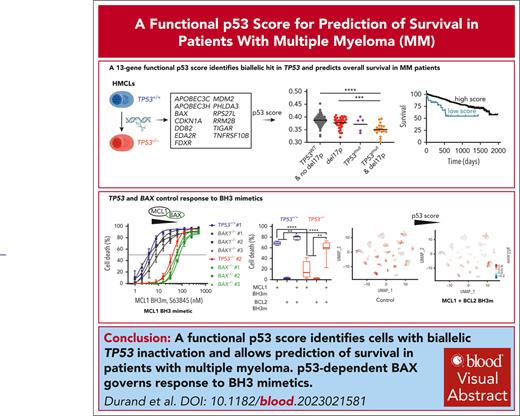
A functional p53 score identifies cells with biallelic TP53 invalidation and predicts survival in myeloma.
p53-regulated BAX, but not BAK, expression governs the death response to BH3 mimetics.
Visual Abstract
To establish a strict p53-dependent gene-expression profile, TP53−/− clones were derived from TP53+/+ and TP53−/mut t(4;14) human myeloma cell lines (HMCLs) using CRISPR/Cas9 technology. From the 17 dysregulated genes shared between the TP53−/− clones from TP53+/+ HMCLs, we established a functional p53 score, involving 13 genes specifically downregulated upon p53 silencing. This functional score segregated clones and myeloma cell lines as well as other cancer cell lines according to their TP53 status. The score efficiently identified samples from patients with myeloma with biallelic TP53 inactivation and was predictive of overall survival in Multiple Myeloma Research Foundation–coMMpass and CASSIOPEA cohorts. At the functional level, we showed that among the 13 genes, p53-regulated BAX expression correlated with and directly affected the MCL1 BH3 mimetic S63845 sensitivity of myeloma cells by decreasing MCL1-BAX complexes. However, resistance to S63845 was overcome by combining MCL1 and BCL2 BH3 mimetics, which displayed synergistic efficacy. The combination of BH3 mimetics was effective in 97% of patient samples with or without del17p. Nevertheless, single-cell RNA sequencing analysis showed that myeloma cells surviving the combination had lower p53 score, showing that myeloma cells with higher p53 score were more sensitive to BH3 mimetics. Taken together, we established a functional p53 score that identifies myeloma cells with biallelic TP53 invalidation, demonstrated that p53-regulated BAX is critical for optimal cell response to BH3 mimetics, and showed that MCL1 and BCL2 BH3 mimetics in combination may be of greater effectiveness for patients with biallelic TP53 invalidation, for whom there is still an unmet medical need.
Introduction
Over the last decade, significant progress has been made in the treatment of patients with multiple myeloma (MM), mainly thanks to the introduction of antibody or cell-based therapies.1-5 Although most patients have an excellent response to treatment, patients with known high-risk features, such as del17p or 1q gain, still have an insufficient response rate.6-8 These chromosomal abnormalities involve at least 2 genes renowned in cancer resistance, TP53 and MCL1, respectively. Loss of p53 function induced by gene deletion and/or mutation is not directly druggable, although it induces targetable neovulnerabilities such as viral permissiveness or loss of cell cycle control.9,10 In contrast, 1q gain induces overexpression of MCL1 that can be targeted with MCL1-specific BH3 mimetics.11,12 The role of p53 loss of function in cancer initiation/development is related to genome guardianship, because the silencing of p53 target genes involved in cell death or cell cycle does not recapitulate p53 loss, contrary to silencing genes involved in DNA repair and integrity.13 In MM, hits in TP53 are not primary events compared with 14q32 translocations, but they greatly impair progression-free survival and overall survival.14,15 This reduced response rate to treatment is instead thought to be related to resistance to tumor death and the loss of cell cycle control. The most frequent hit is monoallelic chromosomal deletion, although biallelic hits are more frequent in very advanced disease such as secondary plasma cell leukemia. Moreover, whole-genome sequencing of ∼400 patients recently identified TP53 biallelic losses in patients with relapsed/refractory disease, emphasizing that complete p53 inactivation plays a major role in resistance to treatment.16 Loss of p53 or mutant-p53 expression might differently impair gene-expression profiling, allowing for de novo expression of genes not directly transactivated by p53.17,18 To assess the direct and specific impact of p53 loss of expression on transcriptome in myeloma cells expressing wild-type p53 protein, we used CRISPR/Cas9 technology to silence p53 in myeloma cell lines having a homogeneous myeloma genomic background: we selected cell lines with t(4;14) translocation, another high-risk hallmark in myeloma.
Materials and methods
HMCLs and patient samples
Samples from patients with MM were obtained from the MYRACLE cohort (NCT03807128).19 Isolation of mononuclear cells and myeloma cells, assessment of chromosomal 1q gain or 17p loss, sequencing of TP53 complementary DNA, and characterization of human myeloma cell lines (HMCLs) were reported previously.12,20-23
Generation of CRISPR/Cas9 clones
NCI-H929, XG7, NAN3, and JIM3 HMCLs were stably transduced with lentivirus expressing mCherry-Cas9 (FUCas9Cherry gifted by Marco Herold, Addgene; plasmid number 70182), before second infection with lentivirus containing the green fluorescent protein gene (GFP) and doxycycline-inducible single-guide RNA (FgH1tUTG gifted from Marco Herold, Addgene; plasmid number 70183). The single-guide RNA sequences were 5′-AGATGGCCATGGCGCGGACG(CGG)-3′ for TP53, 5′-GCCATGCTGGTAGACGTGTA-3′ for BAK1, and 5′-AGTAGAAAAGGGCGACAACC-3′ for BAX.24 GFP+ cells were sorted, single-guide RNA expression was induced with doxycycline and cells were cloned (0.3 cell per well). Control cells and clones were obtained from mCherry+GFP+ cells (containing sgTP53) untreated with doxycycline. To increase the proportion of TP53−/− clones, mCherry+GFP+ NCI-H929 and XG7 cells were additionally treated 48 hours with Nutlin3a (10 μM) before cloning. Three TP53−/−, BAK1−/−, or BAX−/− clones were selected.
Cell death and BH3 profiling assays
RNA-seq and scRNA-seq
Genomic profiling of HMCLs, clones or purified myeloma cells was assessed by RNA sequencing (RNA-seq; 3′–digital gene-expression sequencing [DGE-seq]; GSE245163). For single-cell RNA-seq (scRNA-seq), MM bone marrow mononuclear cells were cultured overnight in RPMI 1640, containing 7% fetal calf serum and 3 ng/mL interleukin-6, with or without a combination of BH3 mimetics before scRNA-seq analysis (10X, Chromium), as described in supplemental Methods, available on the Blood website. Bulk RNA-seq, DGE-seq, or gene expression profiling p53 score was calculated via a rank-based gene set scoring method with the singscore package in R.27 scRNA-seq p53 score was calculated using the AddModuleScore function of Seurat package in R.
Additional materials and methods are detailed in the supplemental Materials and methods.
Results
Silencing TP53 identifies shared p53-regulated genes that provide a functional p53 score
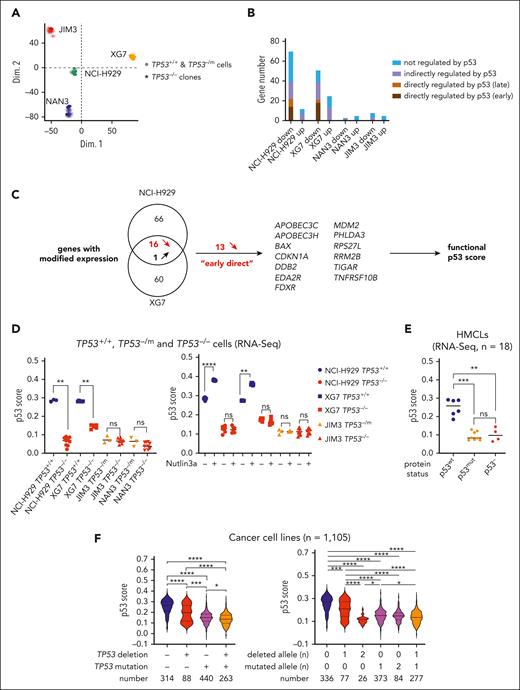
To assess the direct p53 impact in myeloma cells, TP53−/− cells were derived after CRISPR/Cas9 genome editing from t(4;14) HMCL, 2 TP53+/+ NCI-H929 and XG7, and 2 TP53−/mut JIM3 TP53−/R273C and NAN3 TP53−/R248Q. Three TP53−/− clones (TP53−/− numbers 1, 2, and 3) were selected for each cell line, and their expression profiling was compared with that of control cells (mCherry+GFP+TP53+/+ or TP53−/mut bulk cells) and/or control clones (TP53+/+ numbers 1, 2, and 3 or TP53−/mut numbers 1, 2, and 3; Figure 1A; supplemental Table 1). Including TP53, 83 and 77 genes were differentially expressed between TP53+/+ and TP53−/− cells in NCI-H929 and XG7, respectively (false discovery rate < 0.05; supplemental Table 2); 58% and 68% of these genes have been characterized elsewhere as p53-regulated genes,29 that is, early-direct, late p53 targets, with or without proximal p53 binding, or translational, in response to nutlin3a in cell lines28 (Figure 1B). Several genes were differentially expressed between TP53−/mut and TP53−/− cells, but none, except TP53 itself, was shared between the HMCLs. We next focused on the genes whose regulation was shared between the 2 TP53+/+ HMCLs; 16 genes (including TP53) were downregulated in TP53−/− clones, and 1 was upregulated (supplemental Table 2; supplemental Figure 1). We selected 13 genes with decreased expression in TP53−/− clones, known as “early direct” for creating a functional p53 score (Figure 1C).28 Notably, only 5 of these genes were found differentially expressed between 16 TP53wt and 27 TP53abnormal HMCLs using microarray profiling (supplemental Table 2). The 13 genes are located on 12 different chromosome arms, excluding direct impact of myeloma-recurrent chromosomal abnormalities on expression (Table 1). The score discriminated TP53−/− and TP53+/+ clones from NCI-H929 and XG7 but not TP53−/− and TP53−/mut cells from JIM3 and NAN3 (Figure 1D, left). We also characterized the genes differentially expressed between TP53+/+ and TP53−/− clones after nutlin3a treatment; excluding genes with nutlin3a-induced modification of expression in TP53−/− clones (ie, MDM2-dependent and p53-independent regulation), 825 genes (397 up, 428 down in TP53+/+ vs TP53−/− cells) were commonly regulated in NCI-H929 and XG7, including the 16 downregulated genes (supplemental Figure 1; supplemental Table 3). As expected, nutlin3a increased p53 score only in TP53+/+ clones (Figure 1D, right). The score did not discriminate 8 HMCLs expressing a mutated p53 protein from 4 expressing no p53 protein, underscoring that it identifies p53 loss of function, whatever its origin (Figure 1E). In HMCLs profiled using microarray, the score segregated 16 TP53wt from 27 TP53abnormal HMCLs (P < .0001), whatever their myeloma-specific 14q32 translocation background (supplemental Figure 2A-B; supplemental Table 4). Moreover, in 1105 cancer cell lines, the score was significantly associated with TP53 deletion, mutation, or both (Figure 1F, left), with cell lines with biallelic hits having the lowest score (Figure 1F, right; supplemental Table 5), whatever the cell ontology and the patient’s sex (supplemental Figure 2C-D; supplemental Table 5).
A 13-gene functional p53 score established from isogenic TP53 CRISPR/Cas9 HMCLs. (A) Principal component analysis of TP53+/+ or TP53−/mut and TP53−/− cells derived from NCI-H929, XG7, JIM3, and NAN3 HMCLs. Principal component analysis was performed on all cells profiled in triplicate wells using RNA-seq (DGE-seq). RNA profiling was performed on TP53+/+ control (bulk) cells and TP53−/− clones. (B) Number of genes significantly differentially expressed between TP53+/+ control cells and TP53−/− clones. The graph represents the number of genes significantly downregulated or upregulated in TP53−/− NCI-H929, XG7, JIM3, and NAN3 clones compared to their respective TP53+/+ or TP53−/mut control cells (false discovery rate < 0.05). Genes were classified according to their unknown or known p53 transactivation: early-direct, late-direct, late-indirect and not regulated by p53.28 Expression profiling was performed by DGE-seq in triplicate wells (see supplemental Table 2 for complete gene listing). (C) Schematic representation of the functional p53 score construction. Sixteen downregulated genes in TP53−/− clones were shared between NCI-H929 and XG7, and 13 known as early-direct p53 regulated genes were selected for establishing the p53 functional score. (D) The p53 score segregates the clones according to their TP53 status. The score was calculated in TP53+/+ or TP53−/mut and TP53−/− NCI-H929, XG7, JIM3, and NAN3 cells under constitutive (left) or 24-hour nutlin3a (2 μM for XG7 cells and 10 μM for all other clones) culture (right). Statistical analyses were performed using the Mann-Whitney U test. Blue, red, and orange represent TP53+/+, TP53−/−, and TP53−/mutated cells, respectively. (E) The p53 score segregates HMCLs according to their p53 status. The score was calculated in 18 HMCLs characterized by DGE-seq. TP53 sequencing was performed on complementary DNA and p53 expression was determined by western blotting (supplemental Table 4). Statistical analyses were performed using the Kruskal-Wallis test with multiple comparisons. (F) The p53 score is functional in 1105 cancer cell lines with different TP53 status. The score was calculated in 1105 cancer cell lines from DepMap and analyzed according to the presence of TP53 deletion and/or mutation (left) and to the number of TP53 deletion or mutation (right). Statistical analyses were performed using the Kruskal-Wallis test with multiple comparisons. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05. ns, not significant.
A 13-gene functional p53 score established from isogenic TP53 CRISPR/Cas9 HMCLs. (A) Principal component analysis of TP53+/+ or TP53−/mut and TP53−/− cells derived from NCI-H929, XG7, JIM3, and NAN3 HMCLs. Principal component analysis was performed on all cells profiled in triplicate wells using RNA-seq (DGE-seq). RNA profiling was performed on TP53+/+ control (bulk) cells and TP53−/− clones. (B) Number of genes significantly differentially expressed between TP53+/+ control cells and TP53−/− clones. The graph represents the number of genes significantly downregulated or upregulated in TP53−/− NCI-H929, XG7, JIM3, and NAN3 clones compared to their respective TP53+/+ or TP53−/mut control cells (false discovery rate < 0.05). Genes were classified according to their unknown or known p53 transactivation: early-direct, late-direct, late-indirect and not regulated by p53.28 Expression profiling was performed by DGE-seq in triplicate wells (see supplemental Table 2 for complete gene listing). (C) Schematic representation of the functional p53 score construction. Sixteen downregulated genes in TP53−/− clones were shared between NCI-H929 and XG7, and 13 known as early-direct p53 regulated genes were selected for establishing the p53 functional score. (D) The p53 score segregates the clones according to their TP53 status. The score was calculated in TP53+/+ or TP53−/mut and TP53−/− NCI-H929, XG7, JIM3, and NAN3 cells under constitutive (left) or 24-hour nutlin3a (2 μM for XG7 cells and 10 μM for all other clones) culture (right). Statistical analyses were performed using the Mann-Whitney U test. Blue, red, and orange represent TP53+/+, TP53−/−, and TP53−/mutated cells, respectively. (E) The p53 score segregates HMCLs according to their p53 status. The score was calculated in 18 HMCLs characterized by DGE-seq. TP53 sequencing was performed on complementary DNA and p53 expression was determined by western blotting (supplemental Table 4). Statistical analyses were performed using the Kruskal-Wallis test with multiple comparisons. (F) The p53 score is functional in 1105 cancer cell lines with different TP53 status. The score was calculated in 1105 cancer cell lines from DepMap and analyzed according to the presence of TP53 deletion and/or mutation (left) and to the number of TP53 deletion or mutation (right). Statistical analyses were performed using the Kruskal-Wallis test with multiple comparisons. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05. ns, not significant.
Genes of the p53 score
| Gene symbol . | Gene name . | Cytogenetic location . |
|---|---|---|
| APOBEC3C | Apolipoprotein B mRNA–editing enzyme catalytic subunit 3C | 22q13.1 |
| APOBEC3H | Apolipoprotein B mRNA–editing enzyme catalytic subunit 3H | 22q13.1 |
| BAX | BCL2-associated X | 19q13.33 |
| CDKN1A | Cyclin-dependent kinase inhibitor 1A | 6p21.2 |
| DDB2 | Damage-specific DNA-binding protein 2 | 11p11.2 |
| EDA2R | Ectodysplasin A2 receptor | Xq12 |
| FDXR | Ferredoxin reductase | 17q25.1 |
| MDM2 | MDM2 proto-oncogene | 12q15 |
| PHLDA3 | Pleckstrin homology–like domain family A member 3 | 1q32.1 |
| RPS27L | Ribosomal protein S27 like | 15q22.2 |
| RRM2B | Ribonucleotide reductase regulatory TP53–inducible subunit M2B | 8q22.3 |
| TIGAR | TP53-induced glycolysis regulatory phosphatase | 12p13.32 |
| TNFRSF10B | TNF receptor superfamily member 10b | 8p21.3 |
| Gene symbol . | Gene name . | Cytogenetic location . |
|---|---|---|
| APOBEC3C | Apolipoprotein B mRNA–editing enzyme catalytic subunit 3C | 22q13.1 |
| APOBEC3H | Apolipoprotein B mRNA–editing enzyme catalytic subunit 3H | 22q13.1 |
| BAX | BCL2-associated X | 19q13.33 |
| CDKN1A | Cyclin-dependent kinase inhibitor 1A | 6p21.2 |
| DDB2 | Damage-specific DNA-binding protein 2 | 11p11.2 |
| EDA2R | Ectodysplasin A2 receptor | Xq12 |
| FDXR | Ferredoxin reductase | 17q25.1 |
| MDM2 | MDM2 proto-oncogene | 12q15 |
| PHLDA3 | Pleckstrin homology–like domain family A member 3 | 1q32.1 |
| RPS27L | Ribosomal protein S27 like | 15q22.2 |
| RRM2B | Ribonucleotide reductase regulatory TP53–inducible subunit M2B | 8q22.3 |
| TIGAR | TP53-induced glycolysis regulatory phosphatase | 12p13.32 |
| TNFRSF10B | TNF receptor superfamily member 10b | 8p21.3 |
The table indicates the name and chromosomal location of the 13 genes involved in the score.
mRNA, messenger RNA; TNF, tumor necrosis factor.
The 13-gene functional p53 score is prognostic for myeloma patient survival
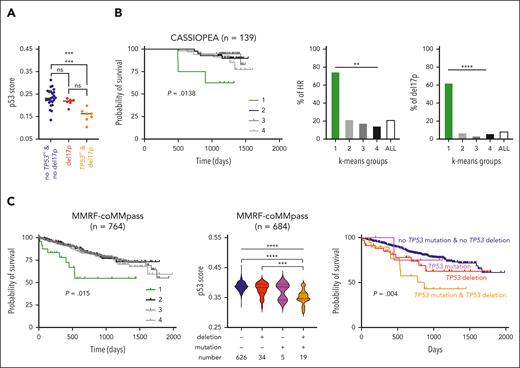
In 38 samples from MYRACLE patients characterized for both del17p (fluorescence in situ hybridization [FISH]) and TP53 mutation (Sanger sequencing), the p53 score significantly discriminated samples with del17p and TP53 mutation (n = 7) from samples without TP53 hit (n = 25; P = .0009; Figure 2A; supplemental Table 6). Because TP53 hits are known to impair overall survival,7 the p53 score was calculated in 2 cohorts of patients at diagnosis, with RNA-seq expression at baseline and overall survival with 139 patients from the CASSIOPEA trial and 764 from the Multiple Myeloma Research Foundation (MMRF)-CoMMpass cohort.30 In both cohorts, K-means method segregated patient samples into 4 groups (supplemental Figure 3A-B, left). Overall survival analysis showed in both cohorts that group 1 with the lowest values (n = 8 and 24 for CASSIOPEA and MMRF-CoMMpass, respectively) had significantly reduced overall survival (P = .0138 and P = .015; Figure 2B-C, left). In CASSIOPEA cohort, group 1 contained 75% of patients classified as high risk (6 of 8) and 62.5% of patients with del17p (5 of 8; Figure 2B, middle and right): patients at high risk (n = 30) or with del17p (n = 12) had significantly reduced overall survival (supplemental Figure 3A).
The 13-gene p53 score is predictive of overall survival. (A) The score discriminates patient samples according to TP53 status. The score was calculated in 38 patient samples characterized by DGE-seq. Del17p and TP53 sequencing were determined by FISH and Sanger sequencing of complementary DNA products, respectively (supplemental Table 6).25 Statistical analysis was performed using the Kruskal-Wallis test with multiple comparisons. (B) The p53 score is predictive of overall survival (OS) of 139 CASSIOPEA patients. Groups of patients were determined according to the score using the K-means method (supplemental Figure 3A). Left graph represents the OS of the 4 groups. Middle and right graphs represent the frequency of patients classified as high risk or with del17p in each K-means groups. Statistical significance was determined using the log-rank (Mantel-Cox) test or χ2 test. (C) OS of 764 MMRF-coMMpass patients according to p53 score. Groups were determined using the K-means method (supplemental Figure 3B). Left graph represents the OS of the 4 groups. Middle graph represents the p53 score in groups separated according to the presence of TP53 deletion or/and mutation in 684 patients, and the right graph represents their OS. Variant allele frequency thresholds were set up at < −0.5 for deletion and >0.35 for mutation. Statistical significance was determined using the log-rank (Mantel-Cox) test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
The 13-gene p53 score is predictive of overall survival. (A) The score discriminates patient samples according to TP53 status. The score was calculated in 38 patient samples characterized by DGE-seq. Del17p and TP53 sequencing were determined by FISH and Sanger sequencing of complementary DNA products, respectively (supplemental Table 6).25 Statistical analysis was performed using the Kruskal-Wallis test with multiple comparisons. (B) The p53 score is predictive of overall survival (OS) of 139 CASSIOPEA patients. Groups of patients were determined according to the score using the K-means method (supplemental Figure 3A). Left graph represents the OS of the 4 groups. Middle and right graphs represent the frequency of patients classified as high risk or with del17p in each K-means groups. Statistical significance was determined using the log-rank (Mantel-Cox) test or χ2 test. (C) OS of 764 MMRF-coMMpass patients according to p53 score. Groups were determined using the K-means method (supplemental Figure 3B). Left graph represents the OS of the 4 groups. Middle graph represents the p53 score in groups separated according to the presence of TP53 deletion or/and mutation in 684 patients, and the right graph represents their OS. Variant allele frequency thresholds were set up at < −0.5 for deletion and >0.35 for mutation. Statistical significance was determined using the log-rank (Mantel-Cox) test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
In the MMRF-CoMMpass cohort, TP53 mutation and 17p deletion were available for 684 patients; 19 displayed deletion and mutation, 34 deletion, and 5 mutation (deletion or mutation thresholds were loss of heterozygosity < −0.5 or variant allele frequency > 0.35). Samples with del17p and mutation had the lowest score (P < .0001) and a lower score than samples with only del17p (P = .0002; Figure 2C, middle). Notably, 90% of samples with biallelic hit were in groups 1 (53%) and 2 (37%), whereas samples with del17p only were more equally distributed among the 4 groups (P = .003; supplemental Figure 3B, middle). Samples with TP53 hit(s) constituted 74% of group 1 and only 5% to 10% of the other groups (supplemental Figure 3B, right). Patients with TP53 mutation and del17p, and not with del17p only, had lower overall survival; the median was 870 days vs not reached in the other groups (Figure 2C, right; P = .004).
These results collectively showed that this score identified samples with biallelic hit in TP53 and provided prognosis in 2 cohorts. We further compared this score with another p53 functional score used for the clinical interpretation of germ line TP53 variants in Li-Fraumeni syndrome, which assesses the increased expression of 10 p53 target genes (none shared with our 13-gene score; supplemental Figure 4) in doxorubicin-treated peripheral blood mononuclear cells.31 This score did not distinguish XG7 or NCI-H929 TP53−/− clones from TP53+/+ clones in the absence of nutlin3a (supplemental Figure 5A-B). It also did not segregate cell lines or patient samples according to their TP53 status (supplemental Figure 5C-D).
TP53 silencing impairs mitochondrial MCL1 priming
As expected, p53-silencing in TP53+/+ cells, but not in TP53−/mut cells, increased resistance to melphalan (lethal dose 50% [LD50] values increased from 11.3 to 16.6 μM, P = .0011 and from 18.7 to 28.3 μM; P = .004 in NCI-H929 and XG7, respectively), and to nutlin3a (LD50 values were ∼2 μM in TP53+/+ clones, whereas no cell death was induced at 10 μM in TP53−/− clones; supplemental Table 7).32,33 The 13 genes involved in the p53 score are known to be involved in the cell cycle, DNA duplication and repair (CDKN1A, DDB2, and RRM2B), defense against viruses or reactive oxygen species (APOBEC3C, APOBEC3H, RPS27L, and TIGAR), apoptosis and mitochondrial metabolism (BAX, TNFRSF10B, and FDXR), signaling pathways (PHLDA3 and EDA2R), and in p53 regulation (MDM2). Given the major role of BAX in apoptosis, we focused on the impact of its expression on mitochondrial fitness. We confirmed that BAX expression was decreased in TP53−/− clones derived from TP53+/+ HMCLs and unchanged in those derived from TP53−/mut HMCLs (supplemental Figure 6A). As previously reported, TP53−/mut HMCLs also underexpressed BAX when compared with TP53wt HMCLs (supplemental Figure 6B).22,25,32 Noteworthy, BAX was the only BCL2-family protein displaying modified expression in both NCI-H929 and XG7 TP53−/− clones. To assess the mitochondrial impact of p53 loss, we used BH3 mimetics specific to MCL1 (S63845), BCL2 (venetoclax) and BCLXL (A1155463), and performed BH3 profiling.12,34
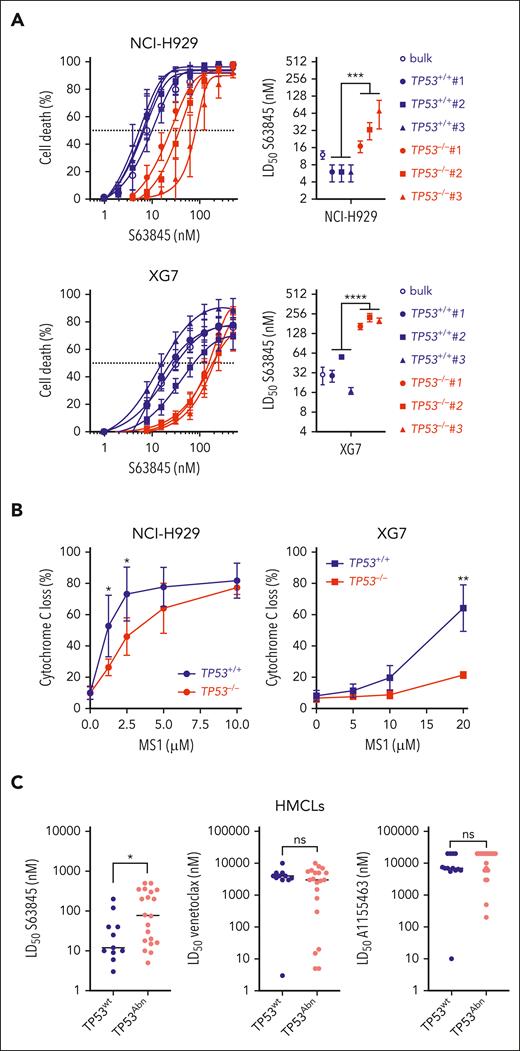
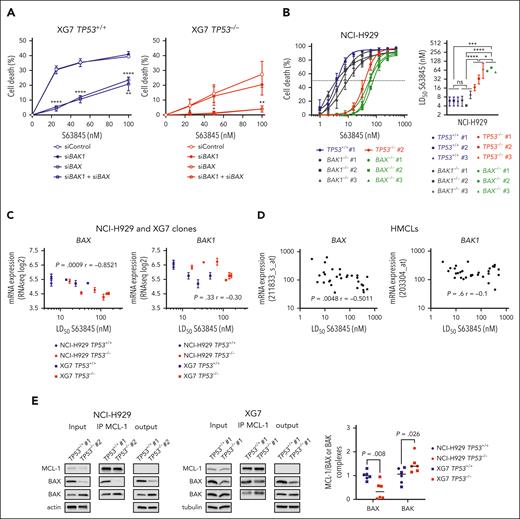
Silencing p53wt in NCI-H929 and XG7 (but not p53mut in JIM3 and NAN3) induced resistance to S63845: LD50 mean values in clones increased from 6 to 40 nM (P = .0002) and from 29 to 138nM, (P < .0001), in NCI-H929 and XG7, respectively (Figure 3A; supplemental Table 7). Impact of p53 on response to venetoclax or A1155463 could not be assessed, cell lines being resistant to both. Global mitochondrial priming, that is, response to BIM or BMF peptides, was unchanged for both NCI-H929 and XG7 TP53−/− clones (supplemental Figure 7), whereas cytochrome-C release in response to MS1 peptide specific to MCL1 was significantly decreased in both NCI-H929 and XG7 TP53−/− clones (Figure 3B), in keeping with reduced sensitivity to S63845. In 31 HMCLs, S63845, but not venetoclax or A1155463, LD50 values were also significantly lower in 11 TP53wt than in 20 TP53Abn HMCLs (median was 12.5 nM vs 77.50 nM; P = .026; Figure 3C; supplemental Table 8). These data collectively showed that TP53 status strongly affected priming to MCL1 in HMCLs.
TP53 loss impairs response to MCL1 BH3 mimetic S63845. (A) CRISPR/Cas9-mediated TP53 inactivation increased resistance to BH3 mimetic specific to MCL1. (Left) BH3 mimetic specific to MCL1 dose response in isogenic TP53+/+ and TP53−/− NCI-H929 and XG7 clones. Graphs represent 3 independent experiments. (Right) LD50 S63845 values in TP53+/+ control cells (bulk) and TP53+/+ and TP53−/− clones in NCI-H929 (top) and XG7 (bottom); see supplemental Table 7. Statistical analysis was performed using the Mann-Whitney U test. (B) CRISPR/Cas9-mediated TP53 inactivation decreased priming to MCL1. Priming to MCL1 was assessed by determining cytochrome-C release induced by MS1 peptide. Cytochrome-C staining was performed using anti–cytochrome-C monoclonal antibody and analyzed by flow cytometry. Graphs represent the mean ± standard deviation (SD) of 3 to 4 independent experiments. Statistical analysis was performed using the Wilcoxon matched pairs signed-rank test. (C) S63845 LD50 values were significantly lower in TP53wt HMCLs. S63845, venetoclax, and A1155463 LD50 values were analyzed in 11 TP53wt and 20 TP53Abn HMCLs (supplemental Table 8). Each point represents 1 HMCL; median values are indicated. For A1155463 and venetoclax, LD50 was arbitrarily set up at 20 000 and 10 000 nM when it did not reach 10 000 or 5000 nM, respectively. Statistical analyses were performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
TP53 loss impairs response to MCL1 BH3 mimetic S63845. (A) CRISPR/Cas9-mediated TP53 inactivation increased resistance to BH3 mimetic specific to MCL1. (Left) BH3 mimetic specific to MCL1 dose response in isogenic TP53+/+ and TP53−/− NCI-H929 and XG7 clones. Graphs represent 3 independent experiments. (Right) LD50 S63845 values in TP53+/+ control cells (bulk) and TP53+/+ and TP53−/− clones in NCI-H929 (top) and XG7 (bottom); see supplemental Table 7. Statistical analysis was performed using the Mann-Whitney U test. (B) CRISPR/Cas9-mediated TP53 inactivation decreased priming to MCL1. Priming to MCL1 was assessed by determining cytochrome-C release induced by MS1 peptide. Cytochrome-C staining was performed using anti–cytochrome-C monoclonal antibody and analyzed by flow cytometry. Graphs represent the mean ± standard deviation (SD) of 3 to 4 independent experiments. Statistical analysis was performed using the Wilcoxon matched pairs signed-rank test. (C) S63845 LD50 values were significantly lower in TP53wt HMCLs. S63845, venetoclax, and A1155463 LD50 values were analyzed in 11 TP53wt and 20 TP53Abn HMCLs (supplemental Table 8). Each point represents 1 HMCL; median values are indicated. For A1155463 and venetoclax, LD50 was arbitrarily set up at 20 000 and 10 000 nM when it did not reach 10 000 or 5000 nM, respectively. Statistical analyses were performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
BAX expression controls response to MCL1-specific BH3 mimetic
To determine the respective role of BAX and BAK in the response to S63845, we transiently silenced BAK1, BAX, or both in TP53+/+ and TP53−/− XG7 cells (supplemental Figure 8A). By contrast to BAK1 silencing, BAX silencing (with or without BAK1 silencing) strongly inhibited cell death: death inhibition was 87% ± 5% (P < .0001) and 91% ± 4% (P = .0385) in TP53+/+ and TP53−/− cells treated with 25 nM S63845, respectively (Figure 4A). CRISPR/Cas9-mediated inactivation of BAX or BAK1 in NCI-H929 cells confirmed that BAX, but not BAK, controlled response to low doses of S63845 (Figure 4B; supplemental Figure 8B; supplemental Table 9): compared with control clones, BAX inactivation induced 10-fold increase in LD50 (mean values, 64 vs 6 nM; P < .0001), whereas BAK1 inactivation did not induce significant LD50 changes (6.7 vs 6 nM). Of note, BAX−/− clones appeared slightly more resistant to S63845 than TP53−/− clones (mean values, 64 vs 40 nM; P = .015), the latter having heterogeneous LD50 values and BAX expression (Figure 4B; supplemental Figure 6). Notably, BAX expression inversely correlated with S63845 LD50 values in TP53+/+ and TP53−/− clones (r = −0.8521; P = .0009, Spearman test), whereas BAK1 expression demonstrated no significant correlation (Figure 4C). In 30 HMCLs, S63845 LD50 values also displayed an inverse correlation with BAX expression (r = −0.5011; P = .0048) but not with BAK1 (P = .6; Figure 4D). No other BCL2 family member displayed significant correlation with S63845 LD50 values in both clones and HMCLs (supplemental Figure 9).
BAX expression governs the response to MCL1 BH3-mimetic S63845. (A) BAX silencing inhibited S63845-induced cell death. XG7 TP53+/+ no. 1 and TP53−/− no. 1 were transiently transfected with siCont, siBAX, siBAK1, or both, before treatment with increasing doses of S63845. The graph represents the mean ± SD of 3 independent experiments. Statistical analyses were performed using the unpaired t test. (B) CRISPR/Cas9-mediated BAX, but not BAK1, inactivation inhibited response to S63845 in NCI-H929 cells. (Left) TP53+/+ no. 1, TP53−/− no. 2, BAX−/− (n = 3), and BAK1−/− (n = 3) clones were treated with increasing doses of S63845 for 24 hours and cell death was determined using flow cytometry. The graph represents the mean ± SD of 3 experiments. (Right) The graph represents the LD50 values of each clone (mean ± SD). Statistical analysis was performed using the Mann-Whitney U test. (C) BAX, but not BAK1, expression correlated with sensitivity to S63845 in NCI-H929 and XG7 clones. Expression of BAX and BAK1 (mean DGE-seq values) were plotted against LD50 S63845 values. Correlation was assessed using the Spearman test. (D) BAX, but not BAK1, expression correlated with sensitivity to S63845 in HMCLs. Expression of BAX and BAK1 (microarray) in 30 HMCLs was plotted against LD50 S63845 value. Correlation was assessed using the Spearman test. (E) CRISPR/Cas9-mediated TP53 inactivation increased MCL1/BAK and decreased MCL1/BAX complexes. For immunoprecipitation (IP) assays, cells were lysed in 1% digitonin-containing buffer and lysates were precleared with protein A conjugated to sepharose beads. For MCL1 IP assay, 700 μg of protein lysate was incubated overnight with an agarose-conjugated monoclonal antibody anti-MCL1 from Santa Cruz Biotechnology. After immunoblotting, MCL1, BAK and BAX levels were determined in the immunoprecipitated fraction and BAX/MCL1 and BAK/MCL1 ratios were calculated to evaluate the abundance of both complexes. One experiment out of 3 is represented. The graph represents the quantities of BAX and BAK bound to MCL1 in TP53+/+ and TP53−/− NCI-H929 and XG7 cells (3 independent experiments). Statistical analyses were performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
BAX expression governs the response to MCL1 BH3-mimetic S63845. (A) BAX silencing inhibited S63845-induced cell death. XG7 TP53+/+ no. 1 and TP53−/− no. 1 were transiently transfected with siCont, siBAX, siBAK1, or both, before treatment with increasing doses of S63845. The graph represents the mean ± SD of 3 independent experiments. Statistical analyses were performed using the unpaired t test. (B) CRISPR/Cas9-mediated BAX, but not BAK1, inactivation inhibited response to S63845 in NCI-H929 cells. (Left) TP53+/+ no. 1, TP53−/− no. 2, BAX−/− (n = 3), and BAK1−/− (n = 3) clones were treated with increasing doses of S63845 for 24 hours and cell death was determined using flow cytometry. The graph represents the mean ± SD of 3 experiments. (Right) The graph represents the LD50 values of each clone (mean ± SD). Statistical analysis was performed using the Mann-Whitney U test. (C) BAX, but not BAK1, expression correlated with sensitivity to S63845 in NCI-H929 and XG7 clones. Expression of BAX and BAK1 (mean DGE-seq values) were plotted against LD50 S63845 values. Correlation was assessed using the Spearman test. (D) BAX, but not BAK1, expression correlated with sensitivity to S63845 in HMCLs. Expression of BAX and BAK1 (microarray) in 30 HMCLs was plotted against LD50 S63845 value. Correlation was assessed using the Spearman test. (E) CRISPR/Cas9-mediated TP53 inactivation increased MCL1/BAK and decreased MCL1/BAX complexes. For immunoprecipitation (IP) assays, cells were lysed in 1% digitonin-containing buffer and lysates were precleared with protein A conjugated to sepharose beads. For MCL1 IP assay, 700 μg of protein lysate was incubated overnight with an agarose-conjugated monoclonal antibody anti-MCL1 from Santa Cruz Biotechnology. After immunoblotting, MCL1, BAK and BAX levels were determined in the immunoprecipitated fraction and BAX/MCL1 and BAK/MCL1 ratios were calculated to evaluate the abundance of both complexes. One experiment out of 3 is represented. The graph represents the quantities of BAX and BAK bound to MCL1 in TP53+/+ and TP53−/− NCI-H929 and XG7 cells (3 independent experiments). Statistical analyses were performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
These results prompted us to directly assess the amount of MCL1 complexed with BAX and BAK. MCL1 immunoprecipitation showed that TP53−/− clones displayed a strong decreased amount of MCL1/BAX complexes (median reduction, 3.07; P = .008) and slightly increased amount of MCL1/BAK (median increase, 1.32; P = .026) when compared with TP53+/+ clones (Figure 4E). S63845 induced the disruption of MCL1-BAX and MCL1-BAK complexes, and BAX-MCL1 complexes were disrupted at lower S63845 concentrations than BAK-MCL1 complexes (supplemental Figure 8C). These data showed that p53 loss reduced the amount of MCL1-BAX complexes.
TP53 or BAX silencing impaired response to BH3 mimetics combination
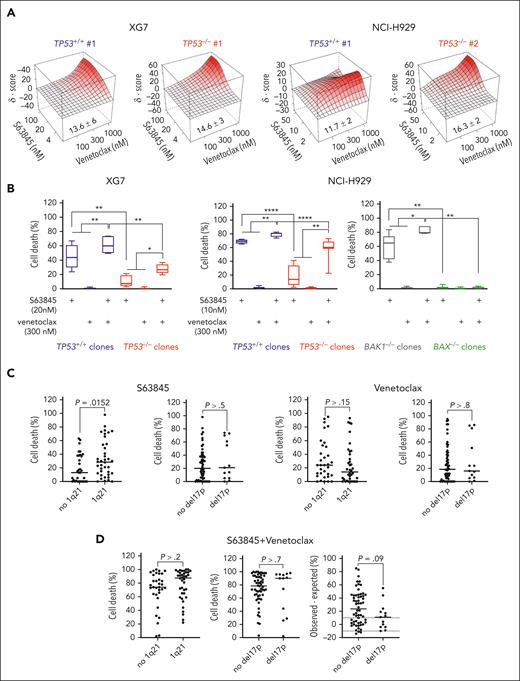
Because the combination of the BH3 mimetics S63845 with venetoclax was reported to be synergistic in myeloma cells, we wondered how p53 loss would affect its efficacy.34 Combination was, indeed, synergistic at high doses in TP53+/+ and TP53−/− clones; Bliss scores were 13.6 ± 6 and 14.6 ± 3 in TP53+/+ no. 1 and TP53−/− no. 1 XG7 clones and 11.7 ± 2 and 16.3 ± 2 in TP53+/+ no. 1 and TP53−/− no. 2 NCI-H929 clones, respectively (Figure 5A). However, at the same doses, maximum cell death induced by combination was significantly lower in TP53−/− clones vs TP53+/+ clones (mean values, 28% ± 6.5% vs 61% ± 11% in XG7 cells [P < .01], and 59% ± 15% vs 79% ± 3.2% in NCI-H929 [P < .0001]; Figure 5B). Of note, combination was more efficient in NCI-H929 cells than in XG7 cells, which had low BCL2 expression (supplemental Figure 6). Finally, BAX but not BAK1 invalidation in NCI-H929 cells, inhibited the response to combination, confirming the major role of BAX in BH3 mimetics response (Figure 5B).
TP53 or BAX silencing impaired response to BH3 mimetics combination. (A) Combination of S63845 and venetoclax was synergistic in TP53+/+ and TP53−/− NCI-H929 and XG7 clones. Clones were cultured for 24 hours with increasing concentrations of S63845 and venetoclax as indicated in the figure. Bliss scores were calculated using SynergyFinder (https://synergyfinder.fimm.fi/). The figure represents the mean of 3 independent experiments. (B) TP53 and BAX controlled the response to BH3 mimetics combination. TP53+/+ (n = 3) and TP53−/− (n = 3) XG7 and NCI-H929 clones as well as BAX−/− (n = 3) and BAK1−/− (n = 3) NCI-H929 clones were cultured for 24 hours with 20 nM (XG7) or 10 nM (NCI-H929) S63845 and/or 300 nM venetoclax, and cell death was assessed using flow cytometry. Statistical analyses were performed using the Mann-Whitney U or Wilcoxon matched pairs signed-rank tests. (C) Gain of 1q21, but not del17p, affected the response to S63845. Response to 25 nM S63845 or 300 nM venetoclax was assessed in 71 myeloma samples, characterized by FISH for both 1q21 gain and del17p (Table 2). The graphs represent cell death induced by each BH3 mimetic in function of 1q21 gain or del17p. Statistical analyses were performed using the Mann-Whitney U test. (D) Deletion of 17p had a trend to decrease the synergy between S63845 and venetoclax. The graphs represent cell death induced by 25 nM S63845 combined with 300 nM venetoclax depending on 1q gain or del17p (left and middle), and the difference between observed cell death (25 nM S63845 combined with 300 nM venetoclax) minus expected cell death (sum of cell death induced by 25 nM S63845 and by 300 nM venetoclax), depending on del17p. Cell death was assessed after 24 hours using flow cytometry. Statistical analyses were performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
TP53 or BAX silencing impaired response to BH3 mimetics combination. (A) Combination of S63845 and venetoclax was synergistic in TP53+/+ and TP53−/− NCI-H929 and XG7 clones. Clones were cultured for 24 hours with increasing concentrations of S63845 and venetoclax as indicated in the figure. Bliss scores were calculated using SynergyFinder (https://synergyfinder.fimm.fi/). The figure represents the mean of 3 independent experiments. (B) TP53 and BAX controlled the response to BH3 mimetics combination. TP53+/+ (n = 3) and TP53−/− (n = 3) XG7 and NCI-H929 clones as well as BAX−/− (n = 3) and BAK1−/− (n = 3) NCI-H929 clones were cultured for 24 hours with 20 nM (XG7) or 10 nM (NCI-H929) S63845 and/or 300 nM venetoclax, and cell death was assessed using flow cytometry. Statistical analyses were performed using the Mann-Whitney U or Wilcoxon matched pairs signed-rank tests. (C) Gain of 1q21, but not del17p, affected the response to S63845. Response to 25 nM S63845 or 300 nM venetoclax was assessed in 71 myeloma samples, characterized by FISH for both 1q21 gain and del17p (Table 2). The graphs represent cell death induced by each BH3 mimetic in function of 1q21 gain or del17p. Statistical analyses were performed using the Mann-Whitney U test. (D) Deletion of 17p had a trend to decrease the synergy between S63845 and venetoclax. The graphs represent cell death induced by 25 nM S63845 combined with 300 nM venetoclax depending on 1q gain or del17p (left and middle), and the difference between observed cell death (25 nM S63845 combined with 300 nM venetoclax) minus expected cell death (sum of cell death induced by 25 nM S63845 and by 300 nM venetoclax), depending on del17p. Cell death was assessed after 24 hours using flow cytometry. Statistical analyses were performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
We next assessed death response to S63845, venetoclax, and their combination in myeloma cells from 71 consecutive patients at diagnosis, progression, or relapse. Median values of cell death induced by 25 nM S63845, 300 nM venetoclax, or their combination were 21%, 18%, and 81%, respectively (Table 2). Presence of 1q gain positively affected the response to S63845 (median cell death was 13% in 33 no 1q samples vs 28.5% in 38 1q samples; P = .0152), as previously reported, but did not affect the response to venetoclax or to the combination, and the presence of del17p did not affect responses to BH3 mimetics (Figure 5C-D).11,35 Regardless of the presence of 1q21 or del17p, combination was effective in 97% of the samples; that is, additive (−10% < observed minus expected death < 10%) in 10 samples (14%) and supra-additive to synergistic (observed minus expected death >10%) in 59 samples (83%). Supra-additivity to synergy of the combination was mainly found in samples without t(11;14) because t(11;14) samples have a high response rate to venetoclax alone (supplemental Figure 10A-B). Combination had a trend to be less efficient in the del17p samples when compared with the no del17p samples: median of observed minus expected value was 11% (range, −10 to 55) vs 23.5% (range, −14 to 85), respectively (P = .09; Figure 5D). Of note, the del17p impact on BH3 combination efficacy was rather restricted to samples without t(11;14); the median of observed minus expected value was 13% (range, −10 to 55) in 10 samples del17p vs 34% (range, −12 to 85) in 32 samples with no del17p (P=.0457; supplemental Figure 10A).
Cell death in MYRACLE samples induced by BH3 mimetics
| Characteristics of patients’ samples . | Myeloma cell death (%) . | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Number . | Disease . | Sample . | Stage . | 1q21, 17p . | t(4;14) . | t(11;14) . | TP53 . | S63845 . | venetoclax . | S63845 + venetoclax . |
| 1 | MM | BM | D | x | x | 0 | 31 | 78 | ||
| 2 | MM | BM | D | 1q21 | x | 4 | 5 | 94 | ||
| 3 | MM | BM | D | t(4;14) | x | 5 | 9 | 73 | ||
| 4 | MM | BM | D | 1q21 | x | x | 33 | 33 | 81 | |
| 5 | MM | BM | D | 1q21 | t(11;14) | 98 | 12 | 99 | ||
| 6 | MM | BM | D | 1q21 | t(4;14) | x | 21 | 44 | 100 | |
| 7 | MM | BM | D | x | x | 0 | 25 | 58 | ||
| 8 | MM | BM | D | del17p | x | 5 | 15 | 44 | ||
| 9 | MM | BM | D | t(11;14) | 63 | 60 | 98 | |||
| 10 | MM | BM | D | del17p | x | 13 | 6 | 74 | ||
| 11 | MM | BM | D | x | x | 18 | 0 | 32 | ||
| 12 | MM | BM | D | 1q21 | x | x | 23 | 8 | 72 | |
| 13 | MM | BM | D | del17p | 5 | 1 | 2 | |||
| 14 | MM | BM | D | 1q21, del17p | t(11;14) | 14 | 85 | 90 | ||
| 15 | MM | BM | D | 1q21 | t(11;14) | 19 | 1 | 59 | ||
| 16 | MM | BM | D | x | 31 | 38 | 73 | |||
| 17 | MM | BM | D | 16 | 53 | 83 | ||||
| 18 | MM | BM | D | del17p | H193R | 0 | 51 | 95 | ||
| 19 | MM | BM | D | x | 0 | 0 | 3 | |||
| 20 | MM | BM | D | x | 17 | 12 | 73 | |||
| 21 | MM | BM | D | t(11;14) | x | 38 | 95 | 94 | ||
| 22 | MM | BM | D | 1q21 | x | x | x | 30 | 12 | 44 |
| 23 | MM | BM | D | t(11;14) | x | 15 | 79 | 85 | ||
| 24 | MM | BM | D | 1q21 | x | x | 81 | 47 | 88 | |
| 25 | MM | BM | D | x | 0 | 21 | 66 | |||
| 26 | MM | BM | D | 1q21 | x | 0 | 3 | 34 | ||
| 27 | MM | BM | D | x | x | x | 44 | 11 | 100 | |
| 28 | MM | BM | D | x | 0 | 24 | 70 | |||
| 29 | MM | BM | D | 1q21 | x | 37 | 29 | 71 | ||
| 30 | MM | BM | D | 1q21, del17p | t(4;14) | x | 73 | 10 | 96 | |
| 31 | MM | BM | D | 1q21 | t(11;14) | 33 | 19 | 95 | ||
| 32 | MM | BM | D | 1q21 | t(4;14) | x | 72 | 0 | 84 | |
| 33 | MM | BM | P | 1q21 | 12 | 0 | 38 | |||
| 34 | MM | BM | P | 1q21 | t(4;14) | 28 | 10 | 98 | ||
| 35 | MM | BM | R | t(11;14) | 56 | 57 | 98 | |||
| 36 | MM | BM | R | 1q21, del17p | x | x | 24 | 86 | 90 | |
| 37 | MM | BM | R | t(11;14) | x | 37 | 35 | 79 | ||
| 38 | MM | BM | R | 0 | 0 | 83 | ||||
| 39 | MM | BM | R | t(4;14) | 39 | 12 | 64 | |||
| 40 | MM | BM | R | 1q21 | t(11;14) | x | 66 | 45 | 98 | |
| 41 | MM | BM | R | 1q21, del17p | t(4;14) | 21 | 5 | 26 | ||
| 42 | MM | BM | R | 1q21 | 30 | 0 | 70 | |||
| 43 | MM | BM | R | 1q21 | t(4;14) | 0 | 0 | 65 | ||
| 44 | MM | BM | R | x | 38 | 7 | 77 | |||
| 45 | MM | BM | R | 1q21 | x | x | 1 | 0 | 22 | |
| 46 | MM | BM | R | 1q21 | t(11;14) | x | 15 | 40 | 65 | |
| 47 | MM | BM | R | 1q21 | x | 49 | 25 | 91 | ||
| 48 | MM | BM | R | 1q21 | x | 41 | 0 | 73 | ||
| 49 | MM | BM | R | del17p | x | 1 | 24 | 29 | ||
| 50 | MM | BM | R | 1q21 | x | 6 | 17 | 59 | ||
| 51 | MM | BM | R | 1q21 | t(11;14) | 75 | 73 | 100 | ||
| 52 | MM | BM | R | 1q21 | x | 9 | 0 | 53 | ||
| 53 | MM | BM | R | del17p | t(11;14) | x | 61 | 81 | 96 | |
| 54 | MM | BM | R | 1q21 | t(4;14) | x | 29 | 27 | 92 | |
| 55 | MM | PE | R | 1q21 | t(4;14) | x | 12 | 0 | 57 | |
| 56 | MM | BM | R | 44 | 41 | 74 | ||||
| 57 | MM | BM | R | x | x | 0 | 18 | 83 | ||
| 58 | MM | BM | R | 1q21 | 2 | 87 | 96 | |||
| 59 | MM | BM | R | x | t(11;14) | x | 2 | 8 | 63 | |
| 60 | MM | BM | R | x | x | 13 | 24 | 98 | ||
| 61 | MM | BM | R | 1q21 | x | x | 32 | 18 | 92 | |
| 62 | MM | BM | R | 1q21, del17p | x | 70 | 16 | 97 | ||
| 63 | MM | BM | R | 1q21 | t(4;14) | 10 | 4 | 87 | ||
| 64 | MM | BM | R | 1q21 | t(11;14) | 54 | 56 | 86 | ||
| 65 | MM | BM | R | 1q21, del17p | 55 | 24 | 98 | |||
| 66 | MM | BM | R | t(11;14) | x | 0 | 92 | 94 | ||
| 67 | MM | BM | R | 1q21, del17p | x | 74 | 0 | 91 | ||
| 68 | MM | BM | R | t(11;14) | x | 26 | 87 | 92 | ||
| 69 | MM | BM | R | 1q21 | t(11;14) | x | 6 | 93 | 99 | |
| 70 | MM | BM | R | x | x | x | 0 | 0 | 21 | |
| 71 | MM | BM | R | x | x | 40 | 28 | 71 | ||
| Characteristics of patients’ samples . | Myeloma cell death (%) . | |||||||||
|---|---|---|---|---|---|---|---|---|---|---|
| Number . | Disease . | Sample . | Stage . | 1q21, 17p . | t(4;14) . | t(11;14) . | TP53 . | S63845 . | venetoclax . | S63845 + venetoclax . |
| 1 | MM | BM | D | x | x | 0 | 31 | 78 | ||
| 2 | MM | BM | D | 1q21 | x | 4 | 5 | 94 | ||
| 3 | MM | BM | D | t(4;14) | x | 5 | 9 | 73 | ||
| 4 | MM | BM | D | 1q21 | x | x | 33 | 33 | 81 | |
| 5 | MM | BM | D | 1q21 | t(11;14) | 98 | 12 | 99 | ||
| 6 | MM | BM | D | 1q21 | t(4;14) | x | 21 | 44 | 100 | |
| 7 | MM | BM | D | x | x | 0 | 25 | 58 | ||
| 8 | MM | BM | D | del17p | x | 5 | 15 | 44 | ||
| 9 | MM | BM | D | t(11;14) | 63 | 60 | 98 | |||
| 10 | MM | BM | D | del17p | x | 13 | 6 | 74 | ||
| 11 | MM | BM | D | x | x | 18 | 0 | 32 | ||
| 12 | MM | BM | D | 1q21 | x | x | 23 | 8 | 72 | |
| 13 | MM | BM | D | del17p | 5 | 1 | 2 | |||
| 14 | MM | BM | D | 1q21, del17p | t(11;14) | 14 | 85 | 90 | ||
| 15 | MM | BM | D | 1q21 | t(11;14) | 19 | 1 | 59 | ||
| 16 | MM | BM | D | x | 31 | 38 | 73 | |||
| 17 | MM | BM | D | 16 | 53 | 83 | ||||
| 18 | MM | BM | D | del17p | H193R | 0 | 51 | 95 | ||
| 19 | MM | BM | D | x | 0 | 0 | 3 | |||
| 20 | MM | BM | D | x | 17 | 12 | 73 | |||
| 21 | MM | BM | D | t(11;14) | x | 38 | 95 | 94 | ||
| 22 | MM | BM | D | 1q21 | x | x | x | 30 | 12 | 44 |
| 23 | MM | BM | D | t(11;14) | x | 15 | 79 | 85 | ||
| 24 | MM | BM | D | 1q21 | x | x | 81 | 47 | 88 | |
| 25 | MM | BM | D | x | 0 | 21 | 66 | |||
| 26 | MM | BM | D | 1q21 | x | 0 | 3 | 34 | ||
| 27 | MM | BM | D | x | x | x | 44 | 11 | 100 | |
| 28 | MM | BM | D | x | 0 | 24 | 70 | |||
| 29 | MM | BM | D | 1q21 | x | 37 | 29 | 71 | ||
| 30 | MM | BM | D | 1q21, del17p | t(4;14) | x | 73 | 10 | 96 | |
| 31 | MM | BM | D | 1q21 | t(11;14) | 33 | 19 | 95 | ||
| 32 | MM | BM | D | 1q21 | t(4;14) | x | 72 | 0 | 84 | |
| 33 | MM | BM | P | 1q21 | 12 | 0 | 38 | |||
| 34 | MM | BM | P | 1q21 | t(4;14) | 28 | 10 | 98 | ||
| 35 | MM | BM | R | t(11;14) | 56 | 57 | 98 | |||
| 36 | MM | BM | R | 1q21, del17p | x | x | 24 | 86 | 90 | |
| 37 | MM | BM | R | t(11;14) | x | 37 | 35 | 79 | ||
| 38 | MM | BM | R | 0 | 0 | 83 | ||||
| 39 | MM | BM | R | t(4;14) | 39 | 12 | 64 | |||
| 40 | MM | BM | R | 1q21 | t(11;14) | x | 66 | 45 | 98 | |
| 41 | MM | BM | R | 1q21, del17p | t(4;14) | 21 | 5 | 26 | ||
| 42 | MM | BM | R | 1q21 | 30 | 0 | 70 | |||
| 43 | MM | BM | R | 1q21 | t(4;14) | 0 | 0 | 65 | ||
| 44 | MM | BM | R | x | 38 | 7 | 77 | |||
| 45 | MM | BM | R | 1q21 | x | x | 1 | 0 | 22 | |
| 46 | MM | BM | R | 1q21 | t(11;14) | x | 15 | 40 | 65 | |
| 47 | MM | BM | R | 1q21 | x | 49 | 25 | 91 | ||
| 48 | MM | BM | R | 1q21 | x | 41 | 0 | 73 | ||
| 49 | MM | BM | R | del17p | x | 1 | 24 | 29 | ||
| 50 | MM | BM | R | 1q21 | x | 6 | 17 | 59 | ||
| 51 | MM | BM | R | 1q21 | t(11;14) | 75 | 73 | 100 | ||
| 52 | MM | BM | R | 1q21 | x | 9 | 0 | 53 | ||
| 53 | MM | BM | R | del17p | t(11;14) | x | 61 | 81 | 96 | |
| 54 | MM | BM | R | 1q21 | t(4;14) | x | 29 | 27 | 92 | |
| 55 | MM | PE | R | 1q21 | t(4;14) | x | 12 | 0 | 57 | |
| 56 | MM | BM | R | 44 | 41 | 74 | ||||
| 57 | MM | BM | R | x | x | 0 | 18 | 83 | ||
| 58 | MM | BM | R | 1q21 | 2 | 87 | 96 | |||
| 59 | MM | BM | R | x | t(11;14) | x | 2 | 8 | 63 | |
| 60 | MM | BM | R | x | x | 13 | 24 | 98 | ||
| 61 | MM | BM | R | 1q21 | x | x | 32 | 18 | 92 | |
| 62 | MM | BM | R | 1q21, del17p | x | 70 | 16 | 97 | ||
| 63 | MM | BM | R | 1q21 | t(4;14) | 10 | 4 | 87 | ||
| 64 | MM | BM | R | 1q21 | t(11;14) | 54 | 56 | 86 | ||
| 65 | MM | BM | R | 1q21, del17p | 55 | 24 | 98 | |||
| 66 | MM | BM | R | t(11;14) | x | 0 | 92 | 94 | ||
| 67 | MM | BM | R | 1q21, del17p | x | 74 | 0 | 91 | ||
| 68 | MM | BM | R | t(11;14) | x | 26 | 87 | 92 | ||
| 69 | MM | BM | R | 1q21 | t(11;14) | x | 6 | 93 | 99 | |
| 70 | MM | BM | R | x | x | x | 0 | 0 | 21 | |
| 71 | MM | BM | R | x | x | 40 | 28 | 71 | ||
Mononuclear cells from the BM or PE of patients with myeloma were cultured for 24 hours with 25 nM S63845, 300 nM venetoclax, or their combination, and myeloma cell death was assessed using flow cytometry. Del17p, 1q21 gain, t(4,14) and t(11;14) were determined by FISH (threshold > 50% of cells). TP53 sequencing was assessed by Sanger sequencing of complementary DNA products. TP53 mutation was considered when it involved the majority of cells.
BM, bone marrow; D, diagnosis; P, progression; PE, pleural effusion; R, relapse; x, not performed.
Myeloma cells resistant to BH3 mimetics displayed a lower p53 score at single-cell level
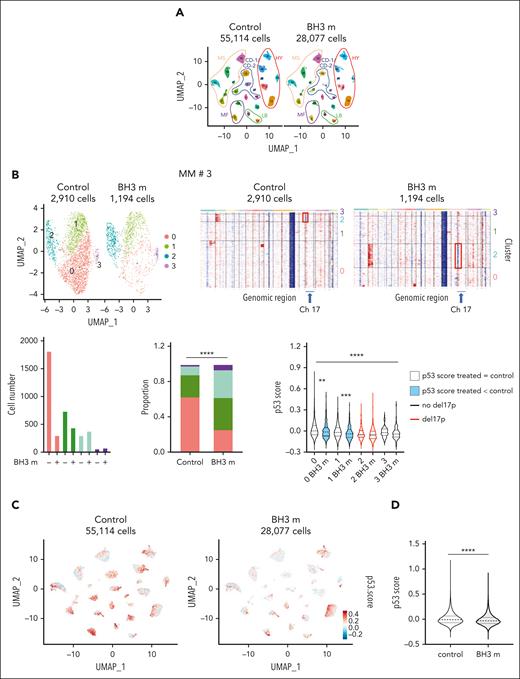
Given the p53 impact on the response to BH3 mimetics and the heterogeneity of myeloma cells in patients, we used scRNA-seq to determine whether cells surviving BH3 mimetics would have a lower p53 score than control cells. Bone marrow (n = 23) or pleural effusion (n = 1) mononuclear cells from 24 patients (supplemental Table 10) were cultured overnight with or without BH3 mimetics combination, and 15 000 living cells were separated and profiled using 10X Chromium. Myeloma cells were identified as described in supplemental Materials and methods. Figure 6A shows the 83 191 myeloma cells from the 24 control (55 114 control cells) and BH3 mimetics–treated samples (28 077 treated cells) and their molecular classification: 21 samples could be classified as CCND1-1 (CD-1), CCND1-2 (CD-2), hyperdiploid (HY), low bone disease (LB), MAF (MF), and MMSET (MS) (supplemental Figure 11).36 The median number of myeloma cells analyzed per sample was 2284 cells and 643 cells in control and BH3 mimetics–treated conditions, respectively (supplemental Figure 12). In 15 of 24 samples, the proportion of cells in paired clusters was significantly modified upon BH3 mimetics treatment as illustrated for sample no. 3 in Figure 6B (P < .0001; all samples are shown in supplemental Figure 12). Del17p status of clusters was determined using Infer copy number variation, clusters with del17p being shown in red (Figure 6B). The p53 score median value in clusters was compared between paired control and treated samples (when cluster contained at least 10 cells in both conditions): in BH3 mimetics–treated samples, the score was unchanged in 51 clusters (66.2%), inferior in 25 clusters (32.5%), and superior in 1 cluster (1.3%). Change in score was not similar in clusters with or without del17p (P = .035, χ2 test), that is, it decreased in 35.4% of clusters with no del17p (23 out of 65) vs 16.7% of clusters with del17p (2 out of 12; supplemental Figure 12). The p53 score value was compared between control and treated cells: the p53 score was 3.13-fold inferior in treated vs control cells (median values were −0.00961 and −0.03004, respectively; P < .0001; Figure 6C-D).
scRNA-seq analysis of response to BH3 mimetics. (A) Uniform Manifold Approximation and Projection (UMAP) representation of myeloma cells from 24 control and BH3 mimetics–treated patient samples using scRNA-seq. Mononuclear cells from 23 bone marrow or 1 pleural effusion from patients with MM were cultured overnight with interleukin-6 in the presence, or absence, of 25 nM S63845 and 300 nM venetoclax (BH3 m). Cell processing and scRNA-seq analysis were performed as described in supplemental Materials and methods. Each patient sample is numbered according to supplemental Table 10, and their molecular classification is indicated. (B) Example of individual scRNA-seq analysis of response to BH3 mimetics. (Top left) UMAP split representation of myeloma cells in merged control and treated samples from patient no. 3; cells are colored according to clusters. (Top right) Infer copy number variation (InferCNV) of control and BH3 mimetics–treated cells. InferCNV was calculated using broadinstitute/infercnv package in R in comparison with normal bone marrow plasma cells. (Bottom left) The graphs represent the number of cells per cluster and their proportion in each condition. (Right) Statistical comparison of p53 score in paired clusters (Mann-Whitney U test). Black and red violins indicate no del17p and del17p, respectively. All samples are shown in supplemental Figure 12. (C-D) The p53 score is decreased in cells surviving to BH3 mimetic combination. (C) UMAP representation of p53 score of myeloma cells split into control (left) and treated (right) samples (blue = low; red = high). The p53 score was calculated independently in each patient samples. (D) The graph represents the violin representation of p53 score in 55 114 control cells and 28 077 BH3 mimetic–treated cells from the 24 samples. Statistical analysis was performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
scRNA-seq analysis of response to BH3 mimetics. (A) Uniform Manifold Approximation and Projection (UMAP) representation of myeloma cells from 24 control and BH3 mimetics–treated patient samples using scRNA-seq. Mononuclear cells from 23 bone marrow or 1 pleural effusion from patients with MM were cultured overnight with interleukin-6 in the presence, or absence, of 25 nM S63845 and 300 nM venetoclax (BH3 m). Cell processing and scRNA-seq analysis were performed as described in supplemental Materials and methods. Each patient sample is numbered according to supplemental Table 10, and their molecular classification is indicated. (B) Example of individual scRNA-seq analysis of response to BH3 mimetics. (Top left) UMAP split representation of myeloma cells in merged control and treated samples from patient no. 3; cells are colored according to clusters. (Top right) Infer copy number variation (InferCNV) of control and BH3 mimetics–treated cells. InferCNV was calculated using broadinstitute/infercnv package in R in comparison with normal bone marrow plasma cells. (Bottom left) The graphs represent the number of cells per cluster and their proportion in each condition. (Right) Statistical comparison of p53 score in paired clusters (Mann-Whitney U test). Black and red violins indicate no del17p and del17p, respectively. All samples are shown in supplemental Figure 12. (C-D) The p53 score is decreased in cells surviving to BH3 mimetic combination. (C) UMAP representation of p53 score of myeloma cells split into control (left) and treated (right) samples (blue = low; red = high). The p53 score was calculated independently in each patient samples. (D) The graph represents the violin representation of p53 score in 55 114 control cells and 28 077 BH3 mimetic–treated cells from the 24 samples. Statistical analysis was performed using the Mann-Whitney U test. ∗∗∗∗P < .0001; ∗∗∗P < .001; ∗∗P < .01; ∗P < .05.
Discussion
In our work, we performed CRISPR/Cas9 TP53 invalidation to study the direct genomic and functional impact of p53 silencing in myeloma cells. Given the myeloma genomic heterogeneity, we chose 2 HMCLs belonging to the same subgroup, the t(4;14) group. Using RNA-seq, 17 genes were significantly regulated upon TP53 invalidation in both TP53+/+ HMCLs, 16 down and 1 up. Using microarray, only 5 genes were underexpressed (MDM2, CDKN1A, DDB2, FDXR, and PHLDA3) and 1 overexpressed (CDKN2A) in TP53abn vs TP53wt HMCLs; by contrast to the 5 underexpressed genes, CDKN2A expression was not modified in isogenic clones. CDKN2A is not known as a p53 target, and its overexpression in TP53abn HMCLs is, in fact, related to CDKN2A/ARF locus loss in some TP53wt HMCLs,21 which is a common hit in TP53wt cancer cells. This difference illustrates the major interest of isogenic cells for providing a specific p53 transcriptome. As expected, none of the genes affected by TP53 invalidation in both NCI-H929 and XG7 were found modified in JIM3 or NAN3 TP53−/− clones, except TP53, showing that p53 with R273C or R248Q mutation had no residual transcriptomic activity. Although the expression of several genes was found modified in either JIM3 or NAN3 TP53−/− clones, the modest changes instead suggested nonspecific regulation.
To establish a functional p53 score, we kept the underexpressed genes in both NCI-H929 and XG7 TP53−/− clones, known as p53 targets: these 13 genes were identified as early transactivated upon p53 activation, highlighting their high sensitivity to p53 level.28 This score segregated TP53+/+ from TP53−/− or TP53−/mut cells, underscoring its efficacy for identifying p53 loss of function, whatever its origin, that is, lack of expression or mutation, and whatever the genomic background. In 1105 cancer cell lines, the score also distinguished cells with monoallelic or biallelic TP53 hit (deletion and/or mutation) from cells without any TP53 abnormalities. Monoallelic hit was associated with a low score, and cells displaying biallelic hit had an even lower score. In patient cells, samples with low score were mostly del17p in CASSIOPEA (TP53 sequence was not assessed) or del17p and TP53 mutated in MYRACLE and MMRF-coMMpass. Indeed, in MMRF-coMMpass cohort, TP53 deletion alone had only a trend to affect the score (P = .18). This discrepancy between patient cells and cell lines might be explained by the low number of del17p samples in patient cohorts and/or by the clonal proportion (100% of cells in cell lines but not in patient samples) or by the biallelic vs monoallelic deletion, with the biallelic deletion being more frequent in cell lines than in patient cells. Taken together, this functional p53 score is highly efficient in identifying samples with biallelic inactivation both in cell lines and patient samples.
The score will not identify samples with TP53 hit in minor subpopulations in contrast to FISH or next-generation sequencing. However, this functional score identified few samples lacking any TP53 hit, suggesting the existence of other p53 pathway dysregulations that merit further investigation. Of interest, the score was also heterogeneous at the single-cell level, suggesting that it identified cells with low to high activation of p53 pathway.
Interestingly, the score was predictive of overall survival in 2 patient cohorts; it, indeed, identified 3% to 5% of patients with a low p53 score, which is consistent with the low occurrence of mutation/biallelic hit in TP53 gene.16,37 In 139 patients from the CASSIOPEA trial (VTD [bortezomib, thalidomide, and dexamethasone] and daratumumab vs VTD), 75% of patients with a low score were classified as being at high risk (Revised International Staging System), and 62.5% had del17p; del17p was associated with reduced survival, but the impact of monoallelic vs biallelic hit was not assessed. In the MMRF-coMMpass cohort, which is similar to MYRACLE, a real-life cohort, only del17p and mutation was predictive of overall survival, and p53 score, indeed, identified patients with biallelic hit (53% of biallelic samples were in group 1). Deletion alone was not associated with significant lower overall survival, in good agreement with its lack of impact on the score.
The p53 score involves a combination of genes that has not been used in other p53 score either derived from shTP53 HMCLs or from MMRF-CoMMpass study.35,38,39 Our p53 functional score also did not share any gene with the “Li-Fraumeni” functional p53 score that was unable to discriminate 1105 cell lines, 12 clones, and 38 patient samples according to their respective TP53 status.
As expected, TP53 silencing in XG7 and NCI-H929 HMCLs increased resistance to melphalan and nutlin3a. Functional impact of monoallelic deletion in isogenic cell lines could not be assessed because both alleles were inactivated in all clones. However, Munawar et al previously demonstrated that the monoallelic deletion significantly impaired drug sensitivity in isogenic AMO1 cells.40
Among the 13 genes of the score, we focused on BAX, given its major role in apoptosis. Although some BCL2 family members such as PMAIP1 (NOXA) and BBC3 (PUMA) are known to be regulated by p53, only BAX was significantly modified in TP53−/− clones in the absence of stress. Indeed, PMAIP1 or BBC3 transactivation requires a higher amount of p53, as illustrated by their activation by nutlin3a that induced p53 accumulation.28,32,33 Although BAX and BAK are considered as redundant, only BAX expression is directly regulated by p53. To decipher its role, we used BH3 mimetics that directly target mitochondria without inducing genomic or signaling stress, in contrast to DNA-damaging drugs such as melphalan. In MM, sensitivity to BH3 mimetics is primarily associated with genomic heterogeneity, with t(11;14) cells being sensitive to BCL2 BH3 mimetics and t(4;14) to MCL1 BH3 mimetics.12 NCI-H929 and XG7 were sensitive to S63845 but resistant to venetoclax, and removal of p53 induced approximately fivefold increase in S63845 LD50 values. Transient or permanent silencing of BAX or BAK1 in XG7 or NCI-H929 demonstrated that BAX (not BAK) expression was essential for the death response to low doses of S63845, whatever the level of BAK expression. These results clearly demonstrated that BAX decrease in TP53−/− clones greatly affected mitochondrial fitness despite a slight increase in the amount of MCL1/BAK complexes; indeed, MCL1 immunoprecipitation assays showed that the amount of MCL1-BAX was greatly decreased in TP53−/− clones and that low concentrations of S63845 totally disrupted MCL1-BAX and partly MCL1-BAK complexes. Although BAX and BAK affinity for MCL1 is quite similar, BAK has been reported to display a higher affinity than BAX that could favor the release of BAX under low concentrations of S63845.41,42 These results showed that BAK is not a requisite for a response to low or intermediate doses of S63845. Moreover, the impact of p53 and BAX (but not BAK) on MCL1-sensitivity was also found in HMCLs: S63845 LD50 values were correlated with TP53 status and with BAX expression level. Overall, these results show that BAK and BAX are not completely redundant and that p53-induced BAX expression is a limiting factor in mitochondrial apoptosis. With most TP53wt HMCLs being resistant to BCL2 or BCLXL inhibitors, p53/BAX impact on other BH3 mimetics response could not be assessed in myeloma. Nonetheless, in other hematological malignancies, loss of TP53 was also reported to impair response to BCL2 or MCL1 BH3 mimetic.43-45 In chronic lymphocytic leukemia, venetoclax with obinutuzumab was efficient but an inferior efficacy was reported in patients with TP53 hit compared with in patients without TP53 hit.46 Although BCL2 and MCL1 BH3 mimetics combination was more efficient in TP53+/+ clones, it overcame the resistance induced by p53 loss in both NCI-H929 and XG7 TP53−/− clones and was highly efficient in 69% of del17p samples from patients (9 of 13). At single-cell level, we showed in 24 paired control and treated samples that the 13-gene p53 score was heterogeneous; cells surviving BH3 mimetics combination in vitro mainly displayed either a decreased or unchanged p53 score, suggesting, again, that cells with high p53 score were more sensitive to BH3 mimetics than cells with lower score. Finally, our data showed that a functional p53 was important for achieving an optimal response to the BH3 mimetics combination, which was, however, not abrogated in TP53-deficient cells.
In summary, we established a functional p53 score that identifies myeloma cells with biallelic TP53 invalidation, demonstrated that p53-regulated BAX is critical for optimal cell response to BH3 mimetics, and showed that BH3 mimetics combination may be of interest for patients with biallelic TP53 invalidation, for whom there is still an unmet medical need.
Acknowledgments
The authors acknowledge the Cytocell-Flow Cytometry and FACS core facility (Structure Fédérative de Recherche (SFR) Bonamy, BioCore, Inserm UMS 016, CNRS UAR 3556, Nantes, France) for its technical expertise and help, member of the Scientific Interest Group Biogenouest and the Labex IGO program supported by the French National Research Agency (n°ANR-11-LABX-0016-01). The authors thank the Genomics Core Facility GenoA, member of Biogenouest and France Genomique and the Bioinformatics Core Facility BiRD, member of Biogenouest and Institut Français de Bioinformatique (ANR-11-INBS-0013) for the use of their resources and their technical support. The authors thank Elise Douillard and Magali Devic for expert technical assistance, and Emma Leguedey for performing experiments. The coMMpass data were generated as part of the Multiple Myeloma Research Foundation Personalized Medicine Initiative.
This work was supported by grants from Fondation Française pour la Recherche contre le Myélome et les Gammapathies (FFRMG), Ligue Nationale Contre le Cancer (LNCC), Institut National du Cancer (INCA; INCa-DGOS-INSERM_12558, INCa-DGOS-INSERM-ITMO Cancer_18011), and Action Cancer 44. The study was supported by SIRIC ILIAD (R.D.) and by HEMA-NexT, n°ANR-16-IDEX-0007 (N.L.L.).
Authorship
Contribution: R.D. participated in the design of the study, performed experiments, analyzed the results, and participated in the writing; G.D. and C.D. derived, profiled CRISPR/Cas9 TP53 cells, and performed experiments; S. Maïga managed the patient cohorts and performed experiments; C.B. performed bioinformatic analysis and p53 scoring; J.-B.A. performed TP53 analysis in MMRF-coMMpass; J.-B.A., J.C., and J.D. performed scRNA-seq analysis; N.L.L. managed the patient cohorts; B.T., C.T., and P.M. participated in the design of the study and managed the patient cohorts; C.G. and Y.L.B. provided samples and supervised fluorescence in situ hybridization experiments; S. Minvielle provided CASSIOPEA RNA-seq data; D.C. and A.M.-A. designed CRISPR/Cas9 strategy and derived CRISPR/Cas9 clones; P.G.-B., M.A., D.C., and S. Minvielle participated in the design of scRNA-seq study and supervised experiments; and C.P.-D. designed the study, analyzed the results, and wrote the article.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Catherine Pellat-Deceunynck, Centre de Recherche en Cancérologie et Immunologie Intégrée Nantes Angers, INSERM, CNRS, Institut de Recherche en Santé de Nantes Université (IRS-UN), 8 quai Moncousu, BP70721, F-44007 Nantes, France; email: catherine.pellat-deceunynck@univ-nantes.fr; and Romane Durand, Centre de Recherche en Cancérologie et Immunologie Intégrée Nantes Angers, INSERM, CNRS, IRS-UN, 8 quai Moncousu, BP70721, F-44007 Nantes, France; email: romane.durand@univ-nantes.fr.
References
Author notes
G.D., C.B., C.D., and S. Maïga contributed equally to this study.
The data reported in this article have been deposited in the Gene Expression Omnibus database (accession number GSE245163) and in the European Genome-Phenome Archive (EGA50000000044).
Data are available on request from the corresponding author, Catherine Pellat-Deceunynck (catherine.pellat-deceunynck@univ-nantes.fr).
The online version of this article contains a data supplement.
There is a Blood Commentary on this article in this issue.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal