Abstract
Introduction: Genome- and transcriptome-wide analyses continue to enhance our understanding of the molecular pathogenesis of cancer. In lymphomas, this has enabled the identification of hundreds of recurrently mutated genes, highlighting genetic heterogeneity and relationships both within and among clinical entities. While the growing availability of lymphoma genomic data sets can be leveraged to integrate genomic analyses into diagnostic testing and clinical trials, the ability to rapidly process genomic data sets in a reproducible manner serves as a barrier to this goal. To this end, we developed a suite of tools Lymphoid Cancer Research modules (LCR-modules) to facilitate the discovery of novel drivers and molecular features in lymphoma cancers and perform quantitative comparisons between disease entities. We demonstrate here how this toolkit enabled a meta-analysis of lymphoma genomic data involving genome-wide profiles of 3330 patients.
Methods: We assembled a collection of whole genome, whole exome, and RNA sequencing data from a combination of controlled-access repositories and ongoing projects at BC Cancer. The scope of genomic analysis of mature B-cell lymphomas (GAMBL) project includes cell lines and patient tumors from all common mature B cell neoplasms, comprising a total of 4612 samples from 3330 patients. To facilitate the project, we developed a suite of open-source and custom bioinformatics tools (https://github.com/LCR-BCCRC/lcr-modules) that leverages the Snakemake workflow management system and includes lymphoma-centric modules for the discovery and annotation of common mutation types, analysis of B-cell receptor repertoires and discovery of novel aSHM targets and relevant non-coding mutations, and RNA-seq analysis with batch correction and normalization. Individual modules are configured to create an automated, scalable, and reproducible workflow that runs each step as dictated by the availability of new data. The cohort-level integrative analysis and comparisons across entities are handled by our custom R package GAMBLR, which facilitates open-ended data analysis and custom visualizations.
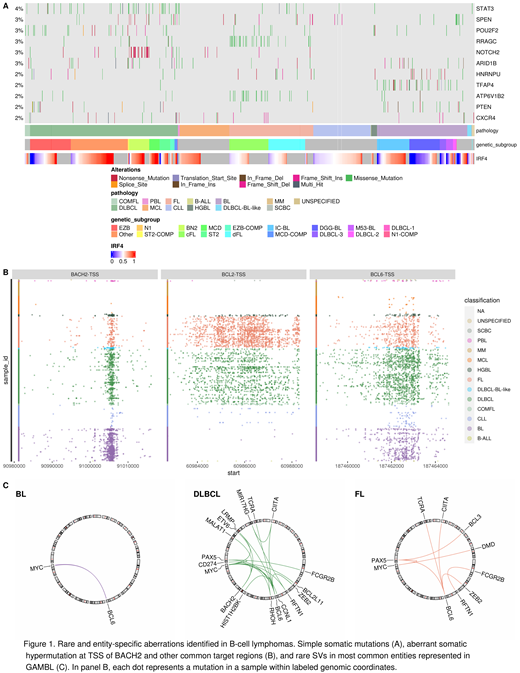
Results: Simple somatic mutations (SSM) were detected using a workflow that utilizes four algorithms to identify high-confidence variants with validated default thresholds for filtering of germline variants and common FFPE-associated artifacts, allowing for processing of samples without matched normal tissue. This automated and reproducible workflow facilitated the discovery of novel genes significantly mutated across lymphomas and broadened our understanding of the scope of aberrant somatic hypermutation (aSHM) and other non-coding mutations. Specifically, HNRNPU, STAT3, TFAP4, RRAGC were found to be mutated at relatively low frequencies, and their presence is a distinct feature of certain lymphomas or novel genetic subgroups within lymphoma types (Figure 1A).
The aSHM analysis and discovery of novel hypermutated regions is handled by a custom tool Rainstorm. As a result, we were able to detect sites preferentially hypermutated in a single entity, such as the transcription start site of BACH2, mutated at lower rates than the other common target sites but significantly more in BL compared to other entities (Figure 1B). Combining aSHM at target sites discovered using our toolkit with other genetic features allowed us to explore and establish novel genetic subgroups within Burkitt lymphoma and follicular lymphoma.
SV analysis can be conducted using Manta, GRIDSS, and JaBbA modules with downstream processing in GAMBLR. In B-cell lymphomas, the most common SVs identified using the automated workflow were targeting MYC, BCL2, and CCND1. Unsurprisingly, the most common translocation partner among B-cell lymphomas was the immunoglobulin heavy chain, but the novel BCL6-FOXP1, CD274-BACH2, BCL6-RHOH translocations in DLBCLs and MYC-BCL6 translocations in BLs were identified, among others (Figure 1C).
Conclusions: We present here the modularized workflow for scalable and automated analysis of genomic and transcriptomic data and demonstrate that it can be successfully deployed across thousands of tumour samples for the discovery of known and novel lymphoma biology. This represents an important advancement in reproducibility that will facilitate clinical translation of genomic discoveries.
Grande: Sage Bionetworks: Current Employment. Coyle: Allakos, Inc.: Consultancy. Steidl: AbbVie: Consultancy; Trillium Therapeutics: Research Funding; Epizyme: Research Funding; Seattle Genetics: Consultancy; Curis Inc.: Consultancy; Bayer: Consultancy; Bristol-Myers Squibb: Research Funding. Scott: Abbvie: Consultancy; NanoString Technologies: Patents & Royalties: Patent describing measuring the proliferation signature in MCL using gene expression profiling.; Celgene: Consultancy; AstraZeneca: Consultancy; Incyte: Consultancy; Janssen: Consultancy, Research Funding; Rich/Genentech: Research Funding; BC Cancer: Patents & Royalties: Patent describing assigning DLBCL COO by gene expression profiling--licensed to NanoString Technologies. Patent describing measuring the proliferation signature in MCL using gene expression profiling. . Morin: Epizyme: Patents & Royalties; Celgene: Consultancy; Foundation for Burkitt Lymphoma Research: Membership on an entity's Board of Directors or advisory committees.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal