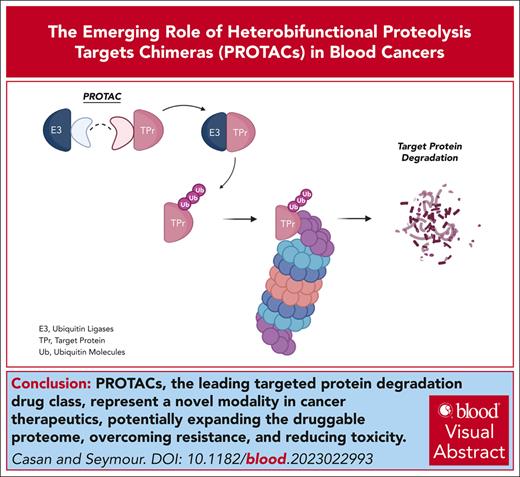
Visual Abstract
Targeted protein degradation (TPD) is a revolutionary approach to targeted therapy in hematological malignancies that potentially circumvents many constraints of existing small-molecule inhibitors. Heterobifunctional proteolysis-targeting chimeras (PROTACs) are the leading TPD drug class, with numerous agents now in clinical trials for a range of blood cancers. PROTACs harness the cell-intrinsic protein recycling infrastructure, the ubiquitin-proteasome system, to completely degrade target proteins. Distinct from targeted small-molecule inhibitor therapies, PROTACs can eliminate critical but conventionally “undruggable” targets, overcome resistance mechanisms to small-molecule therapies, and can improve tissue specificity and off-target toxicity. Orally bioavailable, PROTACs are not dependent on the occupancy-driven pharmacology inherent to inhibitory therapeutics, facilitating substoichiometric dosing that does not require an active or allosteric target binding site. Preliminary clinical data demonstrate promising therapeutic activity in heavily pretreated populations and novel technology platforms are poised to exploit a myriad of permutations of PROTAC molecular design to enhance efficacy and targeting specificity. As the field rapidly progresses and various non-PROTAC TPD drug candidates emerge, this review explores the scientific and preclinical foundations of PROTACs and presents them within common clinical contexts. Additionally, we examine the latest findings from ongoing active PROTAC clinical trials.
Introduction
Scientific advancements have deepened our understanding of cancer pathobiology, catalyzing the development of targeted small-molecule inhibitor (SMI) therapeutics that redefined the standard of care for numerous hematological malignancies. However, the development of structure-based protein targeting molecules is fraught with challenges, and consequently the path from bench to bedside has proven laborious, inefficient, prodigiously expensive, and plagued by a high failure rate.1-4 Moreover, the absence of suitable binding pockets for structure-based targeting renders most of the human proteome effectively “undruggable” with conventional small molecules.5 SMIs are further constrained because their efficacy depends on occupancy-driven pharmacology, necessitating high local drug concentrations and continuous exposure; a requirement that can enhance toxicity and potentially contribute to the emergence of drug resistance.
Targeted protein degradation (TPD) is an alternative strategy to protein inhibition, conceptually enabling complete and sustained eradication of the target protein. TPD therapeutics leverage the cell-intrinsic protein degradation and recycling infrastructure, the ubiquitin-proteasome system (UPS), to achieve target degradation. This mode of action confers important advantages compared with conventional SMIs and, critically, expands the range of targetable proteins. TPD technologies can tractably eliminate many so-called undruggable targets as well as those with acquired resistance to conventional inhibitors. Among several novel protein degrader classes, heterobifunctional proteolysis-targeting chimeras (PROTACs) have emerged as the lead compounds of the TPD era. PROTACs are bivalent molecules composed of 3 functional components: a target protein-binding “warhead” ligand, an E3 ligase–binding ligand, and a linker element that conjoins the 2 ligands. PROTACs facilitate ubiquitination of their target protein, marking it for proteasome-mediated degradation.
In this review, we explore the principles of PROTAC design, their mechanism of action, and the implications for treating hematological malignancies. Additionally, we will examine the existing preclinical and clinical data pertaining to some of the most clinically relevant PROTAC targets currently under development in blood cancer.
The UPS
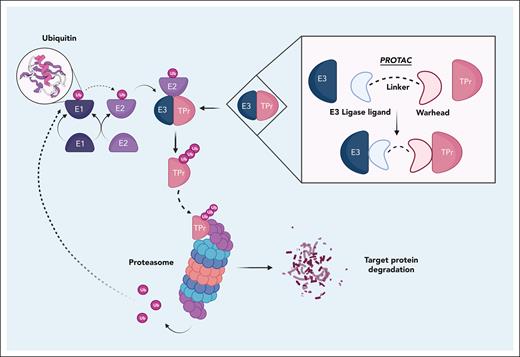
The UPS is an essential cell-intrinsic mechanism for regulating protein homeostasis. Ubiquitin tags mark a protein for degradation by the proteasome 26S subunit in a cascading process involving 3 classes of enzymes (E1-E3; Figure 1). First, ubiquitin-activating (E1) enzymes adenylate ubiquitin before it is translocated to a ubiquitin-conjugating enzyme (E2).6 Ubiquitin-ligases (E3) bind to the target protein, recruiting the target to the E2–ubiquitin complex and catalyzing E2-driven transfer of ubiquitin to the target protein.7-9 Once ubiquitinated, specific proteasomal subunits recognize the tagged protein, triggering protein unfolding and loading into the proteolytic core of the proteasome for degradation.8,9 Deubiquitinating enzymes then liberate the ubiquitin molecules, enabling future reactions.10 Diversity of the UPS enzyme families (especially the E3 ligases, which number >600) provide the substrate specificity to achieve finely regulated protein degradation and, therefore, enormous opportunities for modulation of therapeutic targeting.11 Further detailed reviews of UPS physiology are readily available.12,13
The UPS and PROTAC mechanism of action. The ubiquitin-activating (E1) enzymes adenylate ubiquitin in an ATP-dependent reaction and the ubiquitin molecule is translocated to a ubiquitin-conjugating enzyme (E2). Ubiquitin-ligases (E3) bind to the target protein, recruiting the target to the E2-ubiquitin complex and catalyzing E2-driven transfer of ubiquitin to the target protein. Additional ubiquitin molecules may be attached to form a polyubiquitin chain, with variation in linkage configurations conferring different downstream effects on the protein’s fate. Once ubiquitinated, specific proteasomal subunits recognize the tagged protein via ubiquitin receptors or ubiquitin-associated domains. This initiates the unfolding of the protein and its loading into the proteolytic core of the proteasome, in which it is catalytically degraded. Deubiquitinating enzymes then liberate the ubiquitin molecules, enabling their participation in future ubiquitination reactions. ATP, adenosine triphosphate; Ub, ubiquitin molecules; TPr, target protein.
The UPS and PROTAC mechanism of action. The ubiquitin-activating (E1) enzymes adenylate ubiquitin in an ATP-dependent reaction and the ubiquitin molecule is translocated to a ubiquitin-conjugating enzyme (E2). Ubiquitin-ligases (E3) bind to the target protein, recruiting the target to the E2-ubiquitin complex and catalyzing E2-driven transfer of ubiquitin to the target protein. Additional ubiquitin molecules may be attached to form a polyubiquitin chain, with variation in linkage configurations conferring different downstream effects on the protein’s fate. Once ubiquitinated, specific proteasomal subunits recognize the tagged protein via ubiquitin receptors or ubiquitin-associated domains. This initiates the unfolding of the protein and its loading into the proteolytic core of the proteasome, in which it is catalytically degraded. Deubiquitinating enzymes then liberate the ubiquitin molecules, enabling their participation in future ubiquitination reactions. ATP, adenosine triphosphate; Ub, ubiquitin molecules; TPr, target protein.
PROTACs
The concept of co-opting the UPS to achieve TPD emerged in the 1990s and the first in vitro demonstration of a synthetic PROTAC was published in 2001.14 Early PROTACs used peptidic backbones, which were limited by high molecular weights and modest cell permeability.15 Pure small-molecular PROTACs arose in 2008, improving design flexibility and, thus, the synthesis of clinically viable therapeutic prototypes with improved cellular permeability and pharmacokinetic (PK) properties.16 Additionally, the clinical potential of protein degrader therapeutics had been aptly demonstrated by compounds now termed “molecular glues” such as thalidomide, lenalidomide, and pomalidomide, although the clinical deployment of these agents preceded a detailed understanding of their underlying mechanism of action.17 Recently, considerable investment and effort has generated a multitude of PROTACs, and some have progressed to clinical trial investigation.
PROTAC structure and design principles
As mentioned, PROTACs are tripartite heterobifunctional molecules in which a target protein binding ligand (warhead) is connected to an E3 ligase–binding ligand by a linker component (Figure 1). The PROTAC brings the target protein proximate to the E3 ligase, leading to the formation of a ternary complex between the target, the PROTAC, and the E3 ligase, and the resultant ubiquitination marks the protein for proteasome-mediated destruction.18
Although >600 E3 ligases have been identified in humans, 2 E3 ligases, von Hippel–Lindau (VHL) and cereblon, predominate in existing PROTAC designs. These E3 ligases possess favorable characteristics for recruitment by PROTACs; both are expressed across myriad cell types facilitating multitissue protein targeting, there are numerous existing high-avidity molecular binders, and both can engage a large variety of target proteins.19 However, future PROTAC compounds will likely exploit the panoply of E3 ligases to optimize their target-specific functionality. Using E3 ligases expressed in a narrow range of cell subtypes can enhance targeting tissue specificity and minimize off-target toxicity.20 Additionally, E3 ligases present or overexpressed specifically in malignant cells represent an appealing strategy to target essential wild-type proteins within a highly selective cellular context.21,22 Identifying tissue- and cell-specific E3 ligases has been described as “one of the holy grails” of TPD research.23 New platforms capable of rapid systematic identification and screening of E3 ligase–binding ligands will be essential to realizing the full potential of PROTAC technology.24-27
Far from an inert bridging structure, the linker component of the PROTAC can significantly modulate the interactions between the target protein and E3 ligase, influencing the structure and activity of the ternary complex.28 The burgeoning field of “linkerology” demonstrates that rational approaches to linker design, and adjustment of linker properties such as length, flexibility, atomic composition, and incorporation of functionalized structural components can all modify PROTAC bioactivity characteristics including potency, isoform selectivity, ternary complex “co-operativity” (see discussion hereafter), cellular permeability, and PKs.18,29-33
Emergent technologies can aid identification of novel warheads for use in PROTACs and screen for superior cooperativity between the various PROTAC components. DNA-encoded library approaches are high-throughput screening methods using unique DNA-barcodes linked to warhead candidate molecules. After incubation, unbound ligands are removed, and DNA barcodes linked to remaining high-affinity molecules are amplified and sequenced. This technique can potentially evaluate millions of warhead molecules simultaneously and can be adapted to study linker variations or E3 ligases.34-38
Pharmacological considerations
PROTAC pharmacological properties differ significantly from conventional SMIs. Traditional small-molecule drugs are inhibitory, with efficacy derived from continuous target occupancy requiring continuous local presence at sufficient concentrations. Such “occupancy-driven” pharmacology necessitates stoichiometric dosing as the molecular drug–target interaction occurs at a 1:1 ratio. PROTACs are not bound by this constraint because they degrade rather than inhibit the target. Additionally, after target ubiquitination the PROTAC is released to engage additional target proteins, and PROTACs therefore exhibit “event-driven pharmacology,” achieving therapeutic efficacy even at substoichiometric doses.23
The occupancy-dependence of SMIs demands high-affinity target binding, and the target binding pocket must be active or allosteric, such that functional activity is suppressed upon binding. These requirements impose challenging limits on SMI design and restrict the range of target proteins to those with suitable binding domains. Additionally, these requirements are vulnerable to resistance pathways including target overexpression, acquired mutations modifying the binding site structure, or drug efflux. Overcoming these resistance mechanisms often requires dose escalation, risking untenable toxicity.39 PROTACs and other TPD modalities are not subject to the same constraints. Although target binding obviously remains essential, the binding site need not be active and high affinity binding is not requisite.40 In fact, the binding affinity of the warhead and that of the resultant ternary complex are not necessarily correlated.41-43 The concept relevant to this is “cooperativity,”44 whereby the binding or activity of 1 molecule is influenced by another, and alters the behavior of the entire system. In relation to PROTACs, cooperativity refers to protein-to-protein interactions that particularly affect the nature of the ternary complex. Positive cooperativity describes protein-to-protein interactions that promote ternary complex formation, whereas negative cooperativity denotes the converse. PROTAC designs that leverage positive cooperativity can enhance potency and improve target specificity, enabling differentiation between related proteins or even isoforms.18,45-48
A potential pitfall in PROTAC dosing known as the “hook effect” can lead to a counterintuitive reduction in PROTAC potency at higher doses. When excess PROTAC is present, both the target protein and E3 ligases can become saturated, and surplus PROTAC molecules form inactive bivalent complexes, reducing protein degradation efficacy.49 The hook effect appears to be an intrinsic property of all PROTACs, and although well documented in in vitro studies, there are less in vivo data to guide clinical dosing.50 The hook effect and other PROTAC properties demand an alternative approach to physicochemical assessments. Accordingly, new metrics have been developed, such as the concentration required to achieve 50% of maximal degradation (DC50) and maximal degradation (DCmax) among other descriptors of degradation kinetics, an approach distinct from other pharmacotherapeutics.51,52 Recent data suggest that positive cooperativity can overcome the hook effect by stabilizing the ternary complex.18,53 Such advancements highlight how precise molecule design can modulate and enhance future PROTACs but also reveal the need for further study to ensure optimized dosing in clinical practice. PROTAC delivery remains a major developmental challenge, limiting translational progress of promising preclinical candidates. Compared with conventional SMIs, PROTACs generally have a high molecular weight (700-110 Da), a greater number of hydrogen bond acceptors, highly polar surfaces, and a variety of other characteristics that contribute to PK uncertainty.54 Consequently, PROTACs are typically poorly water soluble, and predisposed to poor cellular penetration and risks of cellular efflux, pharmacokinetically operating beyond the Lipinski “rule of 5” that has traditionally described orally bioavailable compounds.55 Nonetheless, a myriad of PROTACs in clinical trial are indeed orally bioavailable, an important and appealing attribute that demonstrates the feasibility of this formulation. Efforts to interrogate and enhance PROTAC physicochemical properties to optimize their PKs have been described in detail by others.54,56
Pharmacodynamics (PDs) for PROTACs require special consideration and depend significantly on the target protein’s distribution, role in normal physiology, and half-life. As protein degraders, PROTACs may demonstrate a marked discordance between PKs and PDs, particularly when the target protein has a long half-life and/or a slow rate of synthesis, with which prolonged therapeutic effects can be sustained after PROTAC clearance. A potential limitation of PROTACs is off-target toxicity secondary to E3 ligase engagement with nontarget neosubstrates (target proteins that are not normally degraded). Nontarget neosubstrate degradation reduces therapeutic specificity and risks off-target toxicity. This phenomenon is routinely observed (particularly with cereblon based compounds) and is an important consideration in TPD drug development.57-59 Additionally, PROTACs could potentially perturb the ubiquitination and degradation of endogenous substrates of the E3 ligase they leverage. Complications related to this phenomenon would be expected to vary with the target protein, the target tissue E3 ligase expression profile, as well as their degradation capacity, and should be considered as a cause of potential off-target toxicity.
TPD therapeutic candidates require a specialized approach to target validation, molecular design, and PK/PD evaluation. Suitable methodologies are still being optimized but relevant metrics include in vitro target engagement assays, cellular engagement assays, ternary complex formation analysis, cooperativity assessment, and specific approaches to PK/PD studies.60,61
Clinical advantages for PROTACs
PROTACs offer several distinctive advantageous capabilities. The ability to degrade high value but conventionally undruggable proteins including those without enzymatic activity or an active binding pocket is a major advance that profoundly expands the druggable proteome. PROTAC-mediated degradation also facilitates circumvention of common mechanisms of SMI resistance associated with disease progression. Additionally, PROTACs may mitigate toxicity associated with previously identified effective but unsafe SMI drugs, allowing them to be repurposed as PROTAC warheads. It is useful to consider the following particular advantages in the context of current treatment challenges in hematological malignancies.
Drugging the undruggable targets
c-Myc
c-Myc is an archetypal high value cancer target that is currently undruggable with conventional SMIs. Myc family transcription factors (TFs) influence fundamental cellular processes affecting ∼10% to 15% of the genome.62,63 c-Myc plays a pivotal pathological role in many hematological malignancies, and aberrant c-Myc expression often correlates with aggressive tumor behavior and inferior treatment outcomes.64 Suppression of c-Myc results in potent tumor regression in numerous preclinical cancer models.65-67 Yet despite its undisputed therapeutic value, c-Myc targeting currently relies on indirect approaches.68 As a TF, the absence of an active enzymatic binding pocket makes functional inhibition of c-Myc prohibitively challenging; and localization within the nucleus shields c-Myc from antibody therapeutics. With no direct targeting agents, c-Myc is highly attractive for TPD, and although data remain preliminary, the c-Myc–targeting degraders GT19630 and GT19715 have shown high efficiency c-Myc degradation in in vitro models and potent inhibition of tumor growth in murine xenografts of acute myeloid leukemia (AML), including with TP53-mutated disease, and in T acute lymphoblastic leukemia (ALL) cells.69,70 Bromodomain and extraterminal domain proteins epigenetically regulate c-Myc transcription and have been exploited as targets to indirectly downregulate c-Myc in cancer.71 Several PROTACs targeting bromodomain and extraterminal domain family proteins have demonstrated promising preclinical activity in leukemia models (supplemental Table 1), consistently downregulating c-Myc and providing another exciting pathway to treating Myc-driven malignancies.72-77
STAT3
Signal transducer and activator of transcription 3 (STAT3) is a TF that, when constitutively activated, has an important role in many hematological malignancies.78,79 STAT3 has direct effects on cancer cellular proliferation, survival, and invasion, contributing to treatment resistance and weakening antitumor immunity.80,81 As with c-Myc, STAT3 is broadly considered a high-value therapeutic target but also lacks a targetable active binding site. STAT3’s activity was previously thought to be dependent upon dimerization, occurring through its Src homology 2 (SH2) domain.82 Given that direct targeting of STAT3 has proven insurmountable, novel therapies were designed to block the SH2 domain and thus inhibit dimerization.78 These agents were limited by a lack of selectivity, because the SH2 domain is structurally conserved among other STAT proteins, and the discovery that STAT3 monomers retain transcriptional activity such that preventing dimerization can only ever achieve partial efficacy.83,84 SD-36 is a STAT3 PROTAC structurally based on the STAT3 inhibitor SI-109 and achieves profound depletion of wild-type and mutant STAT3 in cell lines.85 SD-36 demonstrates 1000-fold greater potency than its parent SMI, showing exquisite selectivity with no substantive degradation of other STAT proteins. Complete tumor regression was achieved in mouse xenograft models of AML and anaplastic large cell lymphoma at well tolerated doses.86
KT-333 is another STAT3 degrader that retards cellular proliferation and induces apoptosis in multiple cancer cell lines. Selective STAT3 depletion was achieved over 9000 other proteins, and xenograft data demonstrated ∼90% elimination of STAT3 over 48 hours.87 KT-333 is currently being trialed in patients with advanced solid tumors and relapsed/refractory lymphoma (ClinicalTrials.gov identifier: NCT05225584). Preliminary safety and early best response data are presented in Table 1 and STAT3 levels were substantially reduced in circulating peripheral blood mononuclear cells, even at the lowest dose levels.94
Current clinical trials of PROTAC therapeutics in hematological malignancies
| Trial/clinical trial number . | PROTAC . | Target protein . | E3 ligase . | Disease context . | Clinical trial phase . | Patients, n . | Cohort description . | Additional cohort data . | Safety data . | Efficacy data . | References . |
|---|---|---|---|---|---|---|---|---|---|---|---|
| NX-2127-001 NCT04830137 | NX-2127 | BTK/Ikaros/Aiolos | Cereblon | B-cell malignancies | 1 | Total: 54 33 CLL/SLL DLBCL, 6 MCL, 7 MZL, 3 WM, 3 FL, 2 | Median age: 72.5 y F:M, 31.5%:68.5% Median prior therapies: 4 CLL cohort: Median age: 74 y F:M, 33.3%:66.7% Median prior therapies: 5 NHL cohort: Median age: 70 y F:M, 28.6%:71.4% Median prior therapies: 4 | CLL cohort Previous BTKi: 100% Previous pirtobrutinib: 27.3% Previous BTKi & BCL2i: 78.8% BCL2 mutation: 12.1% BTK mutation: 36.4% | DLTs: 2 AE Gr ≥3: Neutropenia: 38.3% Hypertension: 14.9% Anemia: 12.8% Thrombocytopenia: 6.4% COVID-19: 6.4% Atrial fibrillation: 6.4% Pneumonia: 6.4% Leucocytosis: 6.4% Sepsis: 4.3% Increased lymphocyte count: 4.3% Pruritis: 2.1% Dyspnea: 2.1% | CLL evaluable cohort (n = 27) ORR: 40.7% PR: 40.7% SD: 44.4% PD: 14.8% NHL evaluable cohort (n = 17): CR: 11.8% PR: 11.8% SD: 29.4% PD: 47.1% | 88 |
| NX-5948-301 NCT05131022 | NX-5948 | BTK | Cereblon | B-cell malignancies | 1 | Total 26: 7 CLL 7 DLBCL 5 MCL 3 MZL 1 FL 1 PCNSL 2 WM | Median age: 65 y F:M, 28.6%:71.4% Median prior therapies: 4 | CLL cohort: Median age: 64 y F:M 28.6%:71.4% Median prior therapies: 3 TP53 aberrancy: 33.3% BTK mutation (T474): 16.7% PLCG1/2 mutation: 33.3% BCL2 mutation: 33.3% NHL/WM cohort: Median age: 63 y F:M 31.6%:68.4% Median prior therapies: 5 TP53 aberrancy: 20% | No DLTs recorded No treatment-related AE led to discontinuation AE Gr >3: Neutropenia: 19.2% Thrombocytopenia: 7.7% COVID-19: 7.7% Pneumonia: 7.7% Anemia: 3.8% | CLL evaluable cohort (n = 7) 3 PR (42.8%) 2 SD (28.5%) 2 PD (28.5%) NHL evaluable cohort: (n = 19) 2 PMR 1 PR 2 SD 13 PD | 89 89 |
| BGB-16673-101 NCT05006716 NCT05294731 | BGB-16673 | BTK | Undisclosed | B-cell malignancies | 1 | Total: 50 24 CLL/SLL 7 MCL 3 MZL 6 WM 2 DLBCL 6 FL 2 RT | Median age: 70.5 y F:M, 34%:66% Median prior therapies: 4 | CLL cohort unmutated IGHV: 84.2%, TP53 aberrancy: 45.8% Complex CG: 40% Whole cohort: Previous BTKi: 80% Previous BCL2i: 56% BTK mutation present: 29.2% PLCG2 mutation present: 8.3% BCL2 mutation present: 44.4% | No DLTs recorded Any AE Gr ≥3: 38% Treatment-related AE Gr ≥3: 22% Deaths: 4% AE Gr ≥3: Contusion: 30% Diarrhea: 24% Fatigue: 20% Increased amylase: 16% Neutropenia: 16% Increased lipase: 14% Pyrexia: 14% Cough: 12% Headache: 10% Pneumonia: 6% COVID-19 pneumonia: 4% AEs of interest Any bleeding: 42% Any infection: 50% Atrial fibrillation: 0% | Total evaluable cohort (n = 28): ORR: 54% CR: 4% PR/PR-L: 50% MR: 4% SD: 18% PD: 18% Ceased: 7% CLL evaluable cohort: (n = 10) PR/PR-L: 70% SD: 20% Ceased: 10% MCL/MZL/WM evaluable cohort (n = 16) CR: 6.3% PR: 43.8% MR: 6.3% SD: 18.8% PD 18.8% Ceased: 6.3% DLBCL/RT (n = 2) PD: 100% | 90 |
| NCT04861779 | HSK29116 | BTK | Undisclosed | B-cell malignancies | 1 | NA | 91 | ||||
| NCT05780034 | AC0676 | BTK | Undisclosed | B-cell malignancies | 1 | NA | 92 | ||||
| NCT05753501 | ABBV-101 | BTK | Undisclosed | B-cell malignancies | 1 | NA | 93 | ||||
| KT333-TL-101 NCT05225584 | KT-333 | STAT3 | Undisclosed | NHL, CTCL, LGL-L, PTCL | 1 | Total: 29 1 B-cell NHL 1 HL 5 CTCL 1 PTCL 2 LGL-L 19 solid tumor | Median age 65 F:M, 27.6%:72.4% ECOG range, 0-1 ≥3 prior therapies: 86.2% | DLTs: 2 (stomatitis, arthralgia) No treatment-related serious AEs recorded Any AE Gr ≥3: 10% Arthralgia: 3.4% Stomatitis: 3.4% Decreased weight: 3.4% Most common AEs were grade 1/2 constipation, fatigue, nausea, and anemia | Best reported responses HL:PR CTCL cohort, n = 5: PR: 2 SD: 1 Solid tumor cohort SD: 4 | 94 | |
| NCT05233033 | KT-413 | IRAK4 | Cereblon | DLBCL (MYD88 mutated) | 1 | NA | 95 | ||||
| NCT04886622 | DT2216 | BCLXL | VHL | Various malignancies | 1 | NA | 96 | ||||
| NCT05487170 | RNK05047 | BRD4 | HSP90 chaperone complex | DLBCL | 1/2 | NA | 97 |
| Trial/clinical trial number . | PROTAC . | Target protein . | E3 ligase . | Disease context . | Clinical trial phase . | Patients, n . | Cohort description . | Additional cohort data . | Safety data . | Efficacy data . | References . |
|---|---|---|---|---|---|---|---|---|---|---|---|
| NX-2127-001 NCT04830137 | NX-2127 | BTK/Ikaros/Aiolos | Cereblon | B-cell malignancies | 1 | Total: 54 33 CLL/SLL DLBCL, 6 MCL, 7 MZL, 3 WM, 3 FL, 2 | Median age: 72.5 y F:M, 31.5%:68.5% Median prior therapies: 4 CLL cohort: Median age: 74 y F:M, 33.3%:66.7% Median prior therapies: 5 NHL cohort: Median age: 70 y F:M, 28.6%:71.4% Median prior therapies: 4 | CLL cohort Previous BTKi: 100% Previous pirtobrutinib: 27.3% Previous BTKi & BCL2i: 78.8% BCL2 mutation: 12.1% BTK mutation: 36.4% | DLTs: 2 AE Gr ≥3: Neutropenia: 38.3% Hypertension: 14.9% Anemia: 12.8% Thrombocytopenia: 6.4% COVID-19: 6.4% Atrial fibrillation: 6.4% Pneumonia: 6.4% Leucocytosis: 6.4% Sepsis: 4.3% Increased lymphocyte count: 4.3% Pruritis: 2.1% Dyspnea: 2.1% | CLL evaluable cohort (n = 27) ORR: 40.7% PR: 40.7% SD: 44.4% PD: 14.8% NHL evaluable cohort (n = 17): CR: 11.8% PR: 11.8% SD: 29.4% PD: 47.1% | 88 |
| NX-5948-301 NCT05131022 | NX-5948 | BTK | Cereblon | B-cell malignancies | 1 | Total 26: 7 CLL 7 DLBCL 5 MCL 3 MZL 1 FL 1 PCNSL 2 WM | Median age: 65 y F:M, 28.6%:71.4% Median prior therapies: 4 | CLL cohort: Median age: 64 y F:M 28.6%:71.4% Median prior therapies: 3 TP53 aberrancy: 33.3% BTK mutation (T474): 16.7% PLCG1/2 mutation: 33.3% BCL2 mutation: 33.3% NHL/WM cohort: Median age: 63 y F:M 31.6%:68.4% Median prior therapies: 5 TP53 aberrancy: 20% | No DLTs recorded No treatment-related AE led to discontinuation AE Gr >3: Neutropenia: 19.2% Thrombocytopenia: 7.7% COVID-19: 7.7% Pneumonia: 7.7% Anemia: 3.8% | CLL evaluable cohort (n = 7) 3 PR (42.8%) 2 SD (28.5%) 2 PD (28.5%) NHL evaluable cohort: (n = 19) 2 PMR 1 PR 2 SD 13 PD | 89 89 |
| BGB-16673-101 NCT05006716 NCT05294731 | BGB-16673 | BTK | Undisclosed | B-cell malignancies | 1 | Total: 50 24 CLL/SLL 7 MCL 3 MZL 6 WM 2 DLBCL 6 FL 2 RT | Median age: 70.5 y F:M, 34%:66% Median prior therapies: 4 | CLL cohort unmutated IGHV: 84.2%, TP53 aberrancy: 45.8% Complex CG: 40% Whole cohort: Previous BTKi: 80% Previous BCL2i: 56% BTK mutation present: 29.2% PLCG2 mutation present: 8.3% BCL2 mutation present: 44.4% | No DLTs recorded Any AE Gr ≥3: 38% Treatment-related AE Gr ≥3: 22% Deaths: 4% AE Gr ≥3: Contusion: 30% Diarrhea: 24% Fatigue: 20% Increased amylase: 16% Neutropenia: 16% Increased lipase: 14% Pyrexia: 14% Cough: 12% Headache: 10% Pneumonia: 6% COVID-19 pneumonia: 4% AEs of interest Any bleeding: 42% Any infection: 50% Atrial fibrillation: 0% | Total evaluable cohort (n = 28): ORR: 54% CR: 4% PR/PR-L: 50% MR: 4% SD: 18% PD: 18% Ceased: 7% CLL evaluable cohort: (n = 10) PR/PR-L: 70% SD: 20% Ceased: 10% MCL/MZL/WM evaluable cohort (n = 16) CR: 6.3% PR: 43.8% MR: 6.3% SD: 18.8% PD 18.8% Ceased: 6.3% DLBCL/RT (n = 2) PD: 100% | 90 |
| NCT04861779 | HSK29116 | BTK | Undisclosed | B-cell malignancies | 1 | NA | 91 | ||||
| NCT05780034 | AC0676 | BTK | Undisclosed | B-cell malignancies | 1 | NA | 92 | ||||
| NCT05753501 | ABBV-101 | BTK | Undisclosed | B-cell malignancies | 1 | NA | 93 | ||||
| KT333-TL-101 NCT05225584 | KT-333 | STAT3 | Undisclosed | NHL, CTCL, LGL-L, PTCL | 1 | Total: 29 1 B-cell NHL 1 HL 5 CTCL 1 PTCL 2 LGL-L 19 solid tumor | Median age 65 F:M, 27.6%:72.4% ECOG range, 0-1 ≥3 prior therapies: 86.2% | DLTs: 2 (stomatitis, arthralgia) No treatment-related serious AEs recorded Any AE Gr ≥3: 10% Arthralgia: 3.4% Stomatitis: 3.4% Decreased weight: 3.4% Most common AEs were grade 1/2 constipation, fatigue, nausea, and anemia | Best reported responses HL:PR CTCL cohort, n = 5: PR: 2 SD: 1 Solid tumor cohort SD: 4 | 94 | |
| NCT05233033 | KT-413 | IRAK4 | Cereblon | DLBCL (MYD88 mutated) | 1 | NA | 95 | ||||
| NCT04886622 | DT2216 | BCLXL | VHL | Various malignancies | 1 | NA | 96 | ||||
| NCT05487170 | RNK05047 | BRD4 | HSP90 chaperone complex | DLBCL | 1/2 | NA | 97 |
AE, adverse event; BCLxL, B-cell lymphoma extralarge; BCL2, B-cell lymphoma protein 2; BRD, bromodomain; CG, cytogenetics; CR, complete response; CTCL, cutaneous T-cell lymphoma; DLBCL, diffuse large B-cell lymphoma; DLT, dose-limiting toxicity; ECOG, Eastern Cooperative Oncology Group; F, female; FL, follicular lymphoma; Gr, grade; HL, Hodgkin lymphoma; IGHV, immunoglobulin heavy variable; IRAK, interleukin 1 receptor associated kinase; LGL-L, large granular lymphocytic leukemia; M, male; MCL, mantle cell lymphoma; MZL, marginal zone lymphoma; NHL, non-Hodgkin lymphoma; PR, partial response; PR-L, partial response with lymphocytosis; PD, progressive disease; PLCG1/2, phospholipase C gamma-1/2; PTCL, peripheral T-cell lymphoma; RT, Richter transformation; SD, stable disease; SLL, small lymphocytic lymphoma; ST, solid tumor; WM, Waldenstrom macroglobulinemia.
Overcoming resistance, BTK and BCR:ABL
BTK
Bruton tyrosine kinase (BTK) is a cytoplasmic tyrosine kinase, critical to B-cell receptor (BCR) signaling, and important in the pathogenesis of many B-cell malignancies.98-100 SMIs such as ibrutinib, acalabrutinib, zanubrutinib, and pirtobrutinib that bind to the BTK adenosine triphosphate–binding pocket have redefined the standard of care in numerous B-cell cancers, but resistance mutations contributing to disease progression emerge frequently, often involving the BTK inhibitor (BTKi) binding site.101,102 In chronic lymphocytic leukemia (CLL), mutations affecting the cysteine 481 (C481) residue of the adenosine triphosphate–binding pocket are the most common mechanism of ibrutinib resistance.103 The conformational change impairs binding of ibrutinib and other covalent BTKis, restoring the catalytic function of BTK to drive disease progression. Noncovalent BTKis, such as pirtobrutinib, can overcome the C481 mutations but are also susceptible to binding pocket aberrancy.104,105 Numerous additional BTK mutations that confer resistance to both covalent and noncovalent inhibitors have now been documented such as the L528W, T474I, M437R, and V416L mutants. These include “kinase dead” mutants that lack the enzymatic activity of BTK but retain a scaffolding function that maintains aberrant BTK signaling and are unaffected by functional inhibition.102,106,107
For patients with CLL harboring these mutations, treatment options are limited and outcomes poor. The advantages of PROTAC BTK degradation becomes apparent in this context because unlike conventional BTKis, PROTAC activity is not reliant on active site binding or sustained occupancy. PROTACs may therefore provide an alternative for CLL that has become resistant to even the most potent BTK SMIs. Numerous BTK-targeting PROTACs are under active investigation.
The first-in-class BTK PROTAC is NX-2127, a cereblon-recruiting molecule that targets BTK but also degrades the TFs IKZF3 (Aiolos) and IKZF1 (Ikaros).108 Preclinical studies demonstrated marked degradation of wild-type BTK; but NX-2127 also degrades the clinically relevant mutant BTKs C481S, L528W, T474I, M437R, and V416L.109 PROTAC activity against kinase-dead mutants illustrates an important advantage of protein degraders over SMIs; target proteins that exert oncogenic activity through nonenzymatic roles such as kinome scaffolding remain susceptible PROTACs but are an important emerging cause of resistance to SMIs. These data suggest that NX-2127 could overcome mutations that neutralize the most advanced conventional BTKis. A first-in-human phase 1 trial of orally administered NX-2127 has released preliminary results, summarized in Table 1.88 Although the patient cohort was heavily pretreated, the best overall response rate (ORR) in the evaluable CLL cohort was 40.7%, with responses documented in patients that were double refractory to BTKis and B-cell lymphoma 2 (BCL2) inhibitors as well as in patients with noncovalent BTKi-resistant disease, for whom outcomes are historically abysmal.88,110,111 In patients with non-Hodgkin lymphoma/Waldenstrom macroglobulinemia the ORR was 23.6%.
A sister compound NX-5948 possesses greater specificity for BTK, with no anti-Ikaros/Aiolos activity. As with NX-2127, NX-5948 inhibits tumor growth in ibrutinib-resistant xenograft lymphoma mouse models and, importantly, has also been shown to penetrate the blood-brain barrier and improve survival in mouse models of central nervous system diffuse large B-cell lymphoma.112 A clinical trial with NX-5948 is underway (ClinicalTrials.gov identifier: NCT05131022), with reassuring safety data and encouraging early efficacy described in Table 1.89
BGB-16673 is another BTK-targeting PROTAC with preclinical evidence of potent activity against wild-type and drug-resistant mutant BTK.113 Preliminary data from the phase 1 dose escalation study are described in Table 1. In total, 50 patients with a range of B-cell malignancies received BGB-16673 and no patient discontinued therapy because of adverse effects. Substantial reductions in BTK protein levels in the peripheral blood and tumor tissue occurred at even the lowest dose, and the whole cohort ORR was 57%.90 Although these preliminary data represent an early study time point, the response rate in this relapsed/refractory cohort is impressive and an important early milestone for BTK PROTACs.
Numerous additional BTK PROTACs are under clinical investigation but yet to be reported (Table 1).
BCR/ABL
The management of chronic myeloid leukemia (CML) was revolutionized by tyrosine kinase inhibitors (TKIs), which inhibit the CML driver breakpoint cluster region-Abelson murine leukemia viral oncogene homolog 1 (BCR-ABL) fusion protein. Although TKIs confer remarkable benefit for most patients with CML, some develop resistant disease, primarily resulting from BCR-ABL kinase domain (KD) mutations, such as T315i, which abrogate TKI binding.114 Such mutations present a challenging treatment scenario associated with unfavorable patient outcomes and few therapeutic options.115 PROTACs may prove an alternative for patients with KD and other BCR-ABL mutations.88,109,110 BCR-ABL PROTACs have demonstrated activity against T315i-mutated CML, and the V468F mutation that is resistant to the new-generation TKI asciminib.116 In 2022, Ma et al reported a PROTAC termed PMIBcr/Abl-R6, which degrades BCR-ABL irrespective of mutation status.117PMIBcr/Abl-R6 binds to the BCR-ABL oligomerization domain rather than the KD and is therefore unaffected by KD mutations. Interestingly, PMIBcr/Abl-R6 also binds to the E3 ligase MDM2, which ordinarily degrades p53, providing dual functionality; targeting BCR-ABL and simultaneously activating p53 to promote apoptosis. PMIBcr/Abl-R6 demonstrates potent activity in vitro, including against patient-derived CML and ALL cell lines. Activity was retained despite BCR-ABL breakpoint variability and KD mutations including T315i. In an imatinib-resistant, T315i-mutant xenograft model, PMIBcr/Abl-R6 suppressed tumor growth without apparent toxicity in the mice.117 No BCR-ABL PROTAC has yet entered clinical trial, but there is clear therapeutic potential for these agents in TKI-resistant CML and ALL, with numerous PROTAC candidates under development.118
Mitigating on-target toxicity
BCLxL/BCL2
Antiapoptotic proteins
Perturbed regulation of apoptosis is a hallmark of cancer and common driver of oncogenesis in hematological malignancies.119 Antiapoptotic proteins such as B-cell lymphoma 2 (BCL-2) and B-cell lymphoma extra large (BCL-xL) are overexpressed in numerous blood cancers, enhancing cancer cell survival and blunting treatment sensitivity.120,121 Therapeutic modulation of apoptotic balance has proven highly successful in several disease contexts. Navitoclax was an early BH3-mimetic drug that binds antiapoptotic proteins BCL-XL, BCL-2, and BCL-W, and is therapeutically active in CLL, non-Hodgkin lymphoma, and AML.122-124 However, because platelet survival depends on BCL-XL, navitoclax caused challenging on-target thrombocytopenia that significantly constrained its clinical utility.125 PROTAC design can potentially circumvent these limitations as evidenced by DT2216, a VHL-recruiting PROTAC that targets BCL-XL with a navitoclax-based warhead. An exemplar of leveraging differential E3-ligase expression profiles, DT2216 causes only minor BCL-XL degradation in platelets, because VHL is minimally expressed in platelets. In vitro, DT2216 has less impact on platelet viability than navitoclax, and mouse model data showed modest and tolerable thrombocytopenia contrasting with marked thrombocytopenia after navitoclax exposure. Despite lessened toxicity, the antitumor efficacy of DT2216 exceeded that of navitoclax in both cell-based and animal experiments, and this candidate has progressed to investigation in a phase 1 clinical trial (Table 1).20 Furthermore, the other dominant PROTAC E3 ligase, cereblon, is also minimally expressed in platelets, and PZ15227, a BCL-XL targeting PROTAC that engages cereblon, also possesses potent efficacy without inducing severe thrombocytopenia.126 On-target toxicity remains a challenging limitation of SMIs, which cannot be delivered to target cells with precise selectivity. By utilizing differences in E3 ligase expression, as evidenced by DT2216 and PZ15227, PROTACs can achieve greater tissue specificity and thereby mitigate toxicity. As more exploitable E3 ligases are identified and their tissue expression profiles defined, PROTAC design flexibility and disease adaptation will also improve, providing greater safety augmentation.
Other targets
The discussed agents are certainly not an exhaustive catalog of extant PROTACs. Further PROTACs relevant to hematological malignancies are under development and summarized in supplemental Table 1, available on the Blood website.
Mechanisms of resistance to PROTAC agents
Given the limited clinical deployment of PROTACs, patterns of emergent resistance remain largely unknown. However, some data are available and potential resistance mechanisms can be inferred from preclinical studies.
Consistent with known mechanisms of resistance in immunomodulators, most PROTAC resistance involves acquired aberrations in the UPS. Cell lines continually exposed to PROTAC compounds generate resistant clones after alteration or even deletion of key UPS components such as E2 enzymes, E3 ligases, or other regulatory components like the COP9 signalosome or CUL2.127,128 A CRISPR screening approach designed to identify mediators of PROTAC resistance demonstrated concordant results.129 Mutations altering the target protein have been documented to confer resistance (as seen in most conventional SMIs),130 but PROTAC resistance may more commonly arise from mutations affecting UPS components. To counteract this resistance pattern, PROTACs could be deployed sequentially, targeting the same protein but using different E3 ligases, underscoring the critical importance of expanding the range of usable E3 ligases.128 Upregulation of the drug efflux pump, multidrug resistance protein 1, can also confer PROTAC resistance, although this may be overcome by concurrently administering inhibitors of multidrug resistance protein 1.131 As the clinical study of PROTACs progresses, further resistance mechanisms will be revealed, but the distinct nature of PROTACs may allow innovative means of combating resistance that could also synergize with existing treatments.
Emerging TPD technologies
Though clinical PROTAC technology represents the cutting edge of targeted therapeutics, there are already numerous modified PROTAC and alternative TPD designs that seek to improve the range of targetable proteins, delivery formats, safety and specificity of this approach. These technologies are briefly described and summarized in Table 2.
Emerging TPD technologies
| TPD Technology . | Description . | Utility . | Considerations . | References . |
|---|---|---|---|---|
| AbTACs | Use bispecifc antibodies to engage target membrane proteins and trigger internalization and degradation | Enable targeting of membrane proteins (eg, PD1; beyond capability of orthodox PROTACs) | Require cell surface E3 ligases such as ZNRF3 and RNF43 | 132 |
| Oligonucleotide-based PROTACs | Use oligonucleotide motifs as PROTAC warhead Various oligonucleotide formats: single- or double-stranded DNA, RNA, or G-quadruplexes | Enable targeting of TFs and RNA-binding proteins via their nucleic acid–binding domains | Compared to orthodox PROTACs, greater challenges with cell permeability and target cell access | 133,134 |
| Multitargeted PROTACs | Target multiple proteins simultaneously via 1 molecular construct | Tractable multiplexed targeting | Increases complexity of E3 ligase selection Likely increased risk of nontarget neosubtrate degradation, risk of off-target toxicity | 135 |
| BioPROTACs | Engineered fusion proteins, E3 ligase substrate recognition domain modified to express target-specific peptide or protein | Dispenses with warhead identification and ternary complex optimization; can be delivered as mRNA | Unable to be administered orally; delivery is challenging | 136-140 |
| Pre-PROTACs | PROTACs that are only activated by specific stimuli or contexts. Examples: light activated, radiograph activated, and hypoxia or ROS triggers | Improve spatiotemporal and cellular contextual control of PROTAC activity; thus, reduce dose requirements, reduced toxicity | Alternative physical, chemical, or electromagnetic stimuli could be used if suitable sensor moieties are identified | 141-147 |
| MGs | Induce colocalization of target protein and E3 ligase. Example: lenalidomide and pomalidomide. | Smaller molecular size than conventional PROTACs but superior PK | Limited design flexibility | 148-150 |
| Lysosome-mediated TPD LYTACs AUTACs ATTECs AUTOTACs MoDE-As | Leverage lysosomal non-UPS protein cycling cellular processes: (1) autophagy-lysosomal system degrades cytoplasmic proteins, protein aggregates, and organelles; (2) endosome–lysosomal pathway that degrades extracellular proteins | Expands the range of protein, protein structures, and other biomolecules targetable with TPD therapeutics beyond those subject to the UPS | Early in development | 151-153 |
| TPD Technology . | Description . | Utility . | Considerations . | References . |
|---|---|---|---|---|
| AbTACs | Use bispecifc antibodies to engage target membrane proteins and trigger internalization and degradation | Enable targeting of membrane proteins (eg, PD1; beyond capability of orthodox PROTACs) | Require cell surface E3 ligases such as ZNRF3 and RNF43 | 132 |
| Oligonucleotide-based PROTACs | Use oligonucleotide motifs as PROTAC warhead Various oligonucleotide formats: single- or double-stranded DNA, RNA, or G-quadruplexes | Enable targeting of TFs and RNA-binding proteins via their nucleic acid–binding domains | Compared to orthodox PROTACs, greater challenges with cell permeability and target cell access | 133,134 |
| Multitargeted PROTACs | Target multiple proteins simultaneously via 1 molecular construct | Tractable multiplexed targeting | Increases complexity of E3 ligase selection Likely increased risk of nontarget neosubtrate degradation, risk of off-target toxicity | 135 |
| BioPROTACs | Engineered fusion proteins, E3 ligase substrate recognition domain modified to express target-specific peptide or protein | Dispenses with warhead identification and ternary complex optimization; can be delivered as mRNA | Unable to be administered orally; delivery is challenging | 136-140 |
| Pre-PROTACs | PROTACs that are only activated by specific stimuli or contexts. Examples: light activated, radiograph activated, and hypoxia or ROS triggers | Improve spatiotemporal and cellular contextual control of PROTAC activity; thus, reduce dose requirements, reduced toxicity | Alternative physical, chemical, or electromagnetic stimuli could be used if suitable sensor moieties are identified | 141-147 |
| MGs | Induce colocalization of target protein and E3 ligase. Example: lenalidomide and pomalidomide. | Smaller molecular size than conventional PROTACs but superior PK | Limited design flexibility | 148-150 |
| Lysosome-mediated TPD LYTACs AUTACs ATTECs AUTOTACs MoDE-As | Leverage lysosomal non-UPS protein cycling cellular processes: (1) autophagy-lysosomal system degrades cytoplasmic proteins, protein aggregates, and organelles; (2) endosome–lysosomal pathway that degrades extracellular proteins | Expands the range of protein, protein structures, and other biomolecules targetable with TPD therapeutics beyond those subject to the UPS | Early in development | 151-153 |
ATTECs, autophagosome tethering compounds; AUTACs, autophagy-targeting chimeras; AUTOTACs, autophagy-targeting chimeras; LYTACs, lysosome-targeting chimeras; MoDE-As, molecular degraders of extracellular proteins through the asialoglycoprotein receptor; mRNA, messenger RNA; PD1, programmed cell death protein 1; ROS, reactive oxygen species; TPD, targeted protein degradation.
Antibody-based PROTACs (AbTACs)
Conventional PROTACs primarily target intracellular proteins because of the cytosolic localization of the E3 ligases thus far used. AbTACs use recombinant bispecific antibodies and expand the range of degradable targetable proteins, as demonstrated by Cotton et al, who developed an AbTAC targeting the cell-surface immune checkpoint programmed death ligand 1.132
Oligonucleotide-based PROTACs
Multitargeted PROTACs
Recently, groups have developed PROTACs capable of degrading several targets simultaneously, demonstrating favorable potency and PKs when compared with single-targeting PROTACs.135
BioPROTACs
BioPROTACs are engineered E3 ligases whereby the substrate-recognition domain is modified to express a target-binding peptide or protein structure.136 Prototype bioPROTACs targeting c-Myc and KRAS have already been described.137,138 As modified proteins, a bioPROTAC could be delivered as a messenger RNA payload within a nanocarrier, but these technologies remain inchoate.139,140
Pre-PROTACs
Pre-PROTACs are akin to prodrugs, and only activated by specific stimuli or contexts. Numerous pre-PROTACs responding to different triggers have been developed, including light- and radiation–triggered designs.141-145 PROTACs can also be designed to discern tumor-selective chemical stimuli such as hypoxia or elevated levels of reactive oxygen species.146,147
Molecular glues (MGs)
As previously mentioned, MG compounds were in fact the vanguard class of TPD therapeutics; with immunomodulatory drugs (thalidomide, lenalidomide, and pomalidomide) proving transformational in multiple myeloma particularly.148 Structurally distinct from PROTACs, MGs are monovalent molecules that also induce UPS-dependent protein degradation through colocalization of the target protein and E3 ligase. Previously, MGs were discovered serendipitously, but new screening techniques are emerging and their favorable pharmacological properties have prompted substantial investment in the development of manifold new MGs targeting a multitude of proteins.149
Lysosome-mediated TPD
Lysosome-mediated protein degradation is an alternative homeostatic protein cycling system that regulates biomolecules not subject to the UPS.151 Several lysosomal mediated TPD technologies have arisen including lysosome-targeting chimeras, autophagy-targeting chimeras, autophagosome tethering compounds, and molecular degraders of extracellular proteins through the asialoglycoprotein receptor. These platforms remain in early phases of development but will further expand the druggable proteome and complement UPS-leveraging PROTACs. Lysosomal TPD is beyond the scope of this article but has been extensively reviewed recently.152,153
Conclusion and future directions
The epoch of PROTAC therapeutics is only just beginning, but there are cogent reasons to be optimistic about their translational potential. Protein degraders can target a more expansive segment of the proteome, placing known high value but conventionally undruggable targets within reach. PROTAC activity is likely unperturbed by many resistance mutations that abrogate conventional SMI efficacy, and PROTACs offer rational pathways to mitigating off-target and on-target toxicity. Future nuanced design of PROTAC components will further enhance potency and target specificity and we have discussed just some of the emerging technologies that could broaden the opportunities of TPD. However, clinical data are limited, and a rigorous understanding of long-term efficacy, safety, potential off-target effects, and resistance mechanisms is still required for effective translation. Nonetheless, PROTACs may come to define a new and welcome paradigm for targeted therapy in hematological malignancies and beyond.
Acknowledgments
The authors acknowledge that the term “PROTAC” is a registered trademark of Arvinas Operations, Inc (registration number, 5351627). As in the wider literature, the authors have used the term in reference to comparable agents with similar structures and functions. Both the figure and the visual abstract were created with the BioRender platform.
Authorship
Contribution: J.M.L.C. performed the literature review and wrote the manuscript; and J.F.S. conceptualized the article, reviewed, and edited the manuscript.
Conflict-of-interest disclosure: J.F.S. serves on advisory boards, and has received research funding from, AbbVie, Bristol Myers Squibb, Janssen, Roche, and Celgene; has received honoraria from AbbVie, Bristol Myers Squibb, and Roche; and has served on advisory boards for BeiGene, Genor Bio, and Gilead. J.M.L.C. declares no competing financial interests.
Correspondence: John F. Seymour, Division of Haematology/Medical Oncology, Peter MacCallum Cancer Centre and The Royal Melbourne Hospital, 305 Grattan St, Melbourne, VIC 3000, Australia; email: john.seymour@petermac.org.
References
Author notes
The online version of this article contains a data supplement.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal