Key Points
Genetic studies suggest HDAC3-selective suppression may prove useful for treatment of hematological tumors but will not induce apoptosis.
Genetic and pharmacological cosuppression of HDAC1 with HDAC2 induces a potent pro-apoptotic response of tumor cells.
Abstract
Histone deacetylase (HDAC) inhibitors (HDACis) have demonstrated activity in hematological and solid malignancies. Vorinostat, romidepsin, belinostat, and panobinostat are Food and Drug Administration–approved for hematological malignancies and inhibit class II and/or class I HDACs, including HDAC1, 2, 3, and 6. We combined genetic and pharmacological approaches to investigate whether suppression of individual or multiple Hdacs phenocopied broad-acting HDACis in 3 genetically distinct leukemias and lymphomas. Individual Hdacs were depleted in murine acute myeloid leukemias (MLL-AF9;NrasG12D; PML-RARα acute promyelocytic leukemia [APL] cells) and Eµ-Myc lymphoma in vitro and in vivo. Strikingly, Hdac3-depleted cells were selected against in competitive assays for all 3 tumor types. Decreased proliferation following Hdac3 knockdown was not prevented by BCL-2 overexpression, caspase inhibition, or knockout of Cdkn1a in Eµ-Myc lymphoma, and depletion of Hdac3 in vivo significantly reduced tumor burden. Interestingly, APL cells depleted of Hdac3 demonstrated a more differentiated phenotype. Consistent with these genetic studies, the HDAC3 inhibitor RGFP966 reduced proliferation of Eµ-Myc lymphoma and induced differentiation in APL. Genetic codepletion of Hdac1 with Hdac2 was pro-apoptotic in Eµ-Myc lymphoma in vitro and in vivo and was phenocopied by the HDAC1/2-specific agent RGFP233. This study demonstrates the importance of HDAC3 for the proliferation of leukemia and lymphoma cells, suggesting that HDAC3-selective inhibitors could prove useful for the treatment of hematological malignancies. Moreover, our results demonstrate that codepletion of Hdac1 with Hdac2 mediates a robust pro-apoptotic response. Our integrated genetic and pharmacological approach provides important insights into the individual or combinations of HDACs that could be prioritized for targeting in a range of hematological malignancies.
Introduction
Histone deacetylase (HDAC) inhibitors (HDACis) are gaining widespread use for treatment of hematological malignancies.1,2 The majority of HDACis target class I and/or II HDACs3 and it is unclear which isoforms are vital for tumor cell growth and/or survival. Moreover, it is yet to be established whether selective HDACis could improve antitumor efficacy and limit toxicity. HDACs modify the epigenome through regulated chromatin acetylation and are thought to control gene transcription.4 HDACs regulate expression of CDKN1A in many tumor types and are important cofactors in acute myeloid leukemia-1 (AML1)-ETO-driven AML.2,4-6 HDACs have therefore become promising targets for therapeutic intervention aiming to reverse aberrant epigenetic states associated with cancer.7
Several structurally diverse HDACis have been developed representing different chemical families and HDAC specificity.1,3,8 Vorinostat (Zolinza; Merck), romidepsin (Istodax; Celgene), belinostat (Beleodaq; Spectrum Pharmaceuticals), and panobinostat (Farydak; Novartis) are Food and Drug Administration (FDA)–approved for cutaneous/peripheral T-cell lymphoma and refractory multiple myeloma.9-12 There are 11 “classical” mammalian HDACs:3,13,14 class I HDACs (HDAC1, 2, 3, 8) are located primarily within the nucleus; class IIa HDACs (HDAC4, 5, 7, 9) shuttle between the nucleus and the cytoplasm; and class IIb HDACs (HDAC6, 10) contain 2 catalytic domains and are exclusively found in the cytoplasm. HDAC6 has substrate specificity for α-tubulin and class IV (HDAC11) has characteristics of both class I and II HDACs. Vorinostat, panobinostat, and belinostat inhibit HDAC1, 2, 3, and 6, whereas romidepsin has high affinity for HDAC1, 2, and 3.3
HDACis mediate a range of biological responses including: apoptosis; inhibition of cell-cycle progression; cellular differentiation; suppression of angiogenesis; and enhancing antitumor immunity.1 HDACs also regulate function, localization and/or stability of nonhistone proteins.15-17 For example, the acetylation of heat shock protein-90 (HSP90), a molecular chaperone, is regulated by HDAC6.18 As such, HSP90 client oncoproteins, including BCR-ABL and ERBB2, may be degraded via HDACi-mediated HSP90 deacetylation and have been proposed as a major effector of HDACi mechanism of action.13 The combined effects of histone and nonhistone hyperacetylation are likely critical for the therapeutic activity of HDACis.19
HDAC-selective inhibitors are being developed in the hope of mediating potent antitumor responses and reducing toxicities.20 However, whether more selective HDACis will deliver on this premise remains to be determined. Transient depletion of individual HDACs in human tumor cells using small interfering RNA has not conclusively demonstrated whether antitumor actions of broad-acting HDACis can be phenocopied by loss of individual or multiple HDACs.21-24 Knockdown of HDAC3, and to a lesser extent HDAC1 and 2, resulted in growth inhibition in human colon cancer cell lines; however, the biological response was less potent than vorinostat treatment.25 Depletion or pharmacological inhibition of HDAC3 triggered apoptosis in cutaneous T-cell lymphoma and multiple myeloma.21,22 Apoptotic effects in ovarian cancer cell lines following small interfering RNA-mediated knockdown of HDAC2, 4, 8, and 11 have been reported.23 These studies suggest suppression of a single HDAC may have antitumor effects; however, comprehensive screening approaches using multiple cell systems have not been implemented to date.
Here, we used 3 tractable murine hematological cancer models: MLL-AF9;NrasG12D-driven AML; PML-RARα–driven acute promyelocytic leukemia (APL); and Myc-driven B-cell lymphoma (Eµ-Myc) to assess the effects of genetically depleting or pharmacologically inhibiting individual HDAC isoforms on tumor cell growth and survival. We demonstrate a unique sensitivity of all 3 malignancies to depletion of Hdac3, but no other individual Hdac. Furthermore, depletion of both Hdac1 with Hdac2 induced apoptosis in Eµ-Myc lymphoma. These genetic studies were supported by experiments using pharmacological inhibitors of individual or multiple HDAC isoforms that phenocopied the effects of gene knockdown.
Materials
Cell lines
Antibodies to the following proteins were used: HDAC1 (ab7028; Abcam, Cambridge, UK); HDAC2 (ab7029); HDAC3 (ab7030); HDAC6 (no. 2162; Cell Signaling, Danvers, MA; no. ); acetylated tubulin (6-11B-1, T7451; Sigma-Aldrich, Castle Hill, Australia); acetylated H4(K5) (no. 9672; Millipore, Arundel, Australia); acetylated H4(K8) (no. 2594; Cell Signaling); acetylated H3(K14) (Millipore, 06-911); p21 (F-5; Santa Cruz, CA); β-actin (Sigma-Aldrich); and HSP90 (ADI-SPA-830; Sapphire Bioscience, Australia). Vorinostat was from Merck (Boston, MA), RGFP966 and RGFP233 from Repligen Corporation (Waltham, MA), Tubacin from Enzo (Sapphire Biosciences), ACY-1215 from Acetylon Pharmaceuticals (Boston, MA); and QVD was from Sigma-Aldrich.
NIH-3T3 and Phoenix cells were cultured in Dulbecco’s modified Eagle medium, fetal bovine serum, l-glutamine, and penicillin/streptomycin (Invitrogen, Melbourne, Australia). Eµ-Myc lymphoma cells were maintained as previously detailed.24 MLL-AF9;NrasG12D AML cells were generated as described.26 APL cells were generated as described.27 Mouse embryonic fibroblasts (MEFs) were maintained in complete Dulbecco’s modified Eagle medium plus β-mercaptoethanol.
RNA interference design
Short hairpin RNA (shRNA)mir30s targeting murine Hdac isoforms were designed as previously described (see supplemental Table 1 on the Blood Web site).28-30 Clones were sequence verified using the 5′miR30-ZUBER primer (supplemental Table 3). shRNAs used in APL experiments were generated as described (supplemental Table 2).28
Stable transductions of retroviral vectors
Phoenix cells were transfected using calcium phosphate.28,29 Viral supernatants were spun onto RetroNectin-coated plates (Takara, Clontech) followed by spinfection of tumor cells. APL cells were exposed to concentrated viral supernatants (5× cold PEG-it TM, System Biosciences). Flow cytometry was used to assess transduction efficiency (green fluorescence protein [GFP]/Venus, LSR II, Becton Dickinson).
Proliferation assays
Proliferation was assessed by: (1) competitive proliferation assays as described26 and percentages of GFP+/Venus+/dsRed+ cells were assessed using flow cytometry; (2) cell counting, whereby cells were plated (5 × 103 cells/well, 24-well plate), counted daily (4 days), replated (5 × 103/well) and recounted (4 days); (3) CellTrace Violet staining (CTV; 1-5 × 106 cells, Life Technologies, Mulgrave, Australia), fluorescence-activated cell sorter (FACS)-sorted, serially cultured, and assessed daily by flow cytometry (Canto II, Becton Dickinson); and (4) sorted (GFP+) APL cells (1 × 104 cells/plate) were seeded in methylcellulose medium (MethoCult SF M3434; Stem Cell Technology, Vancouver, Canada), incubated (7-10 days), colonies were scored, pooled, and cells used for immunolabeling, morphologic analysis, and serial replating.
Western blotting and qRT-PCR
Whole cell extracts were prepared and analyzed by western blot as described.31 RNA was obtained using Qiagen mini kits (Doncaster, Australia) or Trizol (Life Technologies, Mulgrave, Australia), converted to cDNA using M-MLV Reverse transcriptase (RNase H Minus, Point mutant) and random primers (Promega, Madison, WI, USA). SensiFast SYBR green fluorescent nucleic acid stain (Bioline, Alexandria, Australia) was used with quantitative real-time polymerase chain reaction (qRT-PCR) primers described (supplemental Table 3).
Assessment of apoptosis and cell cycle in Eµ-Myc cells
Apoptosis was assessed by Annexin V positivity (Becton Dickinson, Australia) with propidium iodide (Sigma-Aldrich) or Fluoro-Gold (Santa Cruz, Dallas, TX) by flow cytometry. Cell cycle was assessed using ClickIt-Edu kit (Life Technologies) as per kit instructions.
Immunophenotyping and morphologic analysis
Flow cytometry was performed on APL cells harvested from methylcellulose. Antibodies used for immunophenotyping were: Ly-6G (Gr-1, no. 25-5931-82; eBioscience, San Diego, CA) and CD11b (MAC1, no. 25-0112-82). Cytospins were stained using the May-Grunwald-Giemsa method (Sigma-Aldrich) and assessed as described elsewhere.32
In vivo depletion of Hdac3 in AML, Eµ-Myc lymphoma, and APL
Tet-on competent MLL-AF9;NrasG12D leukemia cells were transduced (pTRMPV-Neo), G418 selected (1 mg/mL, 6 days), transplanted (1 × 106) into sublethally irradiated (5.5 Gy) recipient mice (CD45.1; shRen.713, n = 12; shHdac3.987, n = 14), and fed dox (≥day 2). Whole-body bioluminescent imaging was undertaken (day 8, n = 4/group) as described.33 Bone marrow from leukemic mice was analyzed by flow cytometry for the percentage of Venus+/dsRed+ (shRNA-expressing) cells in donor-derived (CD45.2+) leukemia populations as previously described.33
Eµ-Myc lymphoma cells (no. 107, CD45.2+) were transduced (pTRMPV-Neo; shScr, shHdac3.1659, shHdac3.201), FACS-sorted, inoculated into CD45.1+ mice (5 × 103, n = 12/shRNA); mice were then fed dox (≥day 3). Mice (n = 6/group) were bled (retroorbital), euthanized, and organs removed (day 10); the remainder were euthanized at ethical end points.34 White blood cells (WBCs; Siemens Advia, Baywater, Australia) and tumor burden (Venus+ cells) were assessed in the peripheral blood (PB) was assessed by flow cytometry.
Transduced APL cells (shLuc or shHdac3) were FACS-sorted (GFP+) and inoculated (2 × 105 cells/mouse) into 129 SvEv mice (n = 10/cohort) and followed for leukemia development. Genomic DNA from leukemic animals was purified using a QIAamp DNA kit (QIAGEN, Valencia, CA) and PCR performed using primers specific for pRetroSuper (supplemental Table 3).
All in vivo experiments were approved by the animal ethics committees of individual institutes.
Assessment of RGFP966 and RGFP233 in Eµ-Myc lymphoma and/or APL in vitro
Generation of shHdac3 MEFs
Transgenic mice expressing shHdac3.1659 or shLuc.1309 were produced as described.37 MEFs were generated by intercrossing with CAGs-rtTA3 mice38 and harvesting of E13.5 fetuses. MEFS were plated in 24-well plates (1 × 105/well) and then serially passaged (±dox, 1 µg/mL). Expression of shRNA cassettes was assessed by flow cytometry (Venus+ cells) and target gene knockdown by western blot.
Generation of Hdac−/− Eµ-Myc lymphoma
Conditional Hdacfl/fl mice (Merck) were crossed with Eµ-Myc.Mx-Cre mice to generate Eµ-Myc.Mx-Cre.Hdacfl/fl lymphoma. Tumors were genotyped at end point combined with western blot or qRT-PCR to confirm deletion of individual Hdacs (supplemental Figure 7A-B). Eµ-Myc tumor cells displaying spontaneous Cre-mediated deletion (see Summers et al36 ) of Hdac1, Hdac2, or Hdac6 were used for subsequent experiments and designated Eµ-Myc.Hdac1−/−, Eµ-Myc.Hdac2−/−, or Eµ-Myc.Hdac6−/−.
Statistical analysis
All data including Kaplan-Meier survival curves were analyzed using appropriate statistical tests, including Student t test and one-way analysis of variance (ANOVA) with multiple comparisons (GraphPad Prism software). Flow cytometry data were analyzed using FlowJo Analysis software (Treestar). Data are expressed as mean ± standard error of the mean (SEM) and statistical significance assumed at P < .05.
Results
Systematic depletion of Hdac isoforms in AML, Eµ-Myc lymphoma, and APL uncovers sensitivities to depletion of Hdac3
We have previously observed potential sensitivities of aggressive MLL-AF9;NrasG12D AML to HDAC inhibition.26 Here we evaluated this by depleting Hdacs 1-11 using independently derived shRNAs and competitive assays to monitor their effects on representation of AML cells in vitro. In contrast to any other single Hdac, cells constitutively depleted of Hdac3 exhibited reduced representation during 12 days of serial passaging (Figure 1A). Hdac3 knockdown efficiency was evaluated by qRT-PCR (day 2) and correlated with the biological effects of depletion (Figure 1B). Validation assays showed dox-inducible depletion of Hdac3 caused antiproliferative effects similar to established control shRNAs, including shMyc.2105, shMyb.2652, and shMen1.2310 (Figure 1C) that target Myc, Myb, and Menin, respectively.
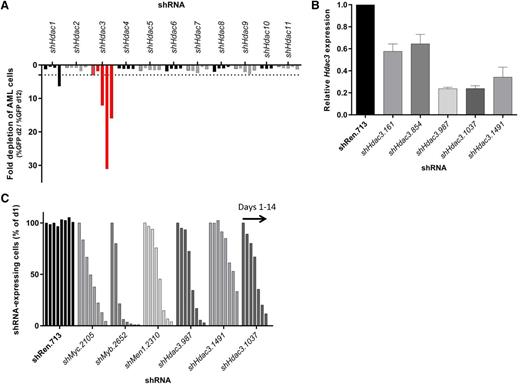
RNAi-mediated screen of all 11 classical Hdac isoforms in AML cells (MLL-AF9;NrasG12D) demonstrates a unique dependency on Hdac3 expression. Eleven Hdac isoforms were depleted in MLL-AF9;NrasG12D AML using multiple shRNAs per Hdac. Transduced AML cells (GFP+, shRNA-expressing) were mixed with nontransduced cells and followed for 10 days for cell representation by flow cytometry. (A) Relative depletion of GFP+ AML cells following constitutive (pLMN) depletion of individual Hdacs during 10 days of serial culture (Hdac3 shown in red). Data are plotted as fold change (GFP% day 2/GFP% day 12). A single experimental screen was undertaken with 4-5 distinct shRNAs per gene represented by individual bars. Dotted line depicts a 3-fold depletion cut off. (B) The efficiency of Hdac3 knockdown in AML cells was validated by qRT-PCR (day 2). Results were normalized to glyceraldehyde-3-phosphate dehydrogenase, whereas relative messenger RNA level in control cells (shRen.713) was set to 1 (n = 3-4 independent biological replicates). Data are presented as mean ± SEM. (C) Dox-inducible depletion of Hdac3 in AML. Each series of bars demonstrates a time course: days 1, 2, 4, 6, 8, 10, 12, and 14. The percentage of shRNA- expressing cells (Venus+/dsRed+) was normalized to day 1. Data are representative of a single experiment using 3 individual shRNAs to Hdac3 (shHdac3.987, shHdac3.1491, shHdac3.1037). Established control shRNAs to Myc, Myb, and MLL/AF9 cofactor Men1 were included for comparison.
RNAi-mediated screen of all 11 classical Hdac isoforms in AML cells (MLL-AF9;NrasG12D) demonstrates a unique dependency on Hdac3 expression. Eleven Hdac isoforms were depleted in MLL-AF9;NrasG12D AML using multiple shRNAs per Hdac. Transduced AML cells (GFP+, shRNA-expressing) were mixed with nontransduced cells and followed for 10 days for cell representation by flow cytometry. (A) Relative depletion of GFP+ AML cells following constitutive (pLMN) depletion of individual Hdacs during 10 days of serial culture (Hdac3 shown in red). Data are plotted as fold change (GFP% day 2/GFP% day 12). A single experimental screen was undertaken with 4-5 distinct shRNAs per gene represented by individual bars. Dotted line depicts a 3-fold depletion cut off. (B) The efficiency of Hdac3 knockdown in AML cells was validated by qRT-PCR (day 2). Results were normalized to glyceraldehyde-3-phosphate dehydrogenase, whereas relative messenger RNA level in control cells (shRen.713) was set to 1 (n = 3-4 independent biological replicates). Data are presented as mean ± SEM. (C) Dox-inducible depletion of Hdac3 in AML. Each series of bars demonstrates a time course: days 1, 2, 4, 6, 8, 10, 12, and 14. The percentage of shRNA- expressing cells (Venus+/dsRed+) was normalized to day 1. Data are representative of a single experiment using 3 individual shRNAs to Hdac3 (shHdac3.987, shHdac3.1491, shHdac3.1037). Established control shRNAs to Myc, Myb, and MLL/AF9 cofactor Men1 were included for comparison.
Next we focused on HDACs inhibited by FDA-approved HDACi, such as vorinostat (HDACs1, 2, 3, and 6) in Eμ-Myc lymphoma. An independent set of Hdac-directed shRNAs to those used in Figure 1 were validated in NIH-3T3 cells using constitutive (pLMS; supplemental Figure 1) or dox-inducible (pTRMPV-Neo; supplemental Figure 2) vectors by western blot. At least 2 shRNAs effectively depleted each individual Hdac and were used in subsequent experiments. Eµ-Myc (no. 4242) lymphoma cells were more sensitive to constitutive depletion of Hdac3 than to knockdown of Hdacs1, 2, or 6 (Figure 2A-B). Additionally, inducible depletion of Hdac3 reproducibly led to the loss of representation of Eµ-Myc (no. 4242) cells using multiple shRNAs (Figure 2C-E). A similar loss-of-representation phenotype was observed following knockdown of Hdac3 in a second Eµ-Myc lymphoma (no. 107; supplemental Figure 3A-B).
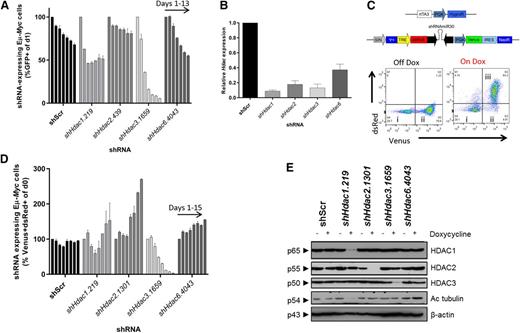
Systematic shRNA-mediated screen of individual Hdac isoforms uncovers sensitivity of Eµ-Myc lymphoma cells to depletion of Hdac3. Eµ-Myc lymphoma cells (no. 4242) were transduced with constitutive (pLMS) or dox-inducible (pTRMPV-Neo) retroviral vectors expressing shRNAs targeting HDACs1, 2, 3, or 6. Cells were isolated by FACS and then serially passaged (±dox) for up to 15 days. The percentage of shRNA-expressing cells (GFP+, pLMS; or Venus+/dsRed+, pTRMPV-Neo) were assessed by flow cytometry (days 1, 3, 5, 7, 9, 11, 13) and data were normalized to day 1, as depicted. (A) Constitutive depletion of individual Hdacs in Eµ-Myc cells in vitro (n = 2 biological replicates). Data are presented as mean ± SEM. (B) qRT-PCR was used to determine the efficiency of Hdac depletion in Eµ-Myc cells (day 3). Results were normalized to L32, whereas relative mRNA level in control cells (shScr) was set to 1 (n = 3-4 biological replicates). Data are presented as mean ± SEM. (C) Schematic representation of the dox-inducible vector system (pTRMPV-Neo). Representative dot plots demonstrating: (i) nontransduced (Venus−) cells; (ii) transduced (Venus+) cells; and (iii) shScr-expressing (Venus+/dsRed+) Eµ-Myc cells from at least 3 independent experiments. (D) Dox-inducible depletion of individual Hdacs in Eµ-Myc tumor cells in vitro (no. 4242; n = 2 independent experiments; days 1, 3, 5, 7, 9, 11, 13, 15). Data are presented as mean ± SEM. (E) Western blotting was used to demonstrate the efficiency of inducible Hdac depletion in Eµ-Myc cells (day 3). Hyperacetylated tubulin (Ac tubulin) was used as a surrogate readout for Hdac6 depletion, whereas changes to its levels in non-shHdac6 samples represent background variability (for example shHdac1). A representative experiment from 3 biological replicates is shown. Molecular weights of individual proteins are to the left of each blot.
Systematic shRNA-mediated screen of individual Hdac isoforms uncovers sensitivity of Eµ-Myc lymphoma cells to depletion of Hdac3. Eµ-Myc lymphoma cells (no. 4242) were transduced with constitutive (pLMS) or dox-inducible (pTRMPV-Neo) retroviral vectors expressing shRNAs targeting HDACs1, 2, 3, or 6. Cells were isolated by FACS and then serially passaged (±dox) for up to 15 days. The percentage of shRNA-expressing cells (GFP+, pLMS; or Venus+/dsRed+, pTRMPV-Neo) were assessed by flow cytometry (days 1, 3, 5, 7, 9, 11, 13) and data were normalized to day 1, as depicted. (A) Constitutive depletion of individual Hdacs in Eµ-Myc cells in vitro (n = 2 biological replicates). Data are presented as mean ± SEM. (B) qRT-PCR was used to determine the efficiency of Hdac depletion in Eµ-Myc cells (day 3). Results were normalized to L32, whereas relative mRNA level in control cells (shScr) was set to 1 (n = 3-4 biological replicates). Data are presented as mean ± SEM. (C) Schematic representation of the dox-inducible vector system (pTRMPV-Neo). Representative dot plots demonstrating: (i) nontransduced (Venus−) cells; (ii) transduced (Venus+) cells; and (iii) shScr-expressing (Venus+/dsRed+) Eµ-Myc cells from at least 3 independent experiments. (D) Dox-inducible depletion of individual Hdacs in Eµ-Myc tumor cells in vitro (no. 4242; n = 2 independent experiments; days 1, 3, 5, 7, 9, 11, 13, 15). Data are presented as mean ± SEM. (E) Western blotting was used to demonstrate the efficiency of inducible Hdac depletion in Eµ-Myc cells (day 3). Hyperacetylated tubulin (Ac tubulin) was used as a surrogate readout for Hdac6 depletion, whereas changes to its levels in non-shHdac6 samples represent background variability (for example shHdac1). A representative experiment from 3 biological replicates is shown. Molecular weights of individual proteins are to the left of each blot.
We recently reported that Hdac1 suppressed the development of APL (PML-RAR), but had little role in maintenance of established APL cells.28 Therefore, we wished to investigate the effects of Hdac3 depletion in this malignancy. Using independently derived shRNAs, depletion of Hdac3 (Figure 3A-B) significantly reduced the clonogenic potential of APL cells (Figure 3C). Interestingly, morphologic analysis of APL cells showed that Hdac3 depletion also induced a more mature cell phenotype (Figure 3Di-ii).
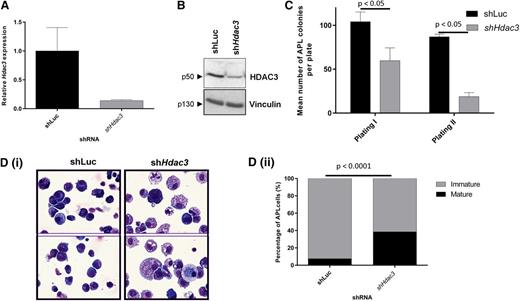
In vitro depletion of Hdac3 reduces growth and triggers differentiation in established APL cells. APL blasts from 129SvEv mice were transduced with constitutive pRetroSuper vectors expressing shHdac3 or control shLuc and sorted for GFP positivity, then assessed for (A) depletion of Hdac3 by qRT-PCR. Values are normalized against glyceraldehyde-3-phosphate dehydrogenase and referred to CTRL (shLuc) (n = 2 biological replicates). Data are presented as mean ± SEM. (B) Depletion of HDAC3 by western blot; Vinculin was used as a loading control. Molecular weights of individual proteins are to the left of each blot. (C) Loss of clonogenicity by serial replating assay (data are presented as mean number of colonies counted 7-10 days after seeding 1 × 104 leukemic cells ± SEM (n = 3 independent experiments). Data are presented as mean ± SEM. Statistical analysis was performed with a paired t test. (D) Differentiation by morphologic analysis of the GFP+/APL cells, harvested after plating in methylcellulose medium (first methylcellulose): (i) representative cytospins; (ii) percentage of mature and immature cells (×60 magnification, May Grünwald-Giemsa staining, Olympus BX51). The prevalence of mature and immature cells was analyzed morphologically in cytological slides and the absolute percentage of mature cells was reported. Experiments were repeated 3 times (biological triplicate). At least 300 cells were scanned for each case. Statistical analysis was performed with the Fisher exact test.
In vitro depletion of Hdac3 reduces growth and triggers differentiation in established APL cells. APL blasts from 129SvEv mice were transduced with constitutive pRetroSuper vectors expressing shHdac3 or control shLuc and sorted for GFP positivity, then assessed for (A) depletion of Hdac3 by qRT-PCR. Values are normalized against glyceraldehyde-3-phosphate dehydrogenase and referred to CTRL (shLuc) (n = 2 biological replicates). Data are presented as mean ± SEM. (B) Depletion of HDAC3 by western blot; Vinculin was used as a loading control. Molecular weights of individual proteins are to the left of each blot. (C) Loss of clonogenicity by serial replating assay (data are presented as mean number of colonies counted 7-10 days after seeding 1 × 104 leukemic cells ± SEM (n = 3 independent experiments). Data are presented as mean ± SEM. Statistical analysis was performed with a paired t test. (D) Differentiation by morphologic analysis of the GFP+/APL cells, harvested after plating in methylcellulose medium (first methylcellulose): (i) representative cytospins; (ii) percentage of mature and immature cells (×60 magnification, May Grünwald-Giemsa staining, Olympus BX51). The prevalence of mature and immature cells was analyzed morphologically in cytological slides and the absolute percentage of mature cells was reported. Experiments were repeated 3 times (biological triplicate). At least 300 cells were scanned for each case. Statistical analysis was performed with the Fisher exact test.
To determine whether Hdac3 depletion was detrimental to growth/survival of nontumor cells, we knocked down Hdac3 in MEFs. Hdac3 depletion had no significant effect on the growth and survival of MEFs up to 13 days following depletion (supplemental Figure 4A; P > .05).
Depletion of Hdac3 induces an antiproliferative response that is not affected by genetic or pharmacological inhibition of apoptosis
After reports suggested that HDAC3 suppression induces tumor cell apoptosis,21,36 we investigated whether depletion of Hdac3 had a pro-apoptotic effect in Eµ-Myc cells. No apoptosis induction above background levels was detected in Hdac3-depleted cells or cells depleted of any other single Hdac isoform (Figure 4A; supplemental Figure 3). In contrast, treatment of Eµ-Myc cells with vorinostat mediated a robust apoptotic response, as we had previously observed (Figure 4A).39 To further demonstrate that induction of apoptosis did not contribute to the loss of cells following knockdown of Hdac3, pro-survival Bcl-2 was overexpressed in Eµ-Myc cells (Eµ-Myc.Bcl-2; supplemental Figure 3C) because this is reported to prevent vorinostat-mediated apoptosis via the intrinsic apoptotic pathway.39 Hdac3 depletion continued to invoke a loss of representation phenotype in Bcl-2–overexpressing cells (Figure 4B-C). Addition of the caspase inhibitor QVD had no effect on suppression of proliferation observed following Hdac3 depletion, but QVD did suppress vorinostat-induced apoptosis (Figure 4D). Surprisingly, we did not detect any acute changes in cell cycle following Hdac3 depletion using sensitive Edu labeling and pulse chase assays (supplemental Figure 5A). Taken together, these data demonstrate that depletion of Hdac3 in Eµ-Myc cells does not induce apoptosis or arrest within any specific phase of the cell cycle, and an intact apoptotic response is not required for the antiproliferative effects of Hdac3 depletion.
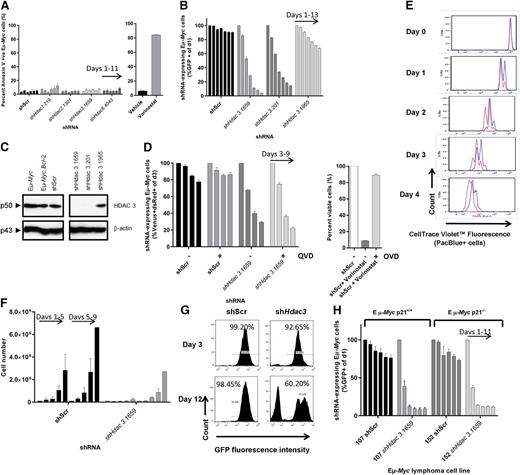
Depletion of Hdac3 in Eµ-Myc lymphoma induces an antiproliferative response that is not affected by genetic or pharmacological apoptosis inhibitors or by loss of p21WAF1/CIP1. Eµ-Myc lymphoma cells were transduced with pTRMPV-Neo vectors expressing shHdac3 or control shScr cassettes, isolated by FACS (100% GFP+), serially passaged (±dox) for up to 15 days. (A) The induction of apoptosis was assessed by phosphatidylserine externalization (Annexin V+) using flow cytometry. As a positive control, we treated Eµ-Myc lymphoma cells with vorinostat (24 hours, 1 µM). Data are presented as percentage of Annexin V positive cells (mean ± SEM, n = 2 biological replicates). Individual bars demonstrate day of analysis (days 1, 3, 5, 7, 9, 11). (B-C) Eµ-Myc lymphoma cells (no. 107) were stably transduced with pMSCV.Bcl-2-mCherry to overexpress pro-survival Bcl-2. Eµ-Myc.Bcl-2 lymphoma cells were then transduced with constitutive vectors expressing shRNAs against Hdac3 or shScr, serially passaged, and assessed by (B) competitive proliferation assay where the percentage of GFP+ cells were normalized to day 1 and individual bars represent days of analysis (days 1, 3, 5, 7, 9, 11, 13; 3 individual shRNAs were tested; 2 biological replicates) and (C) HDAC3 depletion was confirmed by western blot. A representative experiment from 3 biological replicates is shown. Molecular weights of individual proteins are to the left of the blot. (D) Caspase activation was inhibited by treating Eµ-Myc lymphoma cells expressing dox-inducible shRNAs with pan-caspase inhibitor QVD (10 µM, n = 3 biological replicates); cell growth was assessed using competitive proliferation assays (±dox). Data are presented as mean percentage of shRNA-expressing cells (Venus+/dsRed+, dox-treated only, days 3, 5, 7, 9) and normalized to day 3 ± SEM. The activity of QVD was confirmed in Eµ-Myc cells treated with vorinostat (1 µM) ± QVD (presented as percentage viable cells). The proliferation of Eµ-Myc cells depleted of Hdac3 was measured using a carboxyfluorescein diacetate succinimidyl ester–like assay (CTV) and by cell counting/replating assays. (E) Eµ-Myc cells (no. 107) were transduced with constitutive (pLMS) vectors expressing shHdac3.1659 or shScr, immediately stained with CTV, allowed to expand in culture overnight, and then FACS-sorted to a single population of GFP+/CTV+ (PacBlue+) cells. Cells were serially cultured for up to 5 days and underwent daily assessment of their proliferative capacity by flow cytometry (ie, loss of CTV cells over time; shScr, red line; shHdac3, blue line). Individual plots represent daily flow cytometry analysis and are representative of 3 individual biological experiments. (F) Eµ-Myc lymphoma cells constitutively depleted of Hdac3 were seeded (5 × 103 cells; day 1) into 24-well plates (shScr vs shHdac3) followed by daily cell counts (individual bars represent days 1, 2, 3, 4, 5), then replated (5× 103 cells; day 5) and counted daily (n = 3 biological replicates; bars represent days 5, 6, 7, 8, 9). Data are presented as mean ± SEM. (G) Representative histograms demonstrate the percentages of shRNA-expressing (GFP+) Eµ-Myc cells on days 3 and 12 of cell counting/replating assay. (H) Eµ-Myc.Cdkn1a+/+ (no. 107) or Eµ-Myc.Cdkn1a−/− (no. 152) lymphoma cells were depleted of Hdac3 and cell proliferation was assessed by competitive proliferation assay. Individual bars represent percentages of shRNA-expressing (GFP+) Eµ-Myc cells normalized to day 1 (n = 3 biological replicates; individual bars represent days 1, 3, 5, 7, 9, 11). Data are presented as mean ± SEM.
Depletion of Hdac3 in Eµ-Myc lymphoma induces an antiproliferative response that is not affected by genetic or pharmacological apoptosis inhibitors or by loss of p21WAF1/CIP1. Eµ-Myc lymphoma cells were transduced with pTRMPV-Neo vectors expressing shHdac3 or control shScr cassettes, isolated by FACS (100% GFP+), serially passaged (±dox) for up to 15 days. (A) The induction of apoptosis was assessed by phosphatidylserine externalization (Annexin V+) using flow cytometry. As a positive control, we treated Eµ-Myc lymphoma cells with vorinostat (24 hours, 1 µM). Data are presented as percentage of Annexin V positive cells (mean ± SEM, n = 2 biological replicates). Individual bars demonstrate day of analysis (days 1, 3, 5, 7, 9, 11). (B-C) Eµ-Myc lymphoma cells (no. 107) were stably transduced with pMSCV.Bcl-2-mCherry to overexpress pro-survival Bcl-2. Eµ-Myc.Bcl-2 lymphoma cells were then transduced with constitutive vectors expressing shRNAs against Hdac3 or shScr, serially passaged, and assessed by (B) competitive proliferation assay where the percentage of GFP+ cells were normalized to day 1 and individual bars represent days of analysis (days 1, 3, 5, 7, 9, 11, 13; 3 individual shRNAs were tested; 2 biological replicates) and (C) HDAC3 depletion was confirmed by western blot. A representative experiment from 3 biological replicates is shown. Molecular weights of individual proteins are to the left of the blot. (D) Caspase activation was inhibited by treating Eµ-Myc lymphoma cells expressing dox-inducible shRNAs with pan-caspase inhibitor QVD (10 µM, n = 3 biological replicates); cell growth was assessed using competitive proliferation assays (±dox). Data are presented as mean percentage of shRNA-expressing cells (Venus+/dsRed+, dox-treated only, days 3, 5, 7, 9) and normalized to day 3 ± SEM. The activity of QVD was confirmed in Eµ-Myc cells treated with vorinostat (1 µM) ± QVD (presented as percentage viable cells). The proliferation of Eµ-Myc cells depleted of Hdac3 was measured using a carboxyfluorescein diacetate succinimidyl ester–like assay (CTV) and by cell counting/replating assays. (E) Eµ-Myc cells (no. 107) were transduced with constitutive (pLMS) vectors expressing shHdac3.1659 or shScr, immediately stained with CTV, allowed to expand in culture overnight, and then FACS-sorted to a single population of GFP+/CTV+ (PacBlue+) cells. Cells were serially cultured for up to 5 days and underwent daily assessment of their proliferative capacity by flow cytometry (ie, loss of CTV cells over time; shScr, red line; shHdac3, blue line). Individual plots represent daily flow cytometry analysis and are representative of 3 individual biological experiments. (F) Eµ-Myc lymphoma cells constitutively depleted of Hdac3 were seeded (5 × 103 cells; day 1) into 24-well plates (shScr vs shHdac3) followed by daily cell counts (individual bars represent days 1, 2, 3, 4, 5), then replated (5× 103 cells; day 5) and counted daily (n = 3 biological replicates; bars represent days 5, 6, 7, 8, 9). Data are presented as mean ± SEM. (G) Representative histograms demonstrate the percentages of shRNA-expressing (GFP+) Eµ-Myc cells on days 3 and 12 of cell counting/replating assay. (H) Eµ-Myc.Cdkn1a+/+ (no. 107) or Eµ-Myc.Cdkn1a−/− (no. 152) lymphoma cells were depleted of Hdac3 and cell proliferation was assessed by competitive proliferation assay. Individual bars represent percentages of shRNA-expressing (GFP+) Eµ-Myc cells normalized to day 1 (n = 3 biological replicates; individual bars represent days 1, 3, 5, 7, 9, 11). Data are presented as mean ± SEM.
Next we investigated whether depletion of Hdac3 affected the proliferative capacity of lymphoma cells. Eµ-Myc cells depleted of Hdac3 proliferated more slowly than those expressing the control shRNA (Figure 4E). In addition, cell counting/replating assays conclusively showed that the proliferation of Eµ-Myc lymphoma was significantly attenuated when Hdac3 was depleted (Figure 4F). Assessment of cell populations at the experiment’s end demonstrated outgrowth of GFP− cells not expressing shHdac3, suggesting selection of cells without Hdac3 depletion (Figure 4G). Overall, our results suggest that depletion of Hdac3 in Eµ-Myc lymphoma leads to a loss of proliferation phenotype that is independent of apoptosis.
Loss of p21WAF1/CIP1 does not affect the antiproliferative response mediated by Hdac3 depletion
Previous investigators have reported changes to the expression of various cell-cycle/checkpoint proteins, including p15INK4b,40 p21WAF1/CIP1,21,40 p27,41 p53,41,42 p57,43 and/or Rb44,45 following Hdac3 depletion. Therefore, we probed for the expression of all mentioned cell-cycle regulatory proteins in Hdac3-depleted Eµ-Myc cells by western blot (data not shown). Interestingly, only p21WAF1/CIP1 was reproducibly upregulated in a time-dependent manner, such that potent shHdac3.1659 increased p21WAF1/CIP1 levels by day 3, whereas the less potent shHdac3.201 increased p21WAF1/CIP1 expression at day 6 (supplemental Figure 5B). Assessment of Cdkn1a expression by qRT-PCR demonstrated that depletion of Hdac3 had no effect on the transcription of Cdkn1a, suggesting posttranslational regulation/stabilization of p21WAF1/CIP1 (supplemental Figure 5C). Genetic deletion of Cdkn1a (Eµ-Myc.Cdkn1a-/-) did not prevent the antiproliferative effects observed following Hdac3 depletion (Figure 4H; supplemental Figure 5D). These results suggest that p21WAF1/CIP1 regulation/stabilization did not mediate the growth suppressive effects of Hdac3 depletion.
In vivo Hdac3 depletion reduces tumor burden and/or significantly extends the survival of mice bearing AML, Eµ-Myc lymphoma, or APL cells
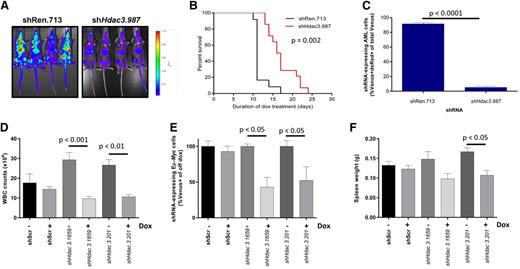
Our in vitro studies reproducibly demonstrated that Hdac3 depletion reduced the proliferation of AML, Eµ-Myc lymphoma, and APL cells; we sought to confirm this in vivo. We inoculated CD45.1+ congenic C57BL/6 mice with luciferase-expressing AML cells transduced with dox-inducible shHdac3.987 (n = 14) or nontargeting shRen.713 (n = 12) and initiated Hdac3 depletion 2 days after transplantation. Remarkably, in vivo depletion of Hdac3 significantly reduced tumor burden (Figure 5A) and provided significant survival benefit in mice bearing AML (Figure 5B). At the terminal disease stage, bone marrow of control mice predominantly showed shRen.713-expressing cells (Venus+/dsRed+), whereas recipients of shHdac3.987-expressing cells showed an outgrowth of AML cells that had evaded shRNA expression (CD45.2+/ Venus+/dsRed−; Figure 5C; supplemental Figure 6A), indicating a strong selection against effective Hdac3 suppression.34
In vivo Hdac3 depletion reduces tumor burden and/or significantly extends the survival of mice bearing AML or Eµ-Myc lymphoma. MLL-AF9;NrasG12D AML cells transduced with dox-inducible shRNA constructs (pTRMPV-Neo) were transplanted into CD45.1+ mice and shRNA expression was induced 2 days after tumor inoculation by addition of dox to food and drinking water (shRen.713, n = 14; shHdac3.987, n = 12). (A) Tumor burden was assessed by bioluminescent imaging following 8 days of dox treatment. (B) Kaplan-Meier curves for survival analysis of mice bearing transplanted AML tumor with indicated pTRMPV-Neo constructs. Day 0 denotes the beginning of dox treatment. Statistical analysis was undertaken using a log-rank (Mantel-Cox) test. (C) Percentage of shRNA-expressing (Venus+/dsRed+) tumor cells in the bone marrow remaining at terminal disease stage were analyzed using a Student t test (P < .0001). Eµ-Myc tumor cells (no. 107) were transduced with dox-inducible pTRMPV-Neo with shRNA cassettes targeting Hdac3 (shHdac3.1659, n = 12; shHdac3.201, n = 12) or shScr control (n = 12), FACS-sorted, and transplanted into CD45.1+ mice (5 × 103 cells per mouse). On day 3 postinoculation, mice (n = 6/group) were fed dox in food and water to initiate expression of shRNAs in vivo. Mice were bled and sacrificed on day 10 to assess (D) WBC count, (E) percentage of tumor cells (Venus+) in PB by flow cytometry, and (F) spleen size in mice bearing Eµ-Myc lymphoma. Data are presented as mean ± SEM. Two biological experiments were undertaken. Data were analyzed using one-way ANOVAs and appropriate post-hoc tests.
In vivo Hdac3 depletion reduces tumor burden and/or significantly extends the survival of mice bearing AML or Eµ-Myc lymphoma. MLL-AF9;NrasG12D AML cells transduced with dox-inducible shRNA constructs (pTRMPV-Neo) were transplanted into CD45.1+ mice and shRNA expression was induced 2 days after tumor inoculation by addition of dox to food and drinking water (shRen.713, n = 14; shHdac3.987, n = 12). (A) Tumor burden was assessed by bioluminescent imaging following 8 days of dox treatment. (B) Kaplan-Meier curves for survival analysis of mice bearing transplanted AML tumor with indicated pTRMPV-Neo constructs. Day 0 denotes the beginning of dox treatment. Statistical analysis was undertaken using a log-rank (Mantel-Cox) test. (C) Percentage of shRNA-expressing (Venus+/dsRed+) tumor cells in the bone marrow remaining at terminal disease stage were analyzed using a Student t test (P < .0001). Eµ-Myc tumor cells (no. 107) were transduced with dox-inducible pTRMPV-Neo with shRNA cassettes targeting Hdac3 (shHdac3.1659, n = 12; shHdac3.201, n = 12) or shScr control (n = 12), FACS-sorted, and transplanted into CD45.1+ mice (5 × 103 cells per mouse). On day 3 postinoculation, mice (n = 6/group) were fed dox in food and water to initiate expression of shRNAs in vivo. Mice were bled and sacrificed on day 10 to assess (D) WBC count, (E) percentage of tumor cells (Venus+) in PB by flow cytometry, and (F) spleen size in mice bearing Eµ-Myc lymphoma. Data are presented as mean ± SEM. Two biological experiments were undertaken. Data were analyzed using one-way ANOVAs and appropriate post-hoc tests.
We next transplanted Eµ-Myc lymphoma (no. 107) transduced with dox-inducible shScr, shHdac3.1659, or shHdac3.201 into CD45.1+ congenic C57BL/6 mice (n = 36) and allowed engraftment before dox treatment (day 3). In vivo depletion of Hdac3 significantly reduced WBC count (Figure 5E), the percentage of Venus+ tumor cells in PB (Figure 5F), and spleen weight (Figure 5G). Consistent with the data from the AML system, we observed outgrowth of nontransduced Eµ-Myc cells (Venus−; supplemental Figure 6B) in mice at ethical end points preventing any survival advantage (data not shown).
We then knocked down Hdac3 (or control Luc) in frankly leukemic APL cells. Sorted GFP+ APL cells were transplanted into 129 SvEv recipient mice and engraftment followed by analysis of PB. Inoculation of GFP+/shHdac3 APL cells resulted in no, or very few, GFP+ cells detected in PB samples, whereas control cells grew exponentially (data not shown). Although all control mice (n = 10) developed APL and were euthanized by 50 days after transplantation, 7 of 10 GFP+/shHdac3 mice remained disease-free for >300 days, whereas the remaining 3 of 10 GFP+/shHdac3 mice developed APL (supplemental Figure 6C). In these mice, APL cells resident at the end of the experiment showed WT Hdac3 expression, suggesting selection against Hdac3 depletion (supplemental Figure 6D). Additionally, PCR analysis from genomic DNA extracted from spleens of 2 leukemic mice confirmed that the leukemic cells originated from nontransduced cells that are present in small numbers (<1%) upon sorting because we were unable to amplify a DNA fragment corresponding to the vector used to transduce cells that should have been stably integrated into the host genome (supplemental Figure 6E).
HDAC3-selective RGFP966 reproduces Hdac3 depletion in Eµ-Myc lymphoma and APL
Next we asked whether the biological effects of Hdac3 depletion could be phenocopied pharmacologically with the HDAC3-selective inhibitor, RGFP966.22,35 Eµ-Myc lymphoma cells did not undergo apoptosis following incubation with RGFP966 (≤1 µM, 48 hours; Figure 6A) but proliferated significantly more slowly than vehicle-treated controls in the presence or absence of pro-survival BCL-2 overexpression (Figure 6B), confirming our genetic depletion studies (see supplemental Figure 7A for biomarker studies). Similarly, RGFP966 did not induce apoptosis in APL cells (supplemental Figure 7B; ≤1 μM) but did reduce clonogenicity and increased maturation (Figure 6C-F). Treatment of wild-type hematopoietic progenitor cells with RGFP966 had only minor effects on cell clonogenicity (data not shown).
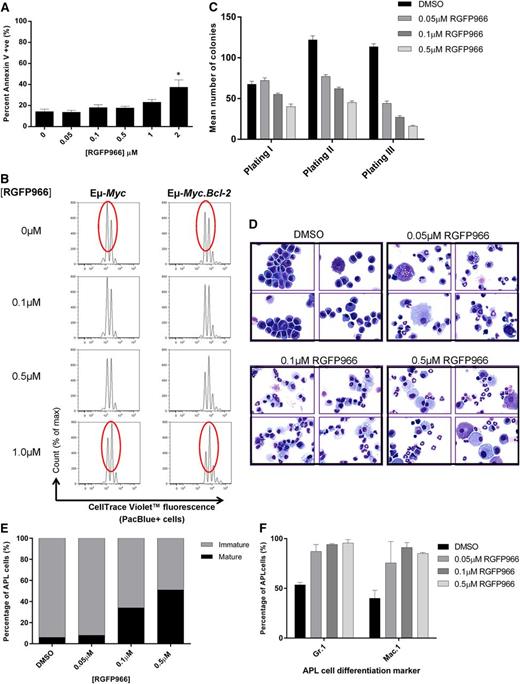
Treatment of Eµ-Myc lymphoma or APL cells with HDAC3-selective RGFP966 mimics HDAC3 depletion. Eµ-Myc lymphoma and APL cells were treated with low micromolar concentrations of RGFP966 (≤2 µM) in vitro and assessed for apoptosis and cell proliferation. (A) Assessment of apoptosis (Annexin V/PI) in Eµ-Myc cells (no. 107) after 48 hours of treatment with RGFP966 using flow cytometry (n = 3 biological replicates; *P < .05). Data were analyzed using a one-way ANOVA. (B) Proliferation of Eµ-Myc (no. 107, n = 3 biological replicates) and Eµ-Myc.Bcl-2 (n = 2 biological replicates) cells was assessed using CTV staining following RGFP966 treatment (≤1 µM, 48 hours). Cells were analyzed for cell division and measured as discrete peaks of decreasing CTV fluorescence (PacBlue+ cells) using flow cytometry. Red ovals highlight the CTV peaks that demonstrate anti-proliferative effects of RGFP966. (C) APL cells were treated with RGFP966 at the indicated concentrations and subjected to colony-forming assays whereby cells were plated in methylcellulose (1 × 104 cells), left for 7-10 days, counted, and replated. Serial replating of RGFP966-treated APL cells led to a dose-dependent reduction in colony forming potential. Data are presented as mean number of colonies counted after 7-10 days ± SEM from 3 biological replicates. (D) Histological assessment of APL cell maturity demonstrates that RGFP966 treatment increases the population of mature vs immature cells in vitro (×60 magnification; ×4 representative cytospins; (top left) dimethylsulfoxide (DMSO); (top right) 0.05 µM RGFP966; (bottom left) 0.1 µM RGFP966; (bottom right) 0.5 µM RGFP966). At least 300 cells were scanned for each case (n = 3 biological replicates). (E) Calculated percentage of mature and immature cells following treatment with 0.05 µM, 0.1 µM, or 0.5 µM RGFP966. In addition, APL cells treated with RGFP966 at indicated concentrations were immunophenotyped by flow cytometry. (F) An increased percentage of cells positive for Gr.1(Ly-6G) or Mac.1 (CD11b) confirms that low concentrations of RGFP966 triggers differentiation in APL cells similar to Hdac3 depletion (n = 2 biological replicates).
Treatment of Eµ-Myc lymphoma or APL cells with HDAC3-selective RGFP966 mimics HDAC3 depletion. Eµ-Myc lymphoma and APL cells were treated with low micromolar concentrations of RGFP966 (≤2 µM) in vitro and assessed for apoptosis and cell proliferation. (A) Assessment of apoptosis (Annexin V/PI) in Eµ-Myc cells (no. 107) after 48 hours of treatment with RGFP966 using flow cytometry (n = 3 biological replicates; *P < .05). Data were analyzed using a one-way ANOVA. (B) Proliferation of Eµ-Myc (no. 107, n = 3 biological replicates) and Eµ-Myc.Bcl-2 (n = 2 biological replicates) cells was assessed using CTV staining following RGFP966 treatment (≤1 µM, 48 hours). Cells were analyzed for cell division and measured as discrete peaks of decreasing CTV fluorescence (PacBlue+ cells) using flow cytometry. Red ovals highlight the CTV peaks that demonstrate anti-proliferative effects of RGFP966. (C) APL cells were treated with RGFP966 at the indicated concentrations and subjected to colony-forming assays whereby cells were plated in methylcellulose (1 × 104 cells), left for 7-10 days, counted, and replated. Serial replating of RGFP966-treated APL cells led to a dose-dependent reduction in colony forming potential. Data are presented as mean number of colonies counted after 7-10 days ± SEM from 3 biological replicates. (D) Histological assessment of APL cell maturity demonstrates that RGFP966 treatment increases the population of mature vs immature cells in vitro (×60 magnification; ×4 representative cytospins; (top left) dimethylsulfoxide (DMSO); (top right) 0.05 µM RGFP966; (bottom left) 0.1 µM RGFP966; (bottom right) 0.5 µM RGFP966). At least 300 cells were scanned for each case (n = 3 biological replicates). (E) Calculated percentage of mature and immature cells following treatment with 0.05 µM, 0.1 µM, or 0.5 µM RGFP966. In addition, APL cells treated with RGFP966 at indicated concentrations were immunophenotyped by flow cytometry. (F) An increased percentage of cells positive for Gr.1(Ly-6G) or Mac.1 (CD11b) confirms that low concentrations of RGFP966 triggers differentiation in APL cells similar to Hdac3 depletion (n = 2 biological replicates).
A pro-apoptotic effect requires the depletion of Hdac1 and Hdac2 in Eµ-Myc lymphoma
We next exploited our ability to delete/deplete multiple Hdacs to determine whether loss of more than a single Hdac isoform could induce a pro-apoptotic response. Here we deleted Hdac1, Hdac2, or Hdac6 (supplemental Figure 8A-C) combined with knockdown of Hdac1, Hdac2, Hdac3, or Hdac6 in Eµ-Myc lymphoma. Combined deletion/depletion of Hdac1 with Hdac2 rapidly reduced the representation of Eµ-Myc cells to a greater extent than single isoform depletion experiments. Hdac6 knockdown had no effect, whereas the antiproliferative effect of Hdac3 depletion was not further potentiated by Hdac1 deletion (Figure 7A) or Hdac2 deletion (data not shown). Reciprocal knockout of Hdac2 combined with Hdac1 depletion confirmed this response (supplemental Figure 8D-F). We then depleted Hdacs1-3 in Eµ-Myc.Hdac6−/− lymphoma and observed that only combined depletion of Hdac6 with Hdac3 reduced the representation of tumor cells (Figure 7B). However, the antiproliferative effects of combined Hdac3 with Hdac6 depletion/deletion were not greater than those observed following Hdac3 depletion alone.
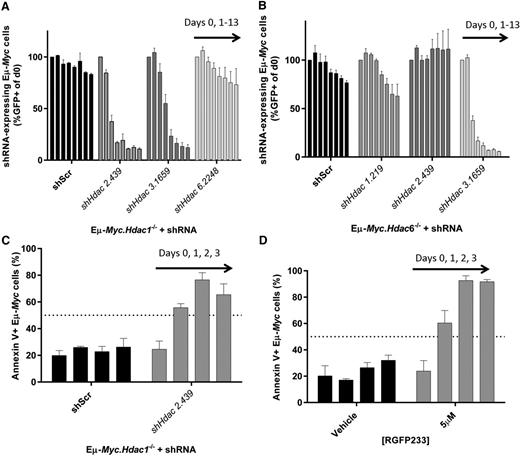
A pro-apoptotic effect requires the depletion of Hdac1 and Hdac2 in Eµ-Myc lymphoma. We produced tractable Eµ-Myc lymphoma cells with conditional knockout of Hdac1, Hdac2, or Hdac6 (Eµ-Myc.Hdac1−/−; Eµ-Myc.Hdac2−/−; Eµ-Myc.Hdac6−/−) and transduced these with shRNAs against Hdac1, Hdac2, Hdac3, or Hdac6. (A) Eµ-Myc.Hdac1−/− lymphoma was transduced with constitutive (pLMS) vectors containing shRNA cassettes against Hdac2, Hdac3, or Hdac6, FACS-sorted to approximately 50% GFP+, and 50% GFP−, and cell representation followed over time using competitive proliferation assays, as described previously. Individual bars represent daily flow cytometry assessment of GFP positive cells (days 0, 1, 3, 5, 7, 9, 11, 13) from 3 independent replicates (mean ± SEM) and normalized to day 0. (B) Effects of constitutively depleting Hdac1, Hdac2, or Hdac3 in Eµ-Myc.Hdac6−/− cells using competitive assays. Each bar represents different days of testing: days 0, 1, 3, 5, 7, 9, 11, 13. Data are presented as mean ± SEM of 3 biological replicates. (C) Eµ-Myc.Hdac1−/− cells transduced with shHdac2.439 were FACS-sorted (GFP+) immediately following transduction and assessed for the induction of apoptosis by Annexin/propidium iodide staining using flow cytometry. Individual bars represent daily mean percentages of Annexin V–positive cells (days 0, 1, 2, 3) ± SEM (3 biological replicates). (D) Eµ-Myc lymphoma cells (no. 107) were treated with HDAC1/2-selective RGFP233 (5 µM) and assessed for apoptosis (Annexin/propidium iodide) by flow cytometry at indicated time points. Data are presented as mean ± SEM (3 biological replicates).
A pro-apoptotic effect requires the depletion of Hdac1 and Hdac2 in Eµ-Myc lymphoma. We produced tractable Eµ-Myc lymphoma cells with conditional knockout of Hdac1, Hdac2, or Hdac6 (Eµ-Myc.Hdac1−/−; Eµ-Myc.Hdac2−/−; Eµ-Myc.Hdac6−/−) and transduced these with shRNAs against Hdac1, Hdac2, Hdac3, or Hdac6. (A) Eµ-Myc.Hdac1−/− lymphoma was transduced with constitutive (pLMS) vectors containing shRNA cassettes against Hdac2, Hdac3, or Hdac6, FACS-sorted to approximately 50% GFP+, and 50% GFP−, and cell representation followed over time using competitive proliferation assays, as described previously. Individual bars represent daily flow cytometry assessment of GFP positive cells (days 0, 1, 3, 5, 7, 9, 11, 13) from 3 independent replicates (mean ± SEM) and normalized to day 0. (B) Effects of constitutively depleting Hdac1, Hdac2, or Hdac3 in Eµ-Myc.Hdac6−/− cells using competitive assays. Each bar represents different days of testing: days 0, 1, 3, 5, 7, 9, 11, 13. Data are presented as mean ± SEM of 3 biological replicates. (C) Eµ-Myc.Hdac1−/− cells transduced with shHdac2.439 were FACS-sorted (GFP+) immediately following transduction and assessed for the induction of apoptosis by Annexin/propidium iodide staining using flow cytometry. Individual bars represent daily mean percentages of Annexin V–positive cells (days 0, 1, 2, 3) ± SEM (3 biological replicates). (D) Eµ-Myc lymphoma cells (no. 107) were treated with HDAC1/2-selective RGFP233 (5 µM) and assessed for apoptosis (Annexin/propidium iodide) by flow cytometry at indicated time points. Data are presented as mean ± SEM (3 biological replicates).
We then investigated whether the effects of suppressing Hdac1 with Hdac2 were due to induction of apoptosis and whether this genetic effect could be phenocopied pharmacologically. The most profound biological response to deleting/depleting Hdac1 with Hdac2 in Eµ-Myc lymphoma was apoptosis (Figure 7C; supplemental Figure 8E-F). Moreover, treatment of Eµ-Myc lymphoma with the HDAC1/2 inhibitor RGFP23322 phenocopied the effects observed following genetic deletion/depletion of Hdac1 with Hdac2 (Figure 7D).
To validate the effects of genetically deleting/depleting Hdac1 with Hdac2 in vivo, we transplanted Eµ-Myc.Hdac1−/− cells transduced with dox-inducible vector pTRMPVIR34 expressing shHdac2.439 or shScr into NSG mice (n = 32). Tumor-bearing mice were fed dox (day 3) and assessed for tumor burden by flow cytometry (days 16, 17, and/or 22; supplemental Figure 8G). Strikingly, combined deletion/depletion of both Hdac1 with Hdac2 significantly reduced tumor burden (Venus+/dsRed+ cells) in the PB (days 16 and 22; supplemental Figure 8H), spleen (day 17; supplemental Figure 8I), and bone marrow (day 17; supplemental Figure 8J). The outgrowth of Venus+/dsRed− cells was observed in mice bearing Eµ-Myc.Hdac1−/− lymphoma, suggesting a selection against Hdac2 depletion (supplemental Figure 8K).
Discussion
Broad clinical use of HDACi targeting multiple HDAC isoforms (vorinostat, belinostat, panobinostat, romidepsin) has been hampered by dose limiting toxicities. Here we investigated whether RNA interference (RNAi)-mediated depletion of a single Hdac was capable of reproducing the anti-proliferative or pro-apoptotic effects of broader spectrum HDACi in 3 murine models of hematological neoplasms: 2 AMLs (MLL-AF9;NrasG12D and PML-RARα APL) and Eµ-Myc B-cell lymphoma. Strikingly, we demonstrated that depletion of Hdac3 consistently attenuated tumor cell proliferation in each tumor model tested in the absence of an apoptotic effect. Additionally, combined deletion/depletion of Hdac1 with Hdac2 was sufficient to induce a pro-apoptotic response in Eµ-Myc lymphoma suggesting that HDACis in development should target HDAC3 alone or both HDAC1 and HDAC2.
Using RNAi technology34,37,46 and an unbiased screening approach, we uncovered unique dependencies of established AML, Eµ-Myc lymphoma, and APL to Hdac3 expression in vitro and in vivo. However, in contrast to a recent report that Hdac3 suppression induces apoptosis in multiple myeloma cells,21 we were unable to detect any significant level of apoptosis following Hdac3 depletion in Eµ-Myc lymphoma. Indeed, loss of Hdac3 in Eµ-Myc tumor cells led to an antiproliferative response that was not attenuated by overexpression of pro-survival BCL-2 or pharmacological inhibition of pro-apoptotic caspases. In agreement, the HDAC3-selective compound RGFP966 mediated an antiproliferative response.
Intriguingly, we were unable to detect acute changes to the cell cycle using sensitive, Edu-based DNA labeling assays. This is consistent with reports demonstrating reduced proliferation of human lung fibroblasts following depletion of ribosomal proteins RP5/RP1147 and delayed tumor growth in Eµ-Myc mice hypomorphic for RPL2448 that could not be attributed to cell-cycle arrest but likely is due to reduced ribosomal content and rates of translation.
Cyclin-dependent kinase inhibitor p21Cip1/Waf1 was upregulated following Hdac3 depletion in Eµ-Myc lymphoma, as observed by others.25,49 In contrast to reports that the antitumor effects of HDACi can be mediated in part by p21Cip1/Waf1,50-52 knockout of Cdkn1a (Eµ-Myc.Cdkn1a−/−) did not attenuate the loss of proliferation phenotype observed following Hdac3 depletion. This is in agreement with our previous report that deletion of Cdkn1a in Eµ-Myc lymphoma did not alter sensitivity to vorinostat-induced cell death or prevent cell-cycle arrest.24
Although all 3 leukemias/lymphomas demonstrated significant antiproliferative effects following Hdac3 depletion in vitro and in vivo, Hdac3 depletion in APL cells also triggered differentiation and led to a more mature phenotype. Furthermore, low concentrations of RGFP966 were able to mimic Hdac3 depletion by reducing clonogenicity and upregulating the myeloid differentiation markers Gr.1 and CD11b (MAC1) concomitant with morphological changes reminiscent of differentiated myeloid cells.28 This suggests that low-dose HDAC3-selective inhibition in patients with APL may promote tumor cell differentiation and enable tumor remissions without the need for toxic chemotherapy, similar to that observed with ATRA or arsenic trioxide.27,53,54
Our data also provide clear genetic evidence that isoform-selective inhibitors targeting alternative HDACs, including HDAC6, may not provide potent antitumor effects. Recent reports highlighted HDAC6 as a key target of broad-acting HDACi 55-57 ; however, our data demonstrate that genetic depletion/deletion of Hdac6 had only minor, if any, growth inhibitory effects in AML or Eµ-Myc lymphoma. In agreement, suppression of HDAC6 did not elicit growth inhibition in Burkitt lymphoma,58 lung carcinoma,59 oncogenic Bcr-Abl–addicted myeloid cells,29 or neuroblastoma.60 Therefore, our data do not support the development and use of HDAC6-selective inhibitors in patients with hematological malignancies modeled in our preclinical studies.
Because depletion of Hdac3 alone was unable to induce apoptosis, we investigated whether deletion of 1 Hdac isoform is able to cooperate with depletion of another Hdac to induce a pro-apoptotic response. By genetically deleting Hdac1, Hdac2, or Hdac6 in Eµ-Myc lymphoma and concomitantly knocking down alternative Hdacs, we uncovered a functional interaction between Hdac1 and Hdac2. Importantly, we observed that suppression of both Hdac1 with Hdac2 was pro-apoptotic in Eµ-Myc lymphoma. This is not surprising considering the functional redundancy reported between these 2 Hdac isoforms.61,62 Pharmacological validation using the HDAC1/2-selective inhibitor RGFP23322 confirmed that suppression of HDAC1 with HDAC2 was sufficient to induce a significant pro-apoptotic response in Eµ-Myc lymphoma.
We have previously demonstrated that FDA-approved compounds, such as vorinostat and panobinostat, that inhibit class I and class II HDACs, and romidepsin that more selectively targets HDACs 1, 2, and 3, all induce apoptosis of Eµ-Myc lymphoma and that this biological response is important for the therapeutic effects of these agents in our preclinical models.29,39,63,64 Collectively, these data suggest that apoptosis mediated by concomitant inhibition of HDACs 1 and 2 is the dominant biological effect mediated by these agents; however, additional biological responses such as suppression of cell proliferation through HDAC3 inhibition may be important in situations where the apoptotic response is inactivated.
In conclusion, this is the first study to provide comprehensive genetic and pharmacological analyses of the sensitivities of 3 distinct hematological tumor types to suppression of individual Hdac isoforms in vitro and in vivo. Using advanced genetic techniques and pharmacological inhibitors, we demonstrated that depletion of Hdac3 reduces the proliferation and/or triggers the differentiation of tumor cells in the absence of an apoptotic response. Moreover, although depletion of Hdac3 did not synergize with loss of any other Hdacs, the combined loss of Hdac1 with Hdac2 induced potent pro-apoptotic effects demonstrated both in vitro and in vivo. Taken together, our data suggest that HDAC3-selective inhibitors may be effective for the treatment of hematological malignancies and that newly developed agents should prioritize HDAC1 and HDAC2 as targets to induce effective antitumor responses.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
The authors acknowledge Ross Dickins, Katie McJunkin, Amy Rappaport, Chris Clarke, and Edwin Hawkins for valuable discussion and reagents; Viki Milovac, Sophie Curcio, Ralph Rossi, and Mandy Ludford-Menting for flow cytometry support; and Lauren Dawes and Kat Papastratos for animal husbandry. We acknowledge Oronza A. Botrugno for technical assistance and Giancarlo Pruneri for hematological analysis.
This work was supported by a National Health and Medical Research Council (NHMRC) Biomedical Fellowship and a Peter MacCallum Cancer Foundation Grant (G.M.M.); a Fondazione Italiana per la ricerca sul cancro (FIRC) fellowship (P.M.); National Institutes of Health (grant CA174793), Burroughs-Wellcome Fund Career Award for Medical Scientists, and Alex’s Lemonade Stand Foundation “A” Award (C.R.V.); and NHMRC Program and Project Grants (Senior Principal Research Fellow), Cancer Council Victoria, Leukemia Foundation of Australia, Victorian Cancer Agency, and Australian Rotary Health Foundation (R.W.J.). Laboratory research is funded by an ERC Starting Grant (36860), Special Research Program of the Austrian Science Fund (grant F4710), and Boehringer Ingelheim (J.Z.); the Italian Association for Cancer Research, FIRC, National Research Council Flagship Project Epigen, and European Community (4D Cell Fate Project 277899) (S.M.).
Authorship
Contribution: G.M.M. designed experiments, executed research, analyzed and interpreted all data, and wrote the manuscript; L.A.C., K.J.F., and K.S. aided in experimental work in Eµ-Myc lymphoma; E.V. and K.S. aided in animal experiments; P.M., F.S., and S.M. designed and executed experiments in acute promyelocytic leukemia cells; J.Z. provided critical experimental reagents; C.R.V. and J.Z. designed and analyzed experiments in acute myeloid leukemia (AML) cells. E.W. and M.R. performed experiments in AML and analyzed data; C.M.H. provided mass spectrometry biomarker analysis; R.W.J., S.M., and J.Z. designed experiments, provided discussion, and assisted in writing the manuscript; and J.R.R. provided RGFP966 and RGFP233 for experimental work.
Conflict-of-interest disclosure: James Rusche is an employee of Repligen Corporation. The other authors declare that there are no other conflicts of interest.
Correspondence: Geoffrey M. Matthews, Department of Medical Oncology, Dana-Farber Cancer Institute, Department of Medicine, Harvard Medical School, 450 Brookline Ave, Harvard Institutes of Medicine Building, Boston, MA 02215; e-mail: geoffreym_matthews@dfci.harvard.edu; Ricky W. Johnstone, Cancer Therapeutics and Cancer Immunology Programs, Peter MacCallum Cancer Centre, St Andrews Place, East Melbourne, VIC 3002, Australia; e-mail: ricky.johnstone@petermac.org; and Saverio Minucci, Drug Development Program and Department of Experimental Oncology, European Institute of Oncology, IEO, 20139 Milan, Italy; e-mail: saverio.minucci@ieo.eu.
References
Author notes
G.M.M. and P.M. contributed equally to this study.
J.Z., S.M., and R.W.J. contributed equally to this study.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal