Abstract
Abstract 1437
Genetic heterogeneity is common not only in solid tumors, but also in leukemias. The analysis of genetic heterogeneity among single cancer cells is vital for a better understanding of cancer evolution and therapeutic failure of systemic cancer therapy. So far, comprehensive genome-wide single cell studies were limited by many technical difficulties. Here, we present a novel approach, combining adapter-linker PCR based whole genome amplification (WGA) with 2nd generation sequencing, that enables comprehensive and comparative genome-wide analysis of single leukemic cells.
WGA, based on adapter-linker PCR (Klein et al PNAS 1999, Stoecklein et al Cancer Cell 2008), of three individually picked cells of the permanent leukemia cell line REH was performed. WGA products, subsequently fragmented to 100 bp or 250 bp, were used for library preparation. After loading one amplified single cell genome per flowcell, DNA was sequenced with paired end (PE) reads (2× 75bp or 2× 100 bp respectively) on a Genome Analyzer IIx or a HiSeq 2000 (Illumina). After alignment with Burrows-Wheeler Aligner (BWA), removal of duplicate read pairs, and identification of SNPs by the Genome Analysis Toolkit (GATK), copy number variants (CNV), loss of heterozygosity (LOH) and allele dropout rates were analyzed, based on the human reference genome (hg19/GRCh37). Results were compared to data obtained by hybridizing pooled gDNA of REH cells of the same passage to a SNP 6.0 array (Affymetrix). Interchromosomal translocations were determined in single cells of the same passage of REH cells by spectral karyotyping (SKY) and compared to sequencing data, analyzed by Geometric Analysis of Structural Variants (GASV).
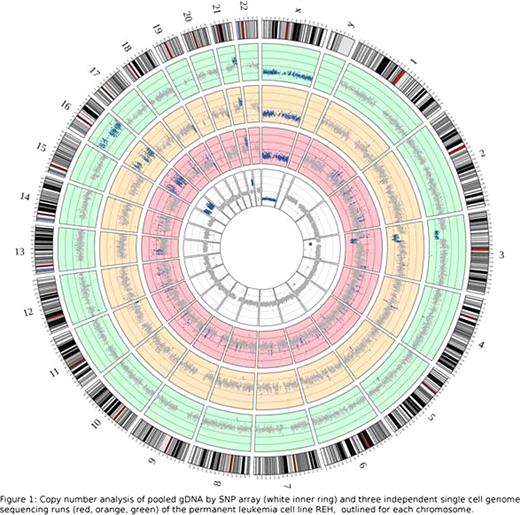
With our approach we obtained up to 600 mio mappable reads per run, evenly spread over the genome, which led to a sequence coverage of up to 67%, with an even higher coverage of coding sequence (76%) and a sequence depth of 16x. Comparison of SNP arraydata with PE sequencing data showed, that they are highly overlapping (99,3%) regarding the detection of normal copy numbers. But also for copy number alterations, consistency between both methods was observed in detecting losses (94.1%) or gains (77.1%) of genomic material (figure 1). Up to 97% of regions of LOH detected by sequencing, were also detected by the SNP array, when analyzed in a resolution of 500K bp. By analyzing the data with higher resolutions of up to 10K bp, an increasing amount of regions of LOH could be detected. However, decreased correlation between SNP array and sequencing data (max. 74.5%) was observed, with high correlation between the sequencing runs (85%). This indicates increased detection of false positive LOH regions by the SNP array and the sequencing approach to be superior in this high resolution. To assess the allele dropout rate as a quality control for the PCR based WGA method, the heterozygous SNPs detected by PE sequencing were compared to those called by the SNP array. High consistency (95%) indicates an allele dropout rate of only 5%.
To analyze the accuracy of our approach in detecting genetic heterogeneity between single cells, we assessed the variability in the SNP profile between the three individual cells. As they are derived from a permanent cell line, they are expected to be highly similar. In fact, the SNPs, that were covered in all three sequencing runs showed a variation of less than 0,1% among the single REH cells. As the SNP array is not applicable to asses copy number neutral variations as translocations, the karyotype of REH cells was assessed by SKY, confirming the predescribed translocations t(4;12), t(4;16), t(5;12), t(16;21) and t(12;21). Breakpoint regions comparable to those defined by SKY, were identified for all 5 translocations by analysis of discordant read pairs with GASV. The detection of additional, exclusively by sequencing identified breakpoints, is currently under intensified investigation, to confirm potentially newly discovered breakpoints and reliably rule out false positive results.
Our approach provides a powerful tool to achieve an unprecedented genome-wide overview on genomic variations of single cells. The robustness of our single cell approach in comparison to the data acquired with pooled gDNA and the homogeneity of our results in the permanent REH cell line clearly shows the reliability of our approach to assess single cell heterogeneity in primary leukemic samples.
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.