but by the same token can lead to lymphomagenesis when oncogenes such as BCL6 are affected. Epigenetic variability is associated with states of increased cell turnover, which may be further enhanced by the actions of AICDA in GC B cells. It is thus intriguing to postulate that the combination of genetic and epigenetic heterogeneity, as a result of clonal variance among proliferating cells in GC B cells might jointly contribute to their malignant transformation.
Another, more speculative reason that hypomethylation may predominate in GC B cells might relate to the increased proliferative rate of these cells and failure to maintain methylation during rapid replication. DNA methylation is produced and maintained by DNMTs and it is clear that among these enzymes, DNMT1 plays an essential role in GC formation. We observed that DNMT1 is strongly induced in GC B cells, that Dnmt1 hypomorph mice displayed a dose dependent impairment in GC formation, and that treatment with the DNMT inhibitor decitabine abrogated GC formation completely. The inefficient methylation maintenance scenario seems less likely to be a major contributor to the GC B-cell hypomethylation because we show that hypomethylation is not a generalized phenomenon (by LC-ESI-MS/MS) but seems instead to be relatively restricted to gene regulatory regions. It is logical to assume that at least part of the reason DNMT1 is up-regulated in GC B cells is to support methylation during rapid cell turnover. However, numerous studies demonstrate that DNMT1 also plays a role in supporting DNA replication and repair (eg, Unterberger et al,39 Ha et al40 ). The fact that phospho-H2AX staining was increased in the nuclei of residual GC cells in Dnmt1 hypomorphic mice is consistent with previously published data, including a report by Unterberger et al revealing that DNMT1 knockdown induces DNA damage responses and H2AX phosphorylation.39 One of the factors required to target DNMT1 to hemimethylated DNA is UHRF1.30 UHRF1 also recruits DNMT1 to sites of pericentromeric heterochromatin replication and can regulate its stability through ubiquitylation.43,44 It would stand to reason that rapidly cycling GC B cells would require high UHRF1 activity. Along these lines we find that the UHRF1 family member UHRF2 is one of the most differentially hypomethylated and expressed genes in GC B cells, and in fact we observed that UHRF1 and UHRF1BP (UHRF1 binding protein) are also strongly up-regulated in GC B cells (not shown). DNMT1 may thus carry out multiple functions in support of the GC phenotype including DNA replication, chromatin condensation/decondensation and maintenance of genomic DNA methylation. It was recently shown that DNMT1 and UHRF1 were required to maintain somatic progenitor cell function in tissues such as epidermis and was observed to maintain proliferative potential and block differentiation.45 GC B cells seem to be even more sensitive to DNMT1 levels than other somatic tissues given their proliferation and DNA repair requirements. In certain ways GC B cells seem to reflect or mimic progenitor cell functions. These are proliferative cells, which remain locked in their phenotype unless induced to differentiate by specific signals, can give rise to terminally differentiated cell types, and represent the probable origin for most B-cell neoplasms. Up-regulation of DNMT1-UHRF complexes in GC B cells may provide a mechanistic link to explain similarities between these phenotypes.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
The authors thank A. Chalmers for her help and technical expertise.
This research was supported by K08 CA127353 (R.S.), Leukemia & Lymphoma Society Translational Research Program Award LLS 6304-11 (R.S.), and Leukemia & Lymphoma Society SCOR LLS 7017-09 (A.M.). A.M. is also supported by the Chemotherapy Foundation, a Leukemia & Lymphoma Society Scholar Award, and a Burroughs Wellcome Clinical Translational Scientist Award. L.A.G. is supported by CA129831 and CA129831-03S1. F.M. is supported by R01CA138234 and U54CA143798. S.D. is a recipient of an HFSP fellowship. T.J.E. and K.L. were supported by the Swedish Cancer Society.
National Institutes of Health
Authorship
Contribution: R.S. generated samples, performed HELP profiling, Q-PCR, MA, IHC, analyzed data, and wrote the manuscript; L.C. performed mouse experiments; L.T. performed flow cytometry, IHC; M.K., S.D., and M.F. analyzed data; G.B. performed flow cytometry; N.W. analyzed data; M.R. contributed to data analysis and method development; T.C. performed phosphor-H2AX staining; T.E. and K.L. performed LUMA assays; J.F. performed quantitation of IHC; A.V. and L.G. performed LCLC-MS; F.M. analyzed data and supervised the project; O.E. analyzed data and wrote the manuscript; and A.M. supervised the project and wrote the manuscript.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Ari Melnick, MD, Division of Hematology/Oncology, Department of Medicine, Weill Cornell Medical College, 1300 York Ave, New York, NY 10065; e-mail: amm2014@med.cornell.edu.