Recent studies have demonstrated the importance of coagulation factor X (FX) in adenovirus (Ad) serotype 5–mediated liver transduction in vivo. FX binds to the adenovirus hexon hypervariable regions (HVRs). Here, we perform a systematic analysis of FX binding to Ad5 HVRs 5 and 7, identifying domains and amino acids critical for this interaction. We constructed a model of the Ad5-FX interaction using crystallographic and cryo-electron microscopic data to identify contact points. Exchanging Ad5 HVR5 or HVR7 from Ad5 to Ad26 (which does not bind FX) diminished FX binding as analyzed by surface plasmon resonance, gene delivery in vitro, and liver transduction in vivo. Exchanging Ad5-HVR5 for Ad26-HVR5 produced deficient virus maturation. Importantly, defined mutagenesis of just 2 amino acids in Ad5-HVR5 circumvented this and was sufficient to block liver gene transfer. In addition, mutation of 4 amino acids in Ad5-HVR7 or a single mutation at position 451 also blocked FX-mediated effects in vitro and in vivo. We therefore define the regions and amino acids on the Ad5 hexon that bind with high affinity to FX thereby better defining adenovirus infectivity pathways. These vectors may be useful for gene therapy applications where evasion of liver transduction is a prerequisite.
Introduction
Adenovirus (Ad)–based vectors are used frequently for preclinical gene delivery and therapy and have been used in more than 25% of gene therapy clinical trials conducted to date. Although adenovirus serotype 5 is the most commonly used serotype, the human and nonhuman adenovirus families are large, and many of these are being exploited in diverse clinical applications, such as cancer gene therapy and vaccination.1,,–4 However, the use of Ad vectors as gene delivery tools has raised several safety concerns. The importance of such issues was highlighted in the recent STEP trial in which patients were vaccinated against human immunodeficiency virus using an Ad5 gene delivery vector. The trial was terminated because the vaccine did not function as expected, but actually increased infection rates in those patients with preexisting antibodies to Ad5. Together with other adverse events in humans transduced with Ad5,5 this highlights the importance of understanding fundamental aspects of Ad biology
In vitro, the interaction of the Ad5 fiber and the coxsackie and adenovirus receptor (CAR) is the major pathway for Ad cell binding.6,7 Similarly, engagement with integrins by the penton base protein mediates internalization after cell binding.8 Although other candidate receptors for Ad5 have emerged since the interaction with CAR was identified,9,10 the role of these receptors in gene transfer after intravascular gene delivery has not been substantiated.
It is well established that Ad5 predominantly transduces rodent liver after intravascular injection,11 however mutations of the Ad5 fiber and/or penton show limited effects on liver gene transfer mediated by Ad5 (reviewed in Nicklin et al12 ). Thereafter, it was suggested that a KKTK motif in the Ad5 fiber shaft was responsible for interaction with cellular heparan sulfate proteoglycans (HSPGs),13,–15 based on early in vitro evidence of Ad5 interactions with HSPGs.16,17 However, mutation of KKTK renders the fiber inflexible, preventing Ad5 from internalization through steric hindrance.18,19 In 2005, Shayakhmetov and colleagues20 reported the interaction of the Ad5 fiber knob with serum-derived factors, specifically coagulation factor (F) IX and complement 4 binding protein (C4BP). Subsequently, the vitamin K–dependent coagulation factors FX, FIX, FVII, and protein C (PC) were shown to enhance transduction of HepG2 hepatoma cells in vitro.21 However, only FX could “rescue” liver transduction in warfarin anticoagulated mice.22 This was demonstrated for both Ad5 and an Ad5 vector ablated in its capacity to bind CAR21,–23 highlighting the pivotal importance of FX in mediating hepatocyte transduction by Ad5 in vivo.
Despite the initial assumption that the interaction of FX occurred via the Ad5 fiber,20 3 recent studies have highlighted the role of a high-affinity interaction of FX with Ad5 hexon in FX-mediated liver gene transfer by Ad5.22,24,25 This is a paradigm shift for adenovirus biology because the hexon was previously thought to only represent a structural component of the virion particle, although it is also considered to be the major site of antigenicity on the Ad5 capsid.4,26,27 We reported the high-affinity interaction of the Gla domain of FX with the hexon and cryoelectron microscopy at 23Å resolution pinpointed the binding to the exposed hypervariable regions (HVRs). This was confirmed using a chimeric adenovirus possessing the entire Ad5 capsid, except the HVRs that were derived from Ad48, a non-FX binding subspecies D virus.22 Working at 40 Å resolution, a subsequent study suggested the FX density was localized to HVR3, 5, and 7 and a virus bearing a large peptide insertion in hexon HVR5 showed markedly reduced FX binding in vitro and failed to deliver a transgene to hepatocytes in vivo.24 A further study demonstrated that liver gene transfer was reduced by insertion of a peptide into HVR5 of the Ad5 hexon.25
To date, no studies have systematically modeled the interaction at sub-40Å resolution and performed defined mutagenesis studies to ascertain which domains and amino acids are integral to the high-affinity interaction with FX. A definitive analysis of FX binding to hexon is critical for both adenovirus biology, its clinical utility and safety, as well as for engineering strategies for future applications of gene-based therapeutics. Here, we generate mutant adenoviral vectors devoid of coagulation factor binding. We show the substantial effect these mutations have on FX binding, cell tethering, and in vivo liver gene transfer.
Methods
Cell lines
Human embryonic kidney HEK293 cells (ATCC) were cultured in Dulbecco modified Eagle medium (DMEM; BioWhittaker) supplemented with 2 mM l-glutamine (Invitrogen), 10% fetal calf serum (FCS; PAA Laboratories), and 1 mM sodium pyruvate (Sigma-Aldrich). Human ovarian carcinoma SKOV3 cells (National Institutes of Health [NIH]) were cultured in RPMI 1640 media (Invitrogen) supplemented with 2 mM l-glutamine, 10% FCS, and 1 mM sodium pyruvate. Human cervical carcinoma HeLa cells were cultured in DMEM supplemented with 2 mM l-glutamine, 10% FCS, and 1 mM sodium pyruvate. Cell lines were maintained at 37°C and 5% CO2.
Modeling the Ad5 hexon-FX interaction
Putative contact residues in hexon were defined by computational analysis of a previously determined high-resolution hexon-FX model calculated by cryo-electron microscopy (cryo-em) and fitting of crystallographic coordinates for hexon and modeled coordinates for FX and hexon.22 These previously published data were generated as follows: briefly frozen-hydrated preparations of both unlabeled Ad5 and Ad5 decorated with FX were imaged in a JEOL 1200EX transmission electron microscope (JEOL). Contrast transfer function correction was applied,28 and 3-dimensional image reconstruction was performed using the polar-Fourier transform method (PFT/EM3DR).29,30 Reconstructions for Ad5 and Ad5-FX were calculated at 26 and 23 Å resolution, respectively, as measured using the bresolve program.31 High-resolution coordinates were docked to the low-resolution cryo-em data using University of California, San Francisco (UCSF) Chimera32 and Situs.22,33 The crystal structure of Ad5 hexon (PDB ID 1P3034 ) was docked into the unlabeled Ad5 reconstruction at each of the 4 possible unique positions within the asymmetric unit of adenovirus. To position FX within our model, a difference map was calculated by subtracting the Ad5-FX reconstruction (filtered to 26 Å resolution) from the unlabeled Ad5 structure. The difference map had sufficient density to permit fitting of coordinates at 3 of the 4 hexon positions. A published high-resolution model was used as there are no crystallographic structures available for FX.35 The docked hexon and FX coordinates were then integrated into single pdb files for each of the 3 models using UCSF Chimera. Interchain contact residues were identified in each model using the CCP4 program contact36 and visualized using UCSF Chimera.
Vector construction
Ad5 HVR swap modifications were generated by amplifying HVR5 and 7 regions from plasmids containing the entire viral genomes of human serotypes 5 and 26 (see supplemental methods for additional details, available on the Blood website; see the Supplemental Materials link at the top of the online article). All hexon point mutant adenoviruses were generated using polymerase chain reaction (PCR) site-directed mutagenesis. PCR fragments incorporating the desired mutations were cloned into pSC-B using the Strataclone Blunt PCR cloning kit (Stratagene) according to the manufacturer's instructions. Modified hexon fragments were then excised by digesting pSC-BHex with BamHI and NdeI and subcloning fragments into pBR.dbb.PmeI-SandI.4 Primers 2-kb DIR: (5′-ACAGTGGAAAGGTCGACGC-3′) and 2-kb REV: (5′-ACTTGACTTTCTAGCTTTCC-3′) were used to amplify a 1.95-kb fragment of the left flanking region of the Ad5 hexon. The 1.95-kb fragment was excised using NdeI and cloned into pBR plasmids containing modified hexon fragments to generate shuttle plasmids for homologous recombination. Adenoviral vectors used in this study were based on the AdEasy, E1/E3 deleted Ad5 adenoviral vector system (Stratagene). PacI-digested pAdeasy-1 plasmid, and PmeI-digested pShuttle-CMV-lacZ were used to generate pAd5CMVlacZ by homologous recombination in BJ5183 bacteria cells. All pBR-based plasmids containing hexon mutants and the 1.95-kb fragment were digested by EcoRI and recombined with AsiSI-digested pAd5CMVlacZ in BJ5183 bacteria cells. All cloning steps were confirmed by sequencing using a 3730 DNA analyzer (Applied Biosystems). Nomenclature used to designate HVR-chimeric Ad5 vectors in previous manuscripts4,22 Ad5HVRxx(yy), where xx is the Ad serotype of the exchanged HVRs and yy represents the specific HVRs that are exchanged, has been adapted to the studies in this manuscript Ad5HVRyy(Adxx). For example, Ad5HVR481 in the previous manuscripts would be designated as Ad5HVR1(Ad48) in this study.
Vector amplification
PacI-digested adenoviral genomes were transfected into HEK293 cells in a 6-well plate using Lipofectamine 2000 (Invitrogen) according to the manufacturer's instructions. Cells and media were recovered when viral foci were observed 7 to 10 days posttransfection. Viral particles (VP) were amplified as described.37 Adenoviral particle titers were calculated by determining protein concentrations using the micro-bicinchoninic acid (BCA) Protein Assay kit (Thermo Scientific) followed by titer calculations using the formula 1 μg protein = 4 × 109 VP as well as end point dilution assays for quantification of plaque forming units (pfu).37
Surface plasmon resonance analysis
Surface plasmon resonance (SPR) was performed using a Biacore T100 (Biacore) as described.21 FX was covalently immobilized onto the flowcell of a CM5 biosensor chip by amine coupling according to the manufacturer's instructions. Subtracted sensorgrams were generated by subtracting the signal from a surface subjected to a blank amine immobilization. Virus in 10 mM N-2-hydroxyethylpiperazine-N′-2-ethanesulfonic acid (HEPES), pH 7.4, 150 mM NaCl, 5 mM CaCl2, 0.005% Tween 20 was passed over the chip at a flow rate of 30 μL/minute. Sensorchips were regenerated between virus application by injection of 10 mM HEPES, pH 7.4, 150 mM NaCl, 3 mM ethylenediaminetetraacetic acid (EDTA), 0.005% Tween 20.
Analysis of virus cell binding capacity
Adenovirus binding experiments were performed in a 24-well format with 2 × 105 SKOV3 cells/well. Cells were cooled to 4°C for 30 minutes and washed twice with cold phosphate-buffered saline (PBS) before adding 1000 VP/cell of Ad5 or hexon-modified adenoviruses in the presence or absence of FX in serum-free media. FX (Haematological Technologies) was used at a final concentration of 1 IU/mL. Cells were incubated with adenovirus for 1 hour at 4°C, washed twice with PBS, then mechanically dislodged from the plate into 200 μL PBS. DNA was extracted from cell pellets using the QIAamp DNA mini kit (QIAGEN) according to the manufacturer's instructions. Viral genomes in 100 ng DNA were quantified by quantitative PCR analysis (7900HT Sequence Detection System; Applied Biosystems) using Power SYBR Green PCR Master mix (Applied Biosystems) and 0.2 μM hexon primers (5′-CGCGGTGCGGCTGGTG-3′ and 5′-TGGCGCATCCCATTCTCC-3′).
In vitro evaluation of cell transduction
Transduction experiments were performed in a 96-well format with 5 × 104 SKOV3 or HeLa cells/well. Cells were infected with Ad5 or hexon mutant adenoviruses at a dose of 1000 VP/cell in serum-free medium in the presence or absence of FX (1 IU/mL). Infected cells were incubated for 3 hours at 37°C, washed with PBS, and maintained until harvesting 48 hours posttransfection. β-Galactosidase activity was quantified using Tropix Galacton Plus and Tropix accelerator II (Applied Biosystems) according to the manufacturer's instructions. β-Galactosidase activity was quantified using a Wallac VICTOR2 (Perkin Elmer Life and Analytical Sciences). Protein concentrations were calculated using BCA assays (Thermo Scientific). Values are expressed as relative light units (RLU)/mg of protein.
In vivo analysis of liver transduction
All animal procedures were approved by the UK Home Office. Male MF-1 mice, aged between 7 and 9 weeks (Harlan), were used for in vivo experiments. Clodronate liposomes (http://clodronateliposomes.org; 200μL) were injected into the tail-vein 24 hours before adenovirus administration. VP (1010) in 100 μL PBS were administered via tail vein injection. All animals were killed 48 hours later. β-Galactosidase transgene expression was analyzed using a β-Gal enzyme-linked immunosorbent assay (ELISA) kit (Roche) according to the manufacturer's instructions. Liver lobes that had been fixed in 2% paraformaldehyde for 16 hours at 4°C were stained for β-galactosidase activity in 5-bromo-4-chloro-3-indolyl-β-D-galactoside (X-gal) staining solution [0.1 M phosphate buffer, pH 7.3, 2 mM MgCl2, 5 mM K3F3(CN)6, 5 mM K4Fe(CN6)6, and 1 mg/mL X-gal] at 37°C overnight.
Results
Modeling of the FX:hexon interaction and identification of FX contact points
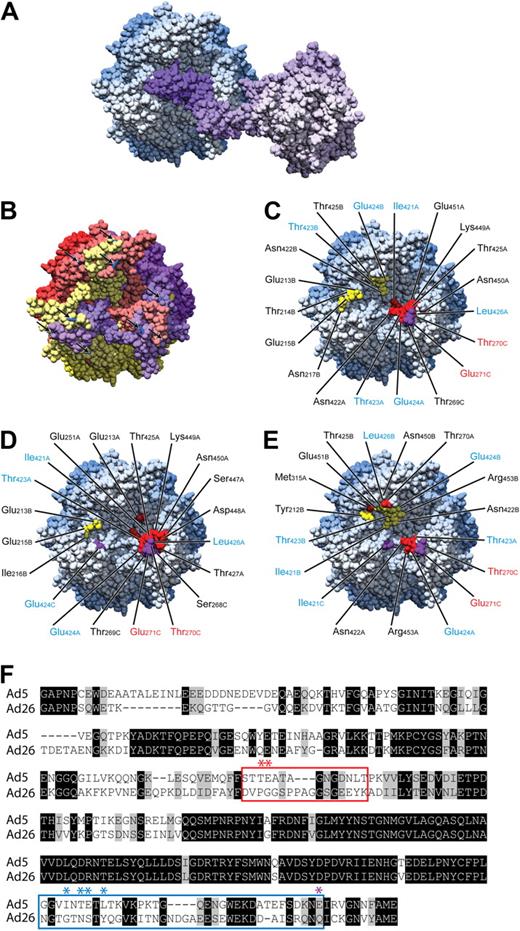
In our previous study,22 we showed that FX bound to the Ad5 hexon HVRs by use of Ad5HVR48, a vector that had all Ad5 HVRs converted to those of Ad48, a virus that did not bind FX. To localize FX binding in more detail, we constructed a high-resolution model of the Ad5-FX interaction. Cryo-em was used to determine low-resolution structures for Ad5 virions complexed with FX and also unlabeled particles. Fitting high-resolution coordinates for Ad5 hexon34,38 and a molecular model of the FX structure35 to these maps led to the synthesis of models of the complex based on density at 3 of the 4 possible unique capsomeres (hexon trimers) within the virion's asymmetric unit (Figure 1C-E). The atomic resolution of 1 representative model viewed perpendicular to the capsid outer surface is shown (Figure 1A). Hexon is colored blue, and FX is colored purple. These models allowed us to determine possible contact residues in hexon that might be important in binding to FX. The list of putative contact residues is not exhaustive owing to the presence of flexible loops in hexon that are not resolved in the published crystallographic coordinates34 (Figure 1B). These models were used as the basis for subsequent mutagenesis to identify residues in the Ad5 hexon critical for FX interaction (Figure 1C-E). We thereby identified HVR5 and 7 as potentially key domains for FX binding. In addition, phylogenetic analysis of the HVR sequences of adenovirus serotypes that bind FX, in comparison to those that did not bind FX, identified an amino acid residue E451 conserved in all binders but replaced by Q in nonbinders.22 E451 is located C terminally to the 4 amino acids mutated in HVR7 (Figure 1F).
Modeling of the Ad5:FX interaction to identify FX contact points in the hexon protein. Models of the interaction between hexon and FX were used to determine candidate contact residues in the Ad5 hexon. (A) An atomic resolution representation of one such model is shown in top view (ie, viewed from the virion exterior). Hexon is shown in blue. FX is shown in purple. (B) Polypeptide chains within the trimeric capsomere are named and colored (A, red; B, yellow; and C, purple) relative to the orientation of FX. Missing regions in the crystal structure of hexon are represented with blue sticks and highlighted with black arrows. (C-E) Contact residues are shown for each of the 3 models calculated and named/colored according to their chain assignment relative to FX. Amino acids targeted for mutagenesis have labels that are colored according to regions identified in panel F. In each panel, hexon is represented such that FX is in approximately the same orientation as in panel C. (F) Amino acid alignment of Ad5 and Ad26 hexon HVR sequences to highlight domains (HVR5, red box; HVR7, blue box) and amino acids targeted for mutagenesis studies (point mutations highlighted by *).
Modeling of the Ad5:FX interaction to identify FX contact points in the hexon protein. Models of the interaction between hexon and FX were used to determine candidate contact residues in the Ad5 hexon. (A) An atomic resolution representation of one such model is shown in top view (ie, viewed from the virion exterior). Hexon is shown in blue. FX is shown in purple. (B) Polypeptide chains within the trimeric capsomere are named and colored (A, red; B, yellow; and C, purple) relative to the orientation of FX. Missing regions in the crystal structure of hexon are represented with blue sticks and highlighted with black arrows. (C-E) Contact residues are shown for each of the 3 models calculated and named/colored according to their chain assignment relative to FX. Amino acids targeted for mutagenesis have labels that are colored according to regions identified in panel F. In each panel, hexon is represented such that FX is in approximately the same orientation as in panel C. (F) Amino acid alignment of Ad5 and Ad26 hexon HVR sequences to highlight domains (HVR5, red box; HVR7, blue box) and amino acids targeted for mutagenesis studies (point mutations highlighted by *).
Substitution of HVR5 or HVR7 from Ad26 diminishes FX binding, FX-mediated cell transduction in vitro, and liver gene transfer in vivo
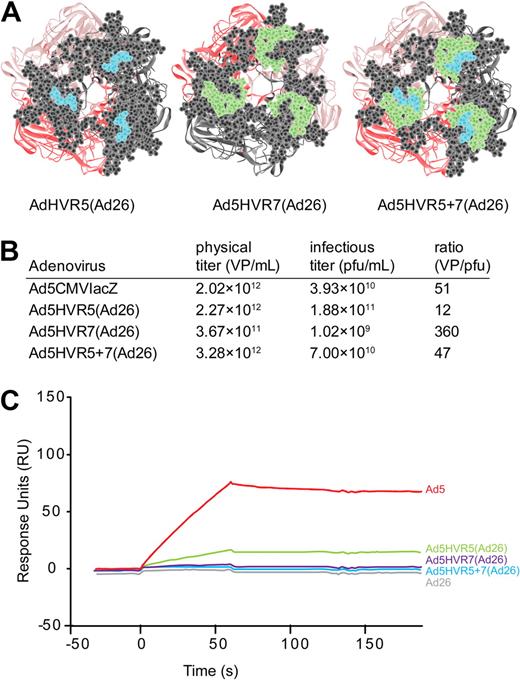
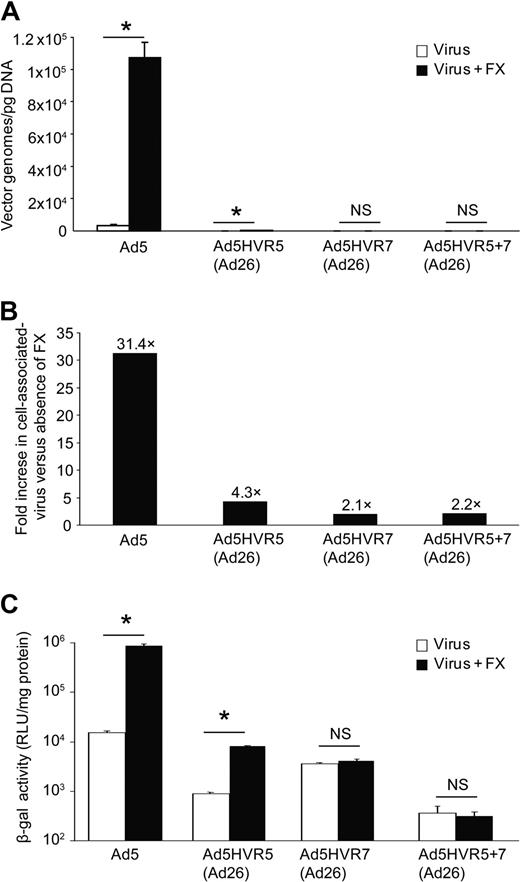
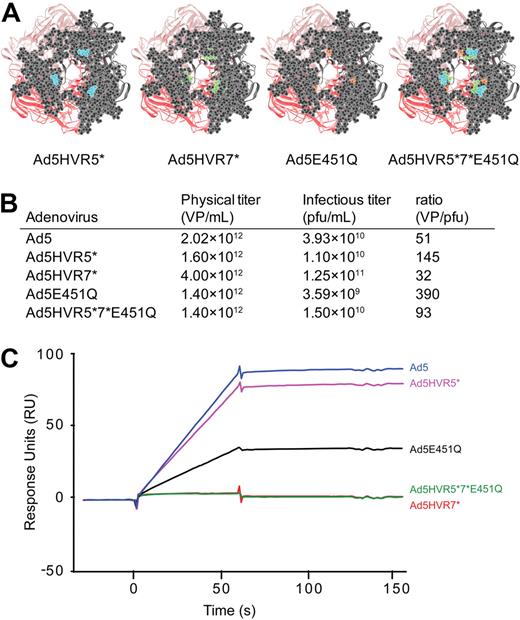
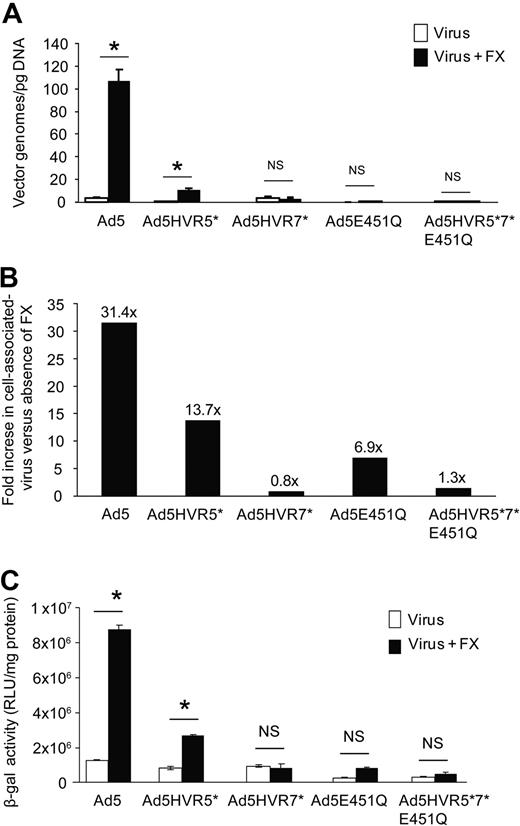
Rather than assess FX interaction by insertion of heterologous peptides, we chose to generate Ad5 vectors with specific HVR domain swaps by engineering HVR5 [Ad5HVR5(Ad26)], HVR7 [Ad5HVR7(Ad26)], or HVR5 and 7 [Ad5HVR5 + 7(Ad26)] substituted with sequences from the non-FX binding serotype Ad26 to determine the contribution of each HVR to FX binding (Figure 2A). All viruses grew efficiently in 293 cells with expected and equivalent titers (Figure 2B). We first assessed direct binding of each virus to FX by SPR. Ad5HVR5(Ad26) showed reduced FX binding compared with parental Ad5 when injected over a FX biosensor chip, while substitution of HVR7 for Ad26 [in virus Ad5HVR7(Ad26)] essentially abolished FX binding by SPR, an effect maintained in the double HVR replacement vector [Ad5HVR5 + 7(Ad26)] (Figure 2C) and comparable to the wild-type Ad26. We next performed cell binding experiments (Figure 3A-B) and transduction experiments (Figure 3C) to assess the effects of HVR substitutions on FX-mediated cell infectivity. In concordance with the SPR data, FX-mediated SKOV3 cell binding was significantly reduced by substitution of HVR5, but was more profoundly reduced by substitution of HVR7 (Figure 3A-B). Moreover, while Ad5HVR5(Ad26) showed a reduced effect of FX on cell binding and transduction compared with Ad5, for both Ad5HVR7(Ad26) and Ad5HVR5 plus 7(Ad26) FX-mediated transduction was abolished (Figure 3C).
Swapping HVR5 and 7 of Ad5 to Ad26 and its effect on FX binding by SPR. (A) Viewlite images of the Ad5 hexon showing HVR5 (blue), HVR7 (green), as well as the remaining HVR amino acids (gray). (B) Virus particle titers and VP/pfu ratios for each virus produced. (C) SPR analysis of virus interaction with immobilized FX (500 RU). Sensorgrams of 1011 VP/mL injected at a flow rate of 30 μL/minute.
Swapping HVR5 and 7 of Ad5 to Ad26 and its effect on FX binding by SPR. (A) Viewlite images of the Ad5 hexon showing HVR5 (blue), HVR7 (green), as well as the remaining HVR amino acids (gray). (B) Virus particle titers and VP/pfu ratios for each virus produced. (C) SPR analysis of virus interaction with immobilized FX (500 RU). Sensorgrams of 1011 VP/mL injected at a flow rate of 30 μL/minute.
In vitro cell binding and transduction of HVR5 and 7 mutant adenoviruses. (A) Cell binding at 4°C to SKOV3 cells in the presence or absence of physiologic concentrations of FX. (B) Fold change in FX-assisted virus association with cell versus absence of FX. (C) SKOV3 cell transduction at 48 hours postinfection after a 3-hour exposure to each virus in the presence or absence of FX in serum-free conditions (P < .01 vs no FX conditions).
In vitro cell binding and transduction of HVR5 and 7 mutant adenoviruses. (A) Cell binding at 4°C to SKOV3 cells in the presence or absence of physiologic concentrations of FX. (B) Fold change in FX-assisted virus association with cell versus absence of FX. (C) SKOV3 cell transduction at 48 hours postinfection after a 3-hour exposure to each virus in the presence or absence of FX in serum-free conditions (P < .01 vs no FX conditions).
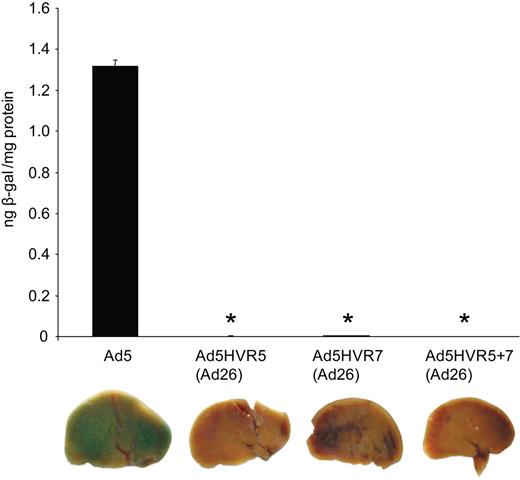
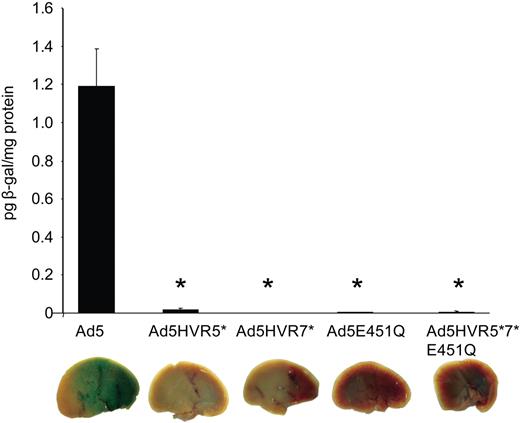
To examine whether ablation of the hexon:FX interaction would manifest in decreased hepatic transduction, MF1 mice were administered intravenously with 1010 VP of the hexon chimeric viruses. Levels of transgene expression (β-galactosidase activity), 48 hours postinjection, were high as expected, however, Ad5HVR5(Ad26), Ad5HVR7(Ad26), and Ad5HVR5 plus 7(Ad26) all showed substantially lower levels of liver transgene expression (Figure 4). Together, these data support the hypothesis that both HVR5 and HVR7 play a critical role in FX binding.
In vivo analysis of Ad5HVR5(Ad26) and Ad5HVR7(Ad26). Male MF-1 mice were injected with 1010 VP of each virus via the tail vein. Mice were killed at 48 hours, and livers were harvested and processed for quantification of β-galactosidase activity and staining (P < .001 vs Ad5).
In vivo analysis of Ad5HVR5(Ad26) and Ad5HVR7(Ad26). Male MF-1 mice were injected with 1010 VP of each virus via the tail vein. Mice were killed at 48 hours, and livers were harvested and processed for quantification of β-galactosidase activity and staining (P < .001 vs Ad5).
Because all hexon mutant viruses contain identical fiber and penton base proteins, we assessed the ability of each to transduce HeLa cells and HepG2 cells in the absence of FX and thus through the CAR-permissive pathway. All mutant viruses efficiently transduced cells, although we noted reductions in infectivity for both viruses containing the HVR5 modifications (supplemental Figure 1). This suggested that both HVR5-containing viruses were potentially deficient in fiber and/or penton base capsid structures. We therefore assessed silver stained gels for virus capsid proteins (supplemental Figure 2). As shown, Ad5HVR5(Ad26) and Ad5HVR5 plus 7(Ad26) both showed profiles consistent with deficient virion maturation. Hence, while both Ad5 HVRs 5 and 7 are important in FX binding, and FX interaction can be eliminated by domain exchanging, Ad26 HVR5 exchange with that of Ad5 potentially prevents virion maturation. In addition, there were several other unidentified protein bands below the hexon band for each Ad with individual point mutations in the hexon.
Point mutagenesis leads to minimally modified Ad5 vectors with abolished liver gene transfer capacity
To interrogate the interaction further and localize critical amino acids in this high affinity interaction, we generated a series of Ad5 vectors containing point mutations at key amino acids within HVR5 (T270P and E271G) and HVR7 (I421G, T423N, E424S, L426Y, and E451Q; Figures 1F, 5A). All point mutant viruses grew to high titers (Figure 5B); CAR-dependent transduction of HeLa and HepG2 cells was efficient for all viruses (supplemental Figure 3), and all virions showed efficient maturation (supplemental Figure 4), indicating that point mutations within HVR5 and HVR7 did not impair CAR-mediated transduction. We observed higher VP/pfu ratios for the Ad5E451Q (7.6× that of Ad5) and Ad5HVR5*7*E451Q (1.8× that of Ad5). The particle:pfu ratios are within the expected ranges for normal individual Ad5 batch variation. SPR analysis revealed a striking effect of some but not all point mutations on FX interactions. The most profound inhibition of FX binding was observed in vectors containing modifications within the HVR7 region (Figure 5C). Interestingly, the introduction of point mutations within HVR5 had a very modest effect on FX binding (Figure 5C). Ad5E451Q bound FX but with lower affinity than HVR5* modification and Ad5CMVLacZ. In vitro cell binding and transduction experiments revealed significantly reduced FX-mediated cell binding and transduction for all mutants compared with Ad5 (Figure 6). However, mutations in the HVR5 region in virus Ad5HVR5* reduced, rather than ablated FX enhancement. Mutations in HVR7 abolished both cell binding and transduction mediated by FX (Figure 6). Importantly, the single point mutation E451Q abolished FX-mediated effects in vitro (Figure 6). When injected intravenously into MF1 mice, all mutant viruses showed substantially decreased levels of hepatic transduction compared with the parental Ad5 vector (Figure 7). Taken together, these studies pinpoint critical amino acids that mediate the FX interaction with the Ad5 hexon.
Effect of point mutagenesis on FX binding by SPR. (A) Viewlite images of mutations in the Ad5 hexon for each virus. (B) Virus titers and VP/pfu ratios for each virus produced. (C) SPR analysis of virus interaction with immobilized FX (500 RU). Sensorgrams of 1011 VP/mL injected at a flow rate of 30 μL/minute.
Effect of point mutagenesis on FX binding by SPR. (A) Viewlite images of mutations in the Ad5 hexon for each virus. (B) Virus titers and VP/pfu ratios for each virus produced. (C) SPR analysis of virus interaction with immobilized FX (500 RU). Sensorgrams of 1011 VP/mL injected at a flow rate of 30 μL/minute.
In vitro analysis of Ad5 hexon point mutants. (A) Cell binding at 4°C to SKOV3 cells in the presence or absence of physiologic concentrations of FX, (*P < .01 vs no FX conditions). (B) Fold change in FX-assisted virus association with cell versus absence of FX. (C) SKOV3 cell transduction at 48-hour postinfection after a 3-hour exposure to each virus in the presence or absence of FX in serum-free conditions (P < .001 vs no FX conditions).
In vitro analysis of Ad5 hexon point mutants. (A) Cell binding at 4°C to SKOV3 cells in the presence or absence of physiologic concentrations of FX, (*P < .01 vs no FX conditions). (B) Fold change in FX-assisted virus association with cell versus absence of FX. (C) SKOV3 cell transduction at 48-hour postinfection after a 3-hour exposure to each virus in the presence or absence of FX in serum-free conditions (P < .001 vs no FX conditions).
In vivo analysis of Ad5 hexon point mutants. MF-1 mice were injected with 1010 VP of each virus via the tail vein. Mice were killed at 48 hours, and livers were harvested and processed for quantification of β-galactosidase activity and staining (P < .001 vs Ad5).
In vivo analysis of Ad5 hexon point mutants. MF-1 mice were injected with 1010 VP of each virus via the tail vein. Mice were killed at 48 hours, and livers were harvested and processed for quantification of β-galactosidase activity and staining (P < .001 vs Ad5).
Discussion
Detailed knowledge of virus infectivity pathways is critical for clinical applications of Ad and for future design of efficient and selective vectors for gene-based therapeutics. Until recently, much of the detail relating to in vivo gene transfer after intravascular delivery of Ad was lacking. Following on from our recent study22 that defined binding of FX to the Ad5 hexon, we now report the major contact points within the hexon HVRs that are critical for this high-affinity interaction. We show an involvement of HVR5 and, more importantly, HVR7 in this interaction. Furthermore, we locate critical amino acid residues in each region that, when mutated, prevent FX binding and associated FX-mediated liver gene transfer. It is interesting to note that SPR analysis on the complete HVR5 and HVR7 domain swaps clearly showed reduced FX association, and this correlated tightly with the observed effects in vitro and in vivo. Surprisingly, we observed virions on silver stains that suggested that exchange of Ad5 HVR5, but not HVR7 for that of Ad26, was incompatible with mature virion assembly, an effect circumvented by defined point mutagenesis instead of complete domain exchange. It is difficult to conclude how HVR modification impacts upon other viral proteins, although it may be due to incompatibilities between the modified hexon and the Ad5-100k scaffolding protein, which is required for hexon trimerization. This in turn may alter hexon interaction with other capsid proteins. However, for the point mutants, SPR demonstrated variable effects on FX binding, while all mutants showed substantial reduction in liver gene transfer. All point mutants were fully functional as evidenced by equal CAR-dependent cell transduction. These data suggest that the domain swaps essentially abolish any effect of FX on Ad5 but the effect on the point mutants is dependent on the specific mutation generated.
A critical issue for gene therapy using intravascular delivery of Ad is the choice of vector system. Because the fiber was thought to be the predominant determinant of cell infectivity, many studies have “pseudotyped” (swapped) fibers from alternate Ad vectors onto the core Ad5 capsid in an attempt to avoid liver transduction (reviewed in Nicklin et al12 ). Clearly, the high-affinity interaction between the Ad5 hexon and FX is maintained in such vectors. Two strategies will therefore avoid this: (1) use of full serotype vectors that possess no binding to FX, as detailed in our previous study,22 or (2) use of Ad5-based vectors with mutated hexon proteins that avoid interaction with FX. Here, we have created minimally mutated vectors where liver gene transfer in mice is eliminated. We used a low viral dose in this study (1010 VP/mouse) as the use of clodronate liposomes increases liver transduction approximately 40-fold.39 It remains unclear as to the involvement of amino acids in the hexon HVRs that are not resolved in the crystal structure in FX binding. From the HVR5 and 7 domain swaps, we can conclude that both regions in their entirety are involved in the high-affinity interaction with FX. In vivo, all mutants show a substantial reduction in FX-mediated liver gene transfer, hence the maintenance of the high-affinity interaction with FX is an important requirement for liver gene transfer in vivo. Recent studies have shown presentation of CAR on red blood cells in human and rats,40,41 hence the in vivo importance of CAR after intravascular delivery likely rests on the interaction of Ad5 with erythrocytes. This hypothesis is supported by our previous work showing CAR interactions are not involved in liver hepatocyte uptake by Ad5.21,22 In an earlier study, we reported that Ad5 vectors with a double point mutation in the Ad5 fiber to block CAR binding abolished agglutination of both rat and human erythrocytes.15 Nevertheless, agglutination is serotype-specific, and it does not take place in murine erythrocytes for Ad5 and is therefore not relevant for this study. Thus, in the future, engineering point mutations in the hexon to ablate FX binding and fiber modifications to remove CAR interactions may show additional utility in vivo.
Modulation of the Ad5:hexon interaction may be important in the development of future gene therapies based on Ad. Emerging evidence highlights the potential of FX in enhancing gene delivery to other cells, including cancer cells,39,42 epithelial cells,43 and vascular endothelial cells (C. Nicol and A.H.B., unpublished, January 2009). While the physiologic and/or pathologic role of these interactions remain to be determined, the high-affinity interaction with FX is likely to have broader relevance than effects on liver gene transfer alone. It is important to note that our previous studies have demonstrated that FX binding to Ad has no/little impact on Ad interaction with Kupffer cells,23 although a recent study showed a small increase in Kupffer cell sequestration of Ad5 in warfarin-treated mice.44 Hence, the vectors we have created in this study will still bind extensively to Kupffer cells. Additional interventions, such as those described recently,44,45 would be required to block Kupffer cell–mediated uptake of Ad5.
In summary, we have identified critical amino acids in the Ad5 hexon that bind FX and created a series of vectors that are devoid of FX binding. This is important information relating to Ad biology and mechanisms of cell infectivity in vitro and in vivo, a key issue that has been deficient despite advancement of Ad to many clinical trials. We propose that minimally modified Ad5 vectors devoid of the high affinity interaction may have broad appeal for gene therapy applications.
An Inside Blood analysis of this article appears at the front of this issue.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
We thank Nicola Britton and Gregor Aitchison at the British Heart Foundation Glasgow Cardiovascular Research Centre (BHF GCRC) for technical assistance, Dirk Spek (Crucell) for technical assistance, and Jort Vellinga (Crucell) for helpful discussions.
This work was supported by the Biotechnology and Biological Sciences Research Council and British Heart Foundation (A.H.B.), and National Institutes of Health grants AI078526, AI066305, and AI066924 (Bethesda, MD; D.B). A.L.P. holds a personal research fellowship from the Caledonian Research Foundation (Edinburgh, United Kingdom). S.N.W. is a recipient of the Philip Gray Fellowship, Katharine Dormandy Trust (London, United Kingdom).
National Institutes of Health
Authorship
Contribution: R.A. performed research and analyzed data; A.C.B., A.L.P., and D.B. performed research; S.N.W. and S.A.N. analyzed data; N.v.R., J.C., J.G., and D.H.B. provided essential reagents; J.H.M. performed research and wrote the paper; A.H.B. supervised the project and wrote the paper; and all authors edited the paper and agreed on the final version.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Andrew H. Baker, British Heart Foundation Glasgow Cardiovascular Research Centre, Division of Cardiovascular and Medical Sciences, 126 University Pl, University of Glasgow, G12 8TA, UK; e-mail: HHab11f@clinmed.gla.ac.ukHH.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal