In this issue of Blood, Boudia and colleagues1 introduce a human induced pluripotent stem cell (iPSC) model of ETO2::GLIS2-driven acute megakaryoblastic leukemia (AMKL). The model recapitulates the human disease, enabling a comprehensive molecular investigation that identifies osteogenic homeobox factor DLX3 as a critical player in the onset of leukemia.
Treatment outcomes of pediatric leukemia have drastically improved in recent decades, with survival rates of childhood acute myeloid leukemia (AML) now reaching 75%.2 However, some subtypes still suffer from a dismal prognosis. One of these subgroups is non–Down syndrome AMKL, an aggressive and rare form of AML most frequently observed in young children and rarely seen in adults. In 2012, a cryptic inversion of chromosome 16 was described.3,4 The inversion leads to expression of the ETO2::GLIS2 (also known as CBFA2T3::GLIS2) oncofusion protein, which was shown to induce self-renewal capacity and differentiation toward the megakaryocyte lineage.5 It was found in almost 20% of the pediatric non–Down syndrome AMKL cases and is associated with a particularly poor outcome, with an overall survival of only 15% to 30%.5 Onset of this condition, often in very young children, implies that the fusion may occur during fetal life, and the developmental susceptibility of the oncofusion has been confirmed in an animal model of the disease.6
The biological factors that trigger onset of the disease are poorly understood, and fetal characteristics are difficult to capture experimentally. Infantile AMKL is a prime example, as it is associated with development. Thus, a modeling strategy should faithfully recapitulate the developmental context and express the oncogene at physiological levels in relevant cell types. Boudia et al used a CRISPR/Cas9 approach to introduce the clinically relevant genetic aberration in human iPSCs. iPSCs are generated by reprogramming somatic cells to a pluripotent state. When these cells differentiate toward the hematopoietic lineage, they recapitulate features of embryonic hematopoiesis. Thus, they provide a human source of cells suitable for gene editing. The genetic complexity and clonal composition of the disease can be captured by generating iPS cell lines from samples from leukemic patients.7 Boudia and colleagues have instead introduced the oncofusion into otherwise healthy iPSCs, allowing them to study the effect of a single driver in a controlled fashion.
Bertuccio et al have previously presented a model of the ETO2::GLIS2-fusion based on iPSCs.8 In that model, the oncofusion was placed under the control of the pan-hematopoietic CD43 promoter in otherwise healthy iPSCs. Although the model replicated some characteristics seen in patients with AMKL, no disease development occurred upon transplantation to mice. Here Boudia and colleagues took on a different approach, using CRISPR/Cas9 technology to introduce the inversion into healthy iPSCs (see figure). By adding guide RNAs targeting the breakpoint regions, precise engineering of the fusion could be achieved. The fusion was thereby placed under the control of the endogenous ETO2 promoter, replicating what is seen in patients. The ETO2::GLIS2 iPS-derived hematopoietic cells could give rise to AML in mice, and phenocopied cellular and molecular features observed in pediatric patients with AMKL. One notable finding was the high expression of the cell surface protein CD56 seen in the model. High expression of CD56, in combination with low or negative expression of CD38, CD45, and HLA-DR, defines a distinct aberrant immunophenotype known as RAM. The RAM phenotype has a dismal prognosis9 and is often specifically associated with ETO2::GLIS2 AMKL. Identification of a RAM phenotype in a diagnostic pediatric AML sample often prompts targeted analysis of the ETO2::GLIS2 translocation when genome-wide testing is not available. Thus, a RAM phenotype has significant clinical relevance, and the fact that the model developed by Boudia et al expresses high levels of CD56 further enhances its clinical value.
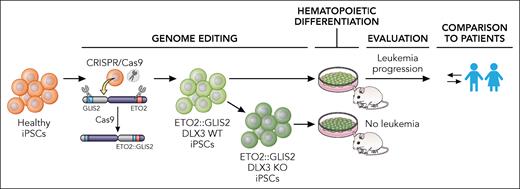
An iPSC model of ETO2::GLIS2-driven acute megakaryoblastic leukemia. Boudia et al genome edited healthy iPSCs to express the ETO2::GLIS2 oncofusion. After differentiation to blood, cells were transplanted to mice and leukemia initiation and progression was evaluated. The features observed were compared with patient samples. The authors also generated ETO2::GLIS2 iPSCs deficient in DLX3 (KO), which resulted in loss of the leukemia-initiating ability in mice. iPSC, induced pluripotent stem cell; KO, knockout; WT, wild-type. Professional illustration by Patrick Lane, ScEYEnce Studios.
An iPSC model of ETO2::GLIS2-driven acute megakaryoblastic leukemia. Boudia et al genome edited healthy iPSCs to express the ETO2::GLIS2 oncofusion. After differentiation to blood, cells were transplanted to mice and leukemia initiation and progression was evaluated. The features observed were compared with patient samples. The authors also generated ETO2::GLIS2 iPSCs deficient in DLX3 (KO), which resulted in loss of the leukemia-initiating ability in mice. iPSC, induced pluripotent stem cell; KO, knockout; WT, wild-type. Professional illustration by Patrick Lane, ScEYEnce Studios.
The authors also monitored the development of disease and found progressive changes at the chromatin level. Aberrant expression of DLX3, a homebox factor also linked to osteosarcoma, was identified as an early event. When DLX3 was overexpressed in normal cord blood cells, a change in the balance between GATA and ETS motifs was seen, features also observed in AMKL patients. In addition, ETO2::GLIS2 iPSCs-derived hematopoietic cells deficient in DLX3 had lost their leukemia-initiating capacity. Because ETO2::GLIS2-driven AMKL is relatively resistant to conventional high-intensity chemotherapy, alternative treatment strategies are needed to improve outcome for these patients. Targeting of the folate receptor 1 (FOLR1) has shown clinical potential,10 and the role of DLX3 in these leukemias, as demonstrated by Boudia et al, could pave the way for novel therapeutic approaches.
The work elegantly demonstrates how human iPSCs can be used to model the initiation of pediatric leukemia. By studying the ETO2::GLIS2 fusion in otherwise healthy cells, the onset of disease could be explored. Despite the challenges in generating transplantable iPS-derived hematopoietic cells, and the addition of only a single mutation, an engraftable leukemia was observed. Of clinical importance, the model may evolve into a drug screening platform for the development of novel treatment strategies.
Conflict-of-interest disclosure: The authors declare no competing financial interests.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal