Abstract
We performed a comprehensive genome-wide miRNA expression profiling of extranodal nasal-type natural killer/T-cell lymphoma (NKTL) using formalin-fixed paraffin-embedded tissue (n = 30) and NK cell lines (n = 6) compared with normal NK cells, with the objective of understanding the pathogenetic role of miRNA deregulation in NKTL. Compared with normal NK cells, differentially expressed miRNAs in NKTL are predominantly down-regulated. Re-expression of down-regulated miRNAs, such as miR-101, miR-26a, miR26b, miR-28-5, and miR-363, reduced the growth of the NK cell line and modulated the expression of their predicted target genes, suggesting the potential functional role of the deregulated miRNAs in the oncogenesis of NKTL. Taken together, the predicted targets whose expression is inversely correlated with the expression of deregulated miRNA in NKTL are significantly enriched for genes involved in cell cycle-related, p53, and MAPK signaling pathways. We also performed immunohistochemical validation for selected target proteins and found overexpression of MUM1, BLIMP1, and STMN1 in NKTL, and notably, a corresponding increase in MYC expression. Because MYC is known to cause repression of miRNA expression, it is possible that MYC activation in NKTL may contribute to the suppression of the miRNAs regulating MUM1, BLIMP1, and STMN1.
Introduction
Extranodal nasal-type natural killer/T-cell lymphoma (NKTL) is an aggressive lymphoma with a strong association with EBV. The pathogenesis of this tumor is poorly understood, but in recent years gene expression profiling (GEP) studies have demonstrated the pathogenetic role of several oncogenic pathways in NKTL, such as AKT, STAT3, NF-κB, Notch-1, and Aurora kinase A.1,2 We recently performed a genome-wide GEP using formalin-fixed paraffin-embedded (FFPE) tissue and, in addition to NF-κB, we also identified deregulation of c-Myc and p53 pathways, and overexpression of survivin in NKTL.3
MicroRNAs (miRNAs) are short, noncoding RNAs that post-transcriptionally regulate the expression of multiple mRNAs. To date, > 1000 human miRNA precursor sequences have been identified and deposited in miRBase.4 miRNAs play a key role in the control of normal biologic processes, including hematopoiesis, and have been implicated in the development of human cancer.5,6 In lymphoid malignancies, miR-155 is overexpressed in Hodgkin lymphoma and non-Hodgkin lymphoma, and dysregulation of miR-16-1 control of cyclinD1 has been reported in mantle cell lymphoma. Yamanaka et al performed northern analysis on NKTL using a limited number of probe sets and found overexpression of miR-155 and miR-21, which results in the activation of AKT signaling.7 Furthermore, quantification of miRNAs can have potential diagnostic and prognostic utility in lymphoma.5,6,8 miRNA expression profiling (MEP) has been increasingly used in cancer research; and in recent years, it has been possible to obtain meaningful and reproducible profiles using FFPE tissue.9 To the best of our knowledge, there have been no reports of genome-wide MEP on NKTL in the published literature.
In this study, we performed the first miRNA expression profiling on a series of NKTL using FFPE tissues in relation to normal NK cells and NK tumor cell lines, with the main objective of understanding the pathogenic role and mechanisms of miRNA dysregulation in NKTL. We also performed a combined analysis of the miRNA profiles, and the gene expression profiles obtained in our previous study using bioinformatics target prediction with subsequent functional validation to identify essential target genes and signaling pathways that are deregulated by miRNA in NKTL.
Methods
Case selection, cell lines, and control tissues
Patients with a diagnosis of NKTL were identified from the archives of the Department of Pathology, National University Hospital, from 1990 to 2010 and classified according to the 2008 WHO lymphoma classification. Cases with no additional tissue available for immunohistochemical or genetic analysis were excluded. A total of 38 cases of NKTL were selected, of which 33 cases were used for tissue microarray construction and 9 cases were subjected to GEP in our previous study.3 According to the WHO criteria, all cases expressed CD3, cytotoxic markers (granzyme B and/or TIA-1), and EBER. Immunoreactivity for CD56, CD8, and CD4 was present in 66% (25 cases), 13% (5 cases), and 5% (2 cases), respectively. The clinical and immunophenotypic data of the cases are summarized in supplemental Table 1 (available on the Blood Web site; see the Supplemental Materials link at the top of the online article).
Thirty cases of NKTL with adequate FFPE tissue and good-quality RNA were selected for miRNA profiling. The study also included 6 NK cell lines (KHYG-1, NK-92, HANK-1, SNT-8, SNK-6, and NK-YS). In addition, 3 paired samples of normal NK cells (unstimulated and stimulated) as well as 2 cases each of normal skin, intestinal, nasal, and lymph node FFPE tissue were also included as control tissue. The study is approved by the Domain Specific Review Board of the National Healthcare Group, Singapore.
NK cell lines and cultures
The NK-tumor cell lines used in this study included NK-92 (ATCC), KHYG-1 (Japanese Collection of Research Bioresources), HANK-1 (gift from Dr Yoshitoyo Kagami), SNK-6, SNT-8 (gift from Dr Norio Shimizu), and NK-YS (gift from Dr YL Kwong). The culture conditions and phenotypic and genotypic characteristics of the NK cell lines, which are well characterized in previous studies,10,11 are summarized in supplemental Table 2. Although 2 of these cells lines are derived from Aggressive Natural Killer-cell leukemia (KHYG and NK-92), only very few miRNAs (24 of 723 miRNAs on chip, 3%) have 2-fold or more difference in expression between these and the other NK/T lymphoma cell lines. They were therefore grouped together as NKTL cell lines for comparison of miRNA expression against tumor samples from patients.
Isolation of normal NK cells from peripheral blood
Highly purified (90%-99%) untouched normal human NK cells were isolated from whole blood samples obtained from healthy donors and buffy coat packs of whole blood samples from the Blood Donation Center, National University Hospital, using the NK cell isolation kit (Miltenyi Biotec) as previously described.3 The isolated NK cells were subsequently stimulated by culturing in the presence of human recombinant IL-2 (Miltenyi Biotec). Cell block preparations of normal NK cells were prepared as previously described.3
RNA extraction from FFPE, NK cell lines, and normal NK cells
Total RNA from NKTL FFPE tissues and FFPE normal tissue controls was isolated using RecoverAll Total Nucleic Acid Isolation (Applied Biosystems) according to the manufacturer's instructions. All the sections were deparaffinized with xylene, subjected to proteinase K digestion, and RNA extracted as per the manufacturer's protocol.
Total RNA was extracted from freshly isolated cells from NK cell lines and normal NK cell samples obtained from healthy donors using miRNeasy mini kit (QIAGEN) protocol with DNaseI treatment included. The concentration and purity of the total RNA extracted were measured using the NanoDrop ND Version 3.0 spectrophotometer (NanoDrop Technologies). RNA quality was assessed with the Agilent 2100 Bioanalyzer (Agilent Technologies) and the RNA 6000 LabChip kit (Agilent Technologies).
miRNA profiling and analysis
miRNA expression was profiled using Agilent human miRNA Microarray Version 2 (Agilent Technologies). Each array contained 60-mer probes representing 723 human and 76 human viral miRNAs from the miRBase Version 10.1. The array experiment was carried out using Agilent miRNA system protocol Version 2.0. Briefly, each RNA sample was labeled with Cyanine3-pCp and hybridized to the Agilent human miRNA microarray using the miRNA Complete Labeling and Hyb Kit (Agilent p/n 5190-0456). The slide was washed using Gene Expression Wash Buffer Kit (Agilent p/n 5188-5327) and then scanned using an Agilent DNA microarray scanner. The raw miRNA expression data were extracted from the scanned image using Agilent Feature Extraction Version 10 software. The raw expression values of miRNA were normalized and analyzed using R Version 2.11.0 and Bioconductor Version 2.8. The microarray data are deposited on the Gene Expression Omnibus (accession number GSE31377).
Transfection of synthetic miRNAs and anti-miRNA inhibitors
miRNA mimics, which are chemically synthesized double-stranded RNA molecules, were designed to mimic endogenous mature miRNAs. They enable detailed study of miRNA biologic effects via gain-of-function experiments.12-14 Cells were transfected with miRNA mimics (Dharmacon RNA Technologies) and anti-miRNA inhibitors (Ambion) at a final concentration of 50nM using DhamarFECT (Dharmacon RNA Technologies) according to the manufacturer's instructions. The control miRNA mimic used was a mimic based on Caenorhabditis elegans miRNA (cel-miR-67). The anti-miRNA inhibitor negative control #1 purchased from Ambion is an RNA oligonucleotide designed to serve as a negative control for experiments involving anti-miRNA inhibitors. Total RNA and protein were collected for assay 2 days after transfection.
Cell growth assay
To generate cell growth curve, cells were harvested and counted at 24-hour intervals. The counting results were validated using the CellTiter 96 AQueous Non-Radioactive Cell Proliferation Assay (Promega) by a linear relationship (r2 = 0.99) between the number of cells and absorbance at 490 nm from each well.
Re-expression of miRNAs using lentivectors
Expression of miRNA precursors were driven by CMV promoters in a HIV-based lentiviral vector purchased from Systems Biosciences. The construct consists of the native stem loop structure of miRNA and 200 to 400 bp of upstream and downstream flanking genomic sequence cloned into the pMIRNA1-plasmid. Packaging of the miRNA constructs in pseudoviral particles was performed using the third-generation packaging system. NK-YS cells were infected with the lentivirus with an efficiency of ∼ 95% as determined by green fluorescent protein measurement by flow cytometry. Empty vector lentivirus was used as a control for the experiments.
Real-time RT-PCR quantification of miRNAs
Total RNAs, including small RNAs, were purified by miRNeasy Mini Kit (QIAGEN). cDNAs were synthesized from total RNA using TaqMan MicroRNA Reverse Transcription Kit with gene-specific primers. Reverse transcription reactions (for final quantity or concentrations) contained 10-ng RNA samples, 0.67μM of dNTP, 1× RT primer, 1× RT buffer, 3.8 U of RNAse inhibitor, and 50 U of reverse transcriptase. The 15-μL reactions were incubated for 30 minutes at 16°C, 30 minutes at 42°C, 5 minutes at 85°C, and then held at 4°C. Real-time RT-PCR quantification of miRNA expression was carried out using TaqManR MicroRNA Assays Kit (Applied Biosystems) according to the manufacturer's protocol. The 20-μL PCR included 1.33 μL RT product, 1× PCR Master mix, and 1× TaqMan-primers mix (Applied Biosystems). Reactions were incubated in a 96-well plate at 95°C for 10 minutes, followed by 40 cycles of 95°C for 15 seconds and 60°C for 1 minute. The threshold cycle (Ct) was determined using default threshold settings. All experiments were done in triplicates. The U6 snRNA was used as a control to normalize miRNA input in the real-time RT-PCR assay.
Real-time RT-PCR quantification of mRNAs
cDNAs from total RNA were obtained by the SuperScriptR III Reverse Transcriptase (Invitrogen) according to the manufacturer's instructions. SYBR PCR Master Mix (Applied Biosystems) was used for quantitative PCR as recommended by the manufacturer. GAPDH was used as a control to normalize mRNA input. All experiments were done in triplicates.
Luciferase reporter assay
A PCR-amplified fragment that contains 2 predicted miR-101 binding sites at the 3′-UTR of STMN1 was cloned into a dual-luciferase expression vector pmirGLO (Promega) to create the STMN1 reporter constructs. The seed and surrounding sequences at binding site #1 (position 278-292 of 3′-UTR, ATGGCTAGTACTGTA) and site #2 (position 437-448, CACAGTGCTGTT) within this construct were separately mutated to CTGGCTAATACGGTA and CGCAGCGCTCTC, respectively, using a Quick-change II site-directed mutagenesis kit (Agilent/Stratagene). The reporter construct containing full-length PRDM1 3′-UTR, the miR-101 and miR-186 precursor expression vectors, and their corresponding control plasmids were purchased from GeneCopoeia, System Biosciences, and Cell Biolabs, respectively. HEK-293T cells were cotransfected, in triplicate wells, with STMN1 reporter construct and miR-101 precursor expression vector at a ratio of 1:6 using Lipofectamine (Invitrogen) for 48 hours. PRDM1 reporter construct and miR-186 precursor vector were similarly transfected at a ratio of 1:200 for 72 hours before harvest. Firefly and Renilla luciferase activities of cell lysates were determined by a dual-luciferase reporter assay system (Promega). The ratio of firefly to Renilla luminescence of cells ectopically expressing miRNAs was compared with that of cells transfected with control miRNA precursor plasmid. miRNA overexpression was confirmed by real-time PCR using TaqMan probes specific to respective miRNAs. Results are presented as averages of 3 independent experiments.
IHC
Immunohistochemistry (IHC) was performed for MUM1, BLIMP1, and STMN1on 4-μm sections from the TMA blocks of NKTL cases. For those cases that were not included in the TMA, 4-μm sections were cut from whole paraffin blocks (5 cases). IHC was also performed on cell blocks of normal NK cells for comparison (see supplemental Table 3 for more details). Appropriate positive tissue controls were used. The immunohistochemical expression for all the antibodies was scored as a percentage of the total tumor cell population per 1-mm core diameter (× 400) by one of the authors (S.-B.N.), as previously described.3 For MUM1and BLIMP1 antibodies, positive expression was defined as nuclear staining in 20% or more of the tumor population. For STMN1, positive expression was defined as cytoplasmic and/or nuclear staining in 20% or more of the tumor population.
Results
miRNA dysregulation in NKTL
We compared the miRNA expression of NKTL FFPE samples (n = 30) with that of normal NK cells and the respective normal FFPE tissue controls from nasal, skin and soft tissue, intestinal tract, and lymph node, as well as that of NK cell lines with normal NK cells (supplemental Tables 4 and 5). Among the miRNAs showing at least 2-fold and statistically significant difference (P < .05) in expression, 2 were found to be up-regulated and 39 were down-regulated in both NK cell lines and FFPE NKTL samples compared with normal NK cells (Table 1). miR-342–5p, miR-26b, miR-363, miR-150, and miR28–5p are the top 5 down-regulated miRNAs, whereas miR-155 and miR-378 are up-regulated in both NK cell lines and FFPE NKTLs.
Deregulated miRNA in NKTL and NK cell lines compared with normal NK cells
| miRNA . | Cell lines versus normal . | NKTL versus normal . | Chromosome location . | Genomic position . | Target genes . | ||
|---|---|---|---|---|---|---|---|
| q-value . | Fold change . | q-value . | Fold change . | ||||
| hsa-miR-342–5p | 0.00047 | 0.17804 | 0.00000 | 0.17231 | 14 | 099645783-099645765 | POFUT1 |
| hsa-miR-26b | 0.00049 | 0.05876 | 0.00000 | 0.02942 | 2 | 218975644-218975625 | BCL2, IGF1, SETD7, FOXP2, CAPRIN1, PSD3, HOXA5, KPNA2, E2F7, ENPEP, EZH2, HMGA1, NAMPT, PIM1, SC4MOL, ACVR1C, AGPAT5, ASCC3, CKS2, CTTNBP2NL, DCDC2, IARS, KIF18A, LARP1, MTM1, NFE2L3, NUP50, SLC7A11 |
| hsa-miR-363 | 0.00049 | 0.05325 | 0.00000 | 0.06310 | X | 133131078-133131095 | BCL2, IGF1, SETD7, FOXP2, PSD3, ATP2A2, BCAT2, DOCK9, SMAD6, ADCY3, ASB7, CHCHD10, FMN2, NFIB, RAB23, RGL1, SLC7A11, YIPF4 |
| hsa-miR-150 | 0.00049 | 0.00483 | 0.00000 | 0.01099 | 19 | 054695901-054695916 | MYB, ELK1, CTH, ENSA |
| hsa-miR-28–5p | 0.00049 | 0.17303 | 0.00000 | 0.14000 | 3 | 189889297-189889278 | IGF1, SETD7, CAPRIN1, HTRA2, MAD2L1, TLN2 |
| hsa-miR-152 | 0.00049 | 0.28067 | 0.00000 | 0.27612 | 17 | 043469539-043469553 | E2F7, ANK2, ATP2A2, B4GALT2, BBC3, CEP55, DPP3, EMP1, HMGB3, IGF1, KLC2 |
| hsa-miR-361–3p | 0.00049 | 0.23188 | 0.00000 | 0.19701 | X | 085045302-085045320 | CD3EAP, OSR2 |
| hsa-miR-22* | 0.00049 | 0.32882 | 0.00000 | 0.34642 | 17 | 001563996-001564012 | No predicted targets |
| hsa-miR-340 | 0.00049 | 0.26426 | 0.00000 | 0.29925 | 5 | 179374967-179374984 | AGPAT5, AHR, CDON, CIT, DEPDC1B, E2F7, FHL2, GK, HECW2, IGF1, ING3, MYO1C, NUPL1, PHLDA1, SLC7A11, TIAM1 |
| hsa-miR-598 | 0.00049 | 0.49291 | 0.00000 | 0.48708 | 8 | 010930141-010930158 | No predicted targets |
| hsa-miR-181a-2* | 0.00049 | 0.23242 | 0.00000 | 0.22641 | 9 | 126494639-126494623 | No predicted targets |
| hsa-miR-132 | 0.00050 | 0.45398 | 0.00000 | 0.42973 | 17 | 001899973-001899987 | HBEGF, BRI3, HN1, TLN2, VDAC2, ADCY3, AHCY, AZIN1, CAPRIN1, FKBP2, NFIB, PPM1G, SCN2A, TRIB1, TTK |
| hsa-miR-194 | 0.00050 | 0.32209 | 0.00024 | 0.32691 | 1 | 218358171-218358185 | HBEGF, TLN2, CTAGE5, LPHN2, PRR7, VDAC2 |
| hsa-miR-768–3p | 0.00050 | 0.13168 | 0.00000 | 0.08066 | 16 | 070349814-070349832 | AHR, CENPE, HOXA4 |
| hsa-miR-873 | 0.00050 | 0.22803 | 0.00000 | 0.22284 | 9 | 028878923-028878939 | FOXK2, MPDU1, TLN2 |
| hsa-miR-338–3p | 0.00052 | 0.13277 | 0.00001 | 0.15864 | 17 | 076714282-076714301 | FKBP1A, ARPC1B |
| hsa-miR-215 | 0.00053 | 0.44650 | 0.00119 | 0.45644 | 1 | 218357881-218357900 | DYRK3, LPAR4, TRIP13 |
| hsa-miR-186 | 0.00054 | 0.26956 | 0.00000 | 0.14583 | 1 | 071305952-071305971 | CDC42, BTF3, PPM1G, SMAD6, ACSL4, BCAT1, BMP2K, EIF2S2, ENPEP, PRDM1, PSD3, PSMD11, PSPH, RGS22, VEGFA, ZCCHC5 |
| hsa-miR-140–3p | 0.00054 | 0.27234 | 0.00000 | 0.16194 | 16 | 068524566-068524552 | FOXK2, UBE2C |
| hsa-miR-140–5p | 0.00054 | 0.30678 | 0.00000 | 0.13482 | 16 | 068524528-068524508 | ARHGAP19, CASP3, ST5, TTK |
| hsa-miR-374b | 0.00055 | 0.32813 | 0.00000 | 0.13347 | X | 073355147-073355164 | TFDP1, CCNE2, EIF2S2, EIF4G1, ENSA, GNB2, HOXA11, HSPA4, HTRA2, LARP1, NFIB, SMAD6 |
| hsa-miR-26a | 0.00056 | 0.14699 | 0.00048 | 0.23283 | 12 | 056504708-056504721 | BCL2, IGF1, SETD7, PSD3, EZH2, HOXA5, KPNA2, E2F7, ENPEP, HMGA1, NAMPT, PIM1, SC4MOL, ASCC3, CKS2, CTTNBP2NL, DCDC2, IARS, KIF18A, LARP1, MTM1, NFE2L3, NUP50, SLC7A11 |
| hsa-let-7g | 0.00057 | 0.21467 | 0.00001 | 0.10030 | 3 | 052277392-052277411 | ACVR1C, AP1S1, CASP3, CDC25A, COL15A1, CYP19A1, DPP3, EZH2, FAM118A, POLR3D, SCD, TARBP2, TTLL4, ATP2A2, BCAP29, BCAT1, CCNF, CD86, DUSP4, HMGA1, RGS16, SOCS1, THRSP, ZCCHC5 |
| hsa-miR-342–3p | 0.00057 | 0.14972 | 0.00000 | 0.07856 | 14 | 099645827-099645812 | ENSA, SLC35F2, TIAM1 |
| hsa-miR-101 | 0.00057 | 0.33272 | 0.00000 | 0.09962 | 1 | 065296713-065296731 | STMN1, BCL2, IGF1, PSD3, EZH2, EMP1, ING3, PANK3, PHLDA1, TRIB1, ACCN2, ASCC3, DDIT4, HNRNPAB, LMNB1, POMP, SCN2A, SELI |
| hsa-miR-192 | 0.00060 | 0.33056 | 0.00044 | 0.32841 | 11 | 064415251-064415268 | DYRK3, LPAR4, TRIP13 |
| hsa-miR-374a | 0.00065 | 0.29720 | 0.00000 | 0.09324 | X | 073423885-073423905 | EIF2S2, ENSA, GK, GNB2, HOXA11, HSPA4, LARP1, NFIB, SMAD6, TFDP1 |
| hsa-miR-876–5p | 0.00070 | 0.43160 | 0.00004 | 0.39927 | 9 | 028853673-028853691 | EME1, FOXM1, DNAJC12, NTRK2, PHLDA1, TFAP2A, ZCCHC5 |
| hsa-miR-22 | 0.00076 | 0.19107 | 0.00000 | 0.19752 | 17 | 001563958-001563975 | BATF3, DDIT4, HOXA4, IPO7, MTHFD2, RFXANK, APBB2, NET1, PPM1G, TIAM1 |
| hsa-miR-10a | 0.00111 | 0.42780 | 0.00008 | 0.36971 | 17 | 044012265-044012284 | BCL6, SOBP, STK24, TIAM1, ZNF367 |
| hsa-miR-590–5p | 0.00168 | 0.45785 | 0.00000 | 0.17107 | 7 | 073243500-073243479 | ING3, NFIB, RBPJ, TIAM1, ZNF367 |
| hsa-miR-30b | 0.00258 | 0.26088 | 0.00000 | 0.09993 | 8 | 135881995-135882010 | ADAM22, DDIT4, NFIB, PPARGC1B, SCN2A, SLC41A2, ASCC3, CCNE2, CELSR3, FGD6, PRDM1, RHEBL1, SCN8A, SMARCD2, SOCS1, AVEN, AZIN1, BCL2, DEPDC4, FRMD6, HOXA11, IL2RA, IRF4, ITSN1, MTA1, PRICKLE1, RGL1, SETD7, STXBP1, SUPT3H, TFDP1 |
| hsa-miR-181c | 0.00324 | 0.40588 | 0.00000 | 0.24845 | 19 | 013846560-013846542 | FKBP1A, NR6A1, CTTNBP2NL, DDIT4, E2F7, HOXA11, NR4A3, APOO, ATP2A2, CDON, FAM3C, IPPK, ITSN1, MAP1A, MINA, NLN, PDIA6, PHLDA1, PRDX3, SCD, SLC25A37 |
| hsa-miR-142–5p | 0.00404 | 0.34288 | 0.00000 | 0.01692 | 17 | 053763643-053763660 | SRI, AHR, HN1, PPM1G, RBBP8, RNH1, SLC41A2 |
| hsa-let-7a | 0.00602 | 0.45269 | 0.00420 | 0.37315 | 11 | 121522486-121522504 | ACVR1C, AP1S1, CASP3, CDC25A, CYP19A1, DPP3, EZH2, SCD, TARBP2, ATP2A2, BCAP29, BCAT1, BRF2, CCNF, COL15A1, DUSP4, FAM118A, HOXB4, POLR3D, RGS16, SOCS1, THRSP, TRIB1, TTLL4, ZCCHC5 |
| hsa-miR-155 | 0.00690 | 11.45284 | 0.00119 | 2.16728 | 21 | 025868188-025868172 | BNC2, SGK3, TLE4, TSHZ3, EIF2C4, FGF7, GPM6B, KLRC3, LHX9, MYLK, PCDH9, PDLIM5, RAB34, RREB1, SOX11, ZNF618 |
| hsa-let-7c | 0.01070 | 0.27938 | 0.00160 | 0.41876 | 21 | 016834050-016834032 | ACVR1C, CASP3, CDC25A, CYP19A1, DPP3, EZH2, SCD, TARBP2, TTLL4, AP1S1, ATP2A2, BCAT1, BRF2, CCNF, COL15A1, DUSP4, FAM118A, HOXB4, POLR3D, RGS16, SOCS1, THRSP, TRIB1 |
| hsa-miR-378 | 0.01145 | 2.53182 | 0.00059 | 4.08760 | 5 | 149092643-149092631 | GPM6B, IGF1R, WDR37 |
| hsa-miR-181a | 0.01665 | 0.38679 | 0.00000 | 0.16954 | 1 | 197094860-197094873 | FKBP1A, NR6A1, CTTNBP2NL, DDIT4, E2F7, HOXA11, NR4A3, APOO, ATP2A2, CDON, FAM3C, IPPK, ITSN1, MAP1A, MINA, NLN, PDIA6, PHLDA1, PLAU, PRDX3, SCD, SLC25A37 |
| hsa-miR-142–3p | 0.01796 | 0.45692 | 0.00000 | 0.01686 | 17 | 053763605-053763626 | TFG, FKBP1A, GNB2, ATP2A2 |
| hsa-miR-15a | 0.04502 | 0.49695 | 0.00000 | 0.13992 | 13 | 049521304-049521323 | CCNE1, CDCA4, CHEK1, MYB, WEE1, CDC25A, KIF23, LPHN2, PDIA6, PPAP2A, SMARCD2, STXBP1, TARBP2, ACSL4, ANKRD13B, BCL2, BTF3, CDC42, E2F7, FKBP1A, FSD1, IARS, LIPE, OTX1, PANK1, PHF19, PIM1, PPIF, PPIL1, PTPN3, SELI, SMYD5, ZCCHC5 |
| miRNA . | Cell lines versus normal . | NKTL versus normal . | Chromosome location . | Genomic position . | Target genes . | ||
|---|---|---|---|---|---|---|---|
| q-value . | Fold change . | q-value . | Fold change . | ||||
| hsa-miR-342–5p | 0.00047 | 0.17804 | 0.00000 | 0.17231 | 14 | 099645783-099645765 | POFUT1 |
| hsa-miR-26b | 0.00049 | 0.05876 | 0.00000 | 0.02942 | 2 | 218975644-218975625 | BCL2, IGF1, SETD7, FOXP2, CAPRIN1, PSD3, HOXA5, KPNA2, E2F7, ENPEP, EZH2, HMGA1, NAMPT, PIM1, SC4MOL, ACVR1C, AGPAT5, ASCC3, CKS2, CTTNBP2NL, DCDC2, IARS, KIF18A, LARP1, MTM1, NFE2L3, NUP50, SLC7A11 |
| hsa-miR-363 | 0.00049 | 0.05325 | 0.00000 | 0.06310 | X | 133131078-133131095 | BCL2, IGF1, SETD7, FOXP2, PSD3, ATP2A2, BCAT2, DOCK9, SMAD6, ADCY3, ASB7, CHCHD10, FMN2, NFIB, RAB23, RGL1, SLC7A11, YIPF4 |
| hsa-miR-150 | 0.00049 | 0.00483 | 0.00000 | 0.01099 | 19 | 054695901-054695916 | MYB, ELK1, CTH, ENSA |
| hsa-miR-28–5p | 0.00049 | 0.17303 | 0.00000 | 0.14000 | 3 | 189889297-189889278 | IGF1, SETD7, CAPRIN1, HTRA2, MAD2L1, TLN2 |
| hsa-miR-152 | 0.00049 | 0.28067 | 0.00000 | 0.27612 | 17 | 043469539-043469553 | E2F7, ANK2, ATP2A2, B4GALT2, BBC3, CEP55, DPP3, EMP1, HMGB3, IGF1, KLC2 |
| hsa-miR-361–3p | 0.00049 | 0.23188 | 0.00000 | 0.19701 | X | 085045302-085045320 | CD3EAP, OSR2 |
| hsa-miR-22* | 0.00049 | 0.32882 | 0.00000 | 0.34642 | 17 | 001563996-001564012 | No predicted targets |
| hsa-miR-340 | 0.00049 | 0.26426 | 0.00000 | 0.29925 | 5 | 179374967-179374984 | AGPAT5, AHR, CDON, CIT, DEPDC1B, E2F7, FHL2, GK, HECW2, IGF1, ING3, MYO1C, NUPL1, PHLDA1, SLC7A11, TIAM1 |
| hsa-miR-598 | 0.00049 | 0.49291 | 0.00000 | 0.48708 | 8 | 010930141-010930158 | No predicted targets |
| hsa-miR-181a-2* | 0.00049 | 0.23242 | 0.00000 | 0.22641 | 9 | 126494639-126494623 | No predicted targets |
| hsa-miR-132 | 0.00050 | 0.45398 | 0.00000 | 0.42973 | 17 | 001899973-001899987 | HBEGF, BRI3, HN1, TLN2, VDAC2, ADCY3, AHCY, AZIN1, CAPRIN1, FKBP2, NFIB, PPM1G, SCN2A, TRIB1, TTK |
| hsa-miR-194 | 0.00050 | 0.32209 | 0.00024 | 0.32691 | 1 | 218358171-218358185 | HBEGF, TLN2, CTAGE5, LPHN2, PRR7, VDAC2 |
| hsa-miR-768–3p | 0.00050 | 0.13168 | 0.00000 | 0.08066 | 16 | 070349814-070349832 | AHR, CENPE, HOXA4 |
| hsa-miR-873 | 0.00050 | 0.22803 | 0.00000 | 0.22284 | 9 | 028878923-028878939 | FOXK2, MPDU1, TLN2 |
| hsa-miR-338–3p | 0.00052 | 0.13277 | 0.00001 | 0.15864 | 17 | 076714282-076714301 | FKBP1A, ARPC1B |
| hsa-miR-215 | 0.00053 | 0.44650 | 0.00119 | 0.45644 | 1 | 218357881-218357900 | DYRK3, LPAR4, TRIP13 |
| hsa-miR-186 | 0.00054 | 0.26956 | 0.00000 | 0.14583 | 1 | 071305952-071305971 | CDC42, BTF3, PPM1G, SMAD6, ACSL4, BCAT1, BMP2K, EIF2S2, ENPEP, PRDM1, PSD3, PSMD11, PSPH, RGS22, VEGFA, ZCCHC5 |
| hsa-miR-140–3p | 0.00054 | 0.27234 | 0.00000 | 0.16194 | 16 | 068524566-068524552 | FOXK2, UBE2C |
| hsa-miR-140–5p | 0.00054 | 0.30678 | 0.00000 | 0.13482 | 16 | 068524528-068524508 | ARHGAP19, CASP3, ST5, TTK |
| hsa-miR-374b | 0.00055 | 0.32813 | 0.00000 | 0.13347 | X | 073355147-073355164 | TFDP1, CCNE2, EIF2S2, EIF4G1, ENSA, GNB2, HOXA11, HSPA4, HTRA2, LARP1, NFIB, SMAD6 |
| hsa-miR-26a | 0.00056 | 0.14699 | 0.00048 | 0.23283 | 12 | 056504708-056504721 | BCL2, IGF1, SETD7, PSD3, EZH2, HOXA5, KPNA2, E2F7, ENPEP, HMGA1, NAMPT, PIM1, SC4MOL, ASCC3, CKS2, CTTNBP2NL, DCDC2, IARS, KIF18A, LARP1, MTM1, NFE2L3, NUP50, SLC7A11 |
| hsa-let-7g | 0.00057 | 0.21467 | 0.00001 | 0.10030 | 3 | 052277392-052277411 | ACVR1C, AP1S1, CASP3, CDC25A, COL15A1, CYP19A1, DPP3, EZH2, FAM118A, POLR3D, SCD, TARBP2, TTLL4, ATP2A2, BCAP29, BCAT1, CCNF, CD86, DUSP4, HMGA1, RGS16, SOCS1, THRSP, ZCCHC5 |
| hsa-miR-342–3p | 0.00057 | 0.14972 | 0.00000 | 0.07856 | 14 | 099645827-099645812 | ENSA, SLC35F2, TIAM1 |
| hsa-miR-101 | 0.00057 | 0.33272 | 0.00000 | 0.09962 | 1 | 065296713-065296731 | STMN1, BCL2, IGF1, PSD3, EZH2, EMP1, ING3, PANK3, PHLDA1, TRIB1, ACCN2, ASCC3, DDIT4, HNRNPAB, LMNB1, POMP, SCN2A, SELI |
| hsa-miR-192 | 0.00060 | 0.33056 | 0.00044 | 0.32841 | 11 | 064415251-064415268 | DYRK3, LPAR4, TRIP13 |
| hsa-miR-374a | 0.00065 | 0.29720 | 0.00000 | 0.09324 | X | 073423885-073423905 | EIF2S2, ENSA, GK, GNB2, HOXA11, HSPA4, LARP1, NFIB, SMAD6, TFDP1 |
| hsa-miR-876–5p | 0.00070 | 0.43160 | 0.00004 | 0.39927 | 9 | 028853673-028853691 | EME1, FOXM1, DNAJC12, NTRK2, PHLDA1, TFAP2A, ZCCHC5 |
| hsa-miR-22 | 0.00076 | 0.19107 | 0.00000 | 0.19752 | 17 | 001563958-001563975 | BATF3, DDIT4, HOXA4, IPO7, MTHFD2, RFXANK, APBB2, NET1, PPM1G, TIAM1 |
| hsa-miR-10a | 0.00111 | 0.42780 | 0.00008 | 0.36971 | 17 | 044012265-044012284 | BCL6, SOBP, STK24, TIAM1, ZNF367 |
| hsa-miR-590–5p | 0.00168 | 0.45785 | 0.00000 | 0.17107 | 7 | 073243500-073243479 | ING3, NFIB, RBPJ, TIAM1, ZNF367 |
| hsa-miR-30b | 0.00258 | 0.26088 | 0.00000 | 0.09993 | 8 | 135881995-135882010 | ADAM22, DDIT4, NFIB, PPARGC1B, SCN2A, SLC41A2, ASCC3, CCNE2, CELSR3, FGD6, PRDM1, RHEBL1, SCN8A, SMARCD2, SOCS1, AVEN, AZIN1, BCL2, DEPDC4, FRMD6, HOXA11, IL2RA, IRF4, ITSN1, MTA1, PRICKLE1, RGL1, SETD7, STXBP1, SUPT3H, TFDP1 |
| hsa-miR-181c | 0.00324 | 0.40588 | 0.00000 | 0.24845 | 19 | 013846560-013846542 | FKBP1A, NR6A1, CTTNBP2NL, DDIT4, E2F7, HOXA11, NR4A3, APOO, ATP2A2, CDON, FAM3C, IPPK, ITSN1, MAP1A, MINA, NLN, PDIA6, PHLDA1, PRDX3, SCD, SLC25A37 |
| hsa-miR-142–5p | 0.00404 | 0.34288 | 0.00000 | 0.01692 | 17 | 053763643-053763660 | SRI, AHR, HN1, PPM1G, RBBP8, RNH1, SLC41A2 |
| hsa-let-7a | 0.00602 | 0.45269 | 0.00420 | 0.37315 | 11 | 121522486-121522504 | ACVR1C, AP1S1, CASP3, CDC25A, CYP19A1, DPP3, EZH2, SCD, TARBP2, ATP2A2, BCAP29, BCAT1, BRF2, CCNF, COL15A1, DUSP4, FAM118A, HOXB4, POLR3D, RGS16, SOCS1, THRSP, TRIB1, TTLL4, ZCCHC5 |
| hsa-miR-155 | 0.00690 | 11.45284 | 0.00119 | 2.16728 | 21 | 025868188-025868172 | BNC2, SGK3, TLE4, TSHZ3, EIF2C4, FGF7, GPM6B, KLRC3, LHX9, MYLK, PCDH9, PDLIM5, RAB34, RREB1, SOX11, ZNF618 |
| hsa-let-7c | 0.01070 | 0.27938 | 0.00160 | 0.41876 | 21 | 016834050-016834032 | ACVR1C, CASP3, CDC25A, CYP19A1, DPP3, EZH2, SCD, TARBP2, TTLL4, AP1S1, ATP2A2, BCAT1, BRF2, CCNF, COL15A1, DUSP4, FAM118A, HOXB4, POLR3D, RGS16, SOCS1, THRSP, TRIB1 |
| hsa-miR-378 | 0.01145 | 2.53182 | 0.00059 | 4.08760 | 5 | 149092643-149092631 | GPM6B, IGF1R, WDR37 |
| hsa-miR-181a | 0.01665 | 0.38679 | 0.00000 | 0.16954 | 1 | 197094860-197094873 | FKBP1A, NR6A1, CTTNBP2NL, DDIT4, E2F7, HOXA11, NR4A3, APOO, ATP2A2, CDON, FAM3C, IPPK, ITSN1, MAP1A, MINA, NLN, PDIA6, PHLDA1, PLAU, PRDX3, SCD, SLC25A37 |
| hsa-miR-142–3p | 0.01796 | 0.45692 | 0.00000 | 0.01686 | 17 | 053763605-053763626 | TFG, FKBP1A, GNB2, ATP2A2 |
| hsa-miR-15a | 0.04502 | 0.49695 | 0.00000 | 0.13992 | 13 | 049521304-049521323 | CCNE1, CDCA4, CHEK1, MYB, WEE1, CDC25A, KIF23, LPHN2, PDIA6, PPAP2A, SMARCD2, STXBP1, TARBP2, ACSL4, ANKRD13B, BCL2, BTF3, CDC42, E2F7, FKBP1A, FSD1, IARS, LIPE, OTX1, PANK1, PHF19, PIM1, PPIF, PPIL1, PTPN3, SELI, SMYD5, ZCCHC5 |
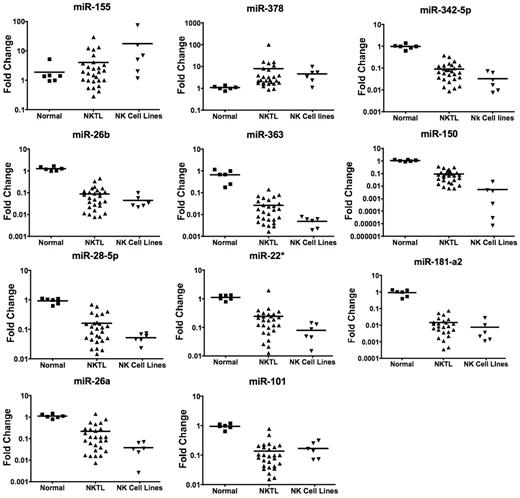
We performed quantitative PCR validation of 11 selected miRNAs, including the top 5 down-regulated miRNAs, 2 up-regulated miRNAs, and a few interesting miRNAs, which may be involved in tumor oncogenesis. On the whole, quantitative PCR results were consistent with MEP data showing overexpression of miR-155 and miR-378 and underexpression of miR-342–5p, miR-26b, miR-363, miR-150 and miR28–5p, miR-22*, miR-181a-2*, miR-26a and miR-101 in NK cell lines and NKTL FFPE samples compared with normal NK cells (Figure 1).
Quantitative RT-PCR validation of miRNA expression profiling data. Eleven miRNAs that were deregulated in NKTL were selected for validation by quantitative RT-PCR. In every case, miRNAs down-regulated in NKTL compared with normal NK cells were also found to be down-regulated by quantitative RT-PCR. Similar observations were made for up-regulated miRNAs. All comparisons are statistically significant (P < .05).
Quantitative RT-PCR validation of miRNA expression profiling data. Eleven miRNAs that were deregulated in NKTL were selected for validation by quantitative RT-PCR. In every case, miRNAs down-regulated in NKTL compared with normal NK cells were also found to be down-regulated by quantitative RT-PCR. Similar observations were made for up-regulated miRNAs. All comparisons are statistically significant (P < .05).
The validity of the MEP platform and results was further verified by comparing the miRNA expressed in our normal and stimulated NK cells with that detected by sequencing methods.15 There is substantial overlap between our data list and the list generated by sequencing method (supplemental Figure 1).
Functional relevance of dysregulated miRNAs in NKTL
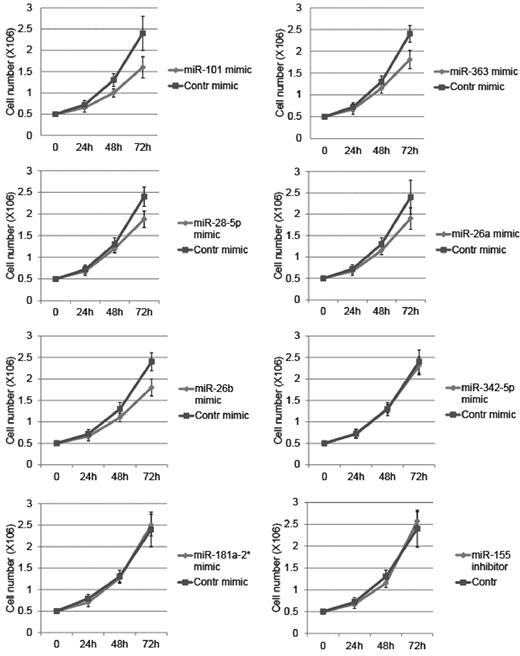
To assess the relevance of the dysregulated miRNAs to the biology of NKTL, we incubated the NKYS cell line with miRNA mimics for down-regulated miRNAs, including miR-101, miR-363, miR-28–5p, miR-26a, miR-26b, miR-342–5p, and miR-181a-2*, and miRNA inhibitor for one of the overexpressed miRNAs, miR-155. The use of miR-101, miR-363, miR-28–5p, miR-26a, and miR-26b mimics substantially reduced growth of NK-YS cells (Figure 2). This suggests that these miRNAs could play a potential role in the growth and proliferation of NKTL.
Effect of deregulated miRNAs on growth of NKTL cells. Growth curves of NK-YS cells transfected with synthetic miRNAs and anti-miRNA inhibitors. Cell number was counted at the indicated time points after transfections. Counting results were validated by CellTiter 96 Cell Proliferation Assay. Error bars represent SD; n = 3.
Effect of deregulated miRNAs on growth of NKTL cells. Growth curves of NK-YS cells transfected with synthetic miRNAs and anti-miRNA inhibitors. Cell number was counted at the indicated time points after transfections. Counting results were validated by CellTiter 96 Cell Proliferation Assay. Error bars represent SD; n = 3.
Next, we identified high-probability predicted target genes of these deregulated miRNAs by intersecting targets predicted by 6 algorithms, including mirBase (http://microrna.sanger.ac.uk), targetScan (http://www.targetscan.org), miRanda (http://www.microrna.org), tarBase (http://diana.cslab.ece.ntua.gr/tarbase), mirtarget2 (http://mirdb.org/miRDB), and pictar (http://pictar.mdc-berlin.de; Table 1). We further assessed the expression of the target genes in those samples that also have GEP done and further narrowed down the relevant target genes to those whose expression is inversely correlated with the expression of the deregulated miRNAs. We selected a number of target genes of the 3 miRNAs (miR-101, miR-26a, and miR-26b), which were shown to alter the growth of NK-YS for further validation. We used lentiviral vectors as an alternate method to express miR-101, miR-26a, and miR-26b in NK-YS. This resulted in a significant increase in the expression of these miRNAs and a corresponding decrease in the expression of STMN1 (Figure 3A), one of the target genes of miR-101, and BCL2, a target gene shared by miR-101, miR-26a, and miR-26b (Figure 3C). On the other hand, IGF1 is only down-regulated on mir-101 expression but not miR-26a or miR-26b expression, although it is also predicted to be targets of all 3 miRNAs (Figure 3C). This inconsistency may be explained by the known discrepancies between predicted target and actual targets. We therefore proceeded to validate several miRNAs and their predicted targets, which may be of relevance in NKTL. In the 3′-UTR of STMN1, there are 2 predicted binding sites of miR-101. To confirm that miR-101 binds to 3′-UTR of STMN1 and affects its expression, and to clarify which of these binding sites are the most important, we performed luciferase assay with different STMN1 3′-UTR constructs; no mutation, first binding sequence mutated (MUT1) or second binding sequence mutated (MUT2) (supplemental Figure 2). Our results showed that the reporter with the fragment of 3′-UTR of STMN1 that contains the 2 seed sequences reduced luciferase activity. The MUT1 reporter construct (with the first seed sequence mutated) showed similar level of luciferase activity as the wild-type reporter construct, whereas the MUT2 reporter construct (with the second seed sequence mutated) demonstrated a restoration to the same level as the empty control reporter. This suggests that the second seed sequence is the critical binding site for miR-101.
Effect of deregulated miRNAs on expression of their target genes. (A) miR-101 represses STMN1. Top: Reintroduction of miR-101 into NKYS cells reduced mRNA levels of STMN1. mRNA levels of STMN1 were determined by quantitative RT-PCR analysis. Cells were transduced by lentivirus produced by miRNA precursor expression vectors or a control vector. Cells were harvested 4 days after transduction. Mature miR-101 transcript was determined by TaqMan miRNA assay. Bottom: 3′-UTR luciferase reporter assay of STMN1. The 293T cells were cotransfected with a reporter construct with or without 3′-UTR of STMN1 that contain 2 potential miR-101 binding sites and an miRNA expression vector with or without hsa-miR-101 precursor sequence. Reporter constructs with point mutations to the seed sequence of either miR-101 target site were similarly cotransfected. Luciferase activity was determined 48 hours after transfection. (B) miR-30b represses PRDM1. Top: Reintroduction of miR-30b into NK-YS cells reduced mRNA levels of PRDM1. Cells were transiently transfected by miRNA-30b mimics. Expression of PRDM1 at 48 hours after transfection was determined by quantitative RT-PCR analysis. Bottom: 3′-UTR luciferase reporter assay of PRDM1. The 293T cells were cotransfected with a reporter construct with or without the whole 3′-UTR of PRDM1 cloned to the distal end of the firefly luciferase gene and an miRNA expression vector with or without the hsa-miR-30b precursor sequence. Luciferase activity was determined 48 hours after transfection. (C) Re-expression of miR-101, miR-26a, or miR-26b in NK-YS cells reduced expression of BCL-2. Cells were transduced by lentivirus produced by miRNA precursor expression vectors or a control vector and were harvested 4 days after transduction for quantitative RT-PCR analysis.
Effect of deregulated miRNAs on expression of their target genes. (A) miR-101 represses STMN1. Top: Reintroduction of miR-101 into NKYS cells reduced mRNA levels of STMN1. mRNA levels of STMN1 were determined by quantitative RT-PCR analysis. Cells were transduced by lentivirus produced by miRNA precursor expression vectors or a control vector. Cells were harvested 4 days after transduction. Mature miR-101 transcript was determined by TaqMan miRNA assay. Bottom: 3′-UTR luciferase reporter assay of STMN1. The 293T cells were cotransfected with a reporter construct with or without 3′-UTR of STMN1 that contain 2 potential miR-101 binding sites and an miRNA expression vector with or without hsa-miR-101 precursor sequence. Reporter constructs with point mutations to the seed sequence of either miR-101 target site were similarly cotransfected. Luciferase activity was determined 48 hours after transfection. (B) miR-30b represses PRDM1. Top: Reintroduction of miR-30b into NK-YS cells reduced mRNA levels of PRDM1. Cells were transiently transfected by miRNA-30b mimics. Expression of PRDM1 at 48 hours after transfection was determined by quantitative RT-PCR analysis. Bottom: 3′-UTR luciferase reporter assay of PRDM1. The 293T cells were cotransfected with a reporter construct with or without the whole 3′-UTR of PRDM1 cloned to the distal end of the firefly luciferase gene and an miRNA expression vector with or without the hsa-miR-30b precursor sequence. Luciferase activity was determined 48 hours after transfection. (C) Re-expression of miR-101, miR-26a, or miR-26b in NK-YS cells reduced expression of BCL-2. Cells were transduced by lentivirus produced by miRNA precursor expression vectors or a control vector and were harvested 4 days after transduction for quantitative RT-PCR analysis.
Next, we validated the relationship between miR-30b (underexpressed in NKTL) and PRDM1 (gene encoding BLIMP1), which has been shown to be important for NK cell maturation and may therefore be of biologic relevance to NKTL. The use of miR-30b mimic in NKYS leads to repression of PRDM1 mRNA expression. The effect of miR-30b on PRDM1 expression is confirmed on luciferase reporter assay when expression of miR-30b precursor inhibited PRDM1 expression (Figure 3B). These results provide further evidence that deregulation of miRNA is of functional and biologic relevance in NKTL.
Downstream pathways affected by dysregulated miRNA
Although previously thought to mediate mainly the inhibition of protein translation, recent studies suggest that miRNAs also extensively down-regulate mRNA expression through mRNA decay.16 We therefore correlated the expression of predicted target genes using data from our previous GEP study3 with the expression of the dysregulated miRNA and identified a total of 226 target genes whose gene expressions were inversely correlated with the expression of the 41 deregulated miRNA (supplemental Figure 3). It is apparent that some miRNAs, such as miR-30b, miR-15a, let-7a, let-7c, and let-7a, regulate multiple target genes, whereas others have only one specific target gene. Conversely, some target genes (eg, E2F7 and EZH2) are regulated by multiple miRNAs, whereas others are regulated by a single miRNA.
There is a significant enrichment among these predicted target genes for genes involved in cell cycle-related pathways, MAPK and p53 signaling pathways (Table 2). This is consistent with our previous findings showing increased expression of cell cycle-related genes and activation of p53 pathway in NKTL.3
Enriched KEGG pathways among predicted gene targets of deregulated miRNAs in NKTL
| KEGGID . | P . | Odds ratio . | Count . | Size . | Term . | Genes . |
|---|---|---|---|---|---|---|
| 770 | .00005 | 22.96432681 | 4 | 15 | Pantothenate and CoA biosynthesis | BCAT1, BCAT2, PANK1, PANK3 |
| 290 | .00044 | 23.4 | 3 | 11 | Valine, leucine, and isoleucine biosynthesis | BCAT1, BCAT2, IARS |
| 4115 | .00055 | 6.206116464 | 6 | 68 | p53 signaling pathway | CASP3, CCNE1, CHEK1, IGF1, CCNE2, BBC3 |
| 4110 | .00057 | 4.531014493 | 8 | 123 | Cell cycle | CCNE1, CDC25A, CHEK1, MAD2L1, TFDP1, TTK, WEE1, CCNE2 |
| 4114 | .00155 | 4.293859649 | 7 | 112 | Oocyte meiosis | ADCY3, ADCY8, CCNE1, IGF1, IGF1R, MAD2L1, CCNE2 |
| 4914 | .00175 | 4.853855006 | 6 | 85 | Progesterone-mediated oocyte maturation | ADCY3, ADCY8, CDC25A, IGF1, IGF1R, MAD2L1 |
| 5215 | .00945 | 3.844590369 | 5 | 87 | Prostate cancer | BCL2, CCNE1, IGF1, IGF1R, CCNE2 |
| 4510 | .00994 | 2.730212766 | 8 | 196 | Focal adhesion | BCL2, CDC42, ELK1, IGF1, IGF1R, MYLK, VEGFA, TLN2 |
| 4912 | .01513 | 3.38227185 | 5 | 98 | GnRH signaling pathway | ADCY3, ADCY8, CDC42, HBEGF, ELK1 |
| 4614 | .02164 | 8.205761317 | 2 | 17 | Renin-angiotensin system | ENPEP, NLN |
| 5210 | .03373 | 3.113924051 | 4 | 84 | Colorectal cancer | BCL2, CASP3, IGF1R, ACVR1C |
| 4150 | .03533 | 3.868 75 | 3 | 51 | mTOR signaling pathway | IGF1, VEGFA, DDIT4 |
| 5414 | .03884 | 2.963230862 | 4 | 88 | Dilated cardiomyopathy | ADCY3, ADCY8, ATP2A2, IGF1 |
| 4010 | .04517 | 1.984836601 | 8 | 263 | MAPK signaling pathway | CASP3, CDC42, DUSP4, ELK1, FGF7, STMN1, NTRK2, ACVR1C |
| 52 | .04708 | 5.119341564 | 2 | 26 | Galactose metabolism | GCK, B4GALT2 |
| 450 | .04708 | 5.119341564 | 2 | 26 | Seleno amino acid metabolism | AHCY, CTH |
| KEGGID . | P . | Odds ratio . | Count . | Size . | Term . | Genes . |
|---|---|---|---|---|---|---|
| 770 | .00005 | 22.96432681 | 4 | 15 | Pantothenate and CoA biosynthesis | BCAT1, BCAT2, PANK1, PANK3 |
| 290 | .00044 | 23.4 | 3 | 11 | Valine, leucine, and isoleucine biosynthesis | BCAT1, BCAT2, IARS |
| 4115 | .00055 | 6.206116464 | 6 | 68 | p53 signaling pathway | CASP3, CCNE1, CHEK1, IGF1, CCNE2, BBC3 |
| 4110 | .00057 | 4.531014493 | 8 | 123 | Cell cycle | CCNE1, CDC25A, CHEK1, MAD2L1, TFDP1, TTK, WEE1, CCNE2 |
| 4114 | .00155 | 4.293859649 | 7 | 112 | Oocyte meiosis | ADCY3, ADCY8, CCNE1, IGF1, IGF1R, MAD2L1, CCNE2 |
| 4914 | .00175 | 4.853855006 | 6 | 85 | Progesterone-mediated oocyte maturation | ADCY3, ADCY8, CDC25A, IGF1, IGF1R, MAD2L1 |
| 5215 | .00945 | 3.844590369 | 5 | 87 | Prostate cancer | BCL2, CCNE1, IGF1, IGF1R, CCNE2 |
| 4510 | .00994 | 2.730212766 | 8 | 196 | Focal adhesion | BCL2, CDC42, ELK1, IGF1, IGF1R, MYLK, VEGFA, TLN2 |
| 4912 | .01513 | 3.38227185 | 5 | 98 | GnRH signaling pathway | ADCY3, ADCY8, CDC42, HBEGF, ELK1 |
| 4614 | .02164 | 8.205761317 | 2 | 17 | Renin-angiotensin system | ENPEP, NLN |
| 5210 | .03373 | 3.113924051 | 4 | 84 | Colorectal cancer | BCL2, CASP3, IGF1R, ACVR1C |
| 4150 | .03533 | 3.868 75 | 3 | 51 | mTOR signaling pathway | IGF1, VEGFA, DDIT4 |
| 5414 | .03884 | 2.963230862 | 4 | 88 | Dilated cardiomyopathy | ADCY3, ADCY8, ATP2A2, IGF1 |
| 4010 | .04517 | 1.984836601 | 8 | 263 | MAPK signaling pathway | CASP3, CDC42, DUSP4, ELK1, FGF7, STMN1, NTRK2, ACVR1C |
| 52 | .04708 | 5.119341564 | 2 | 26 | Galactose metabolism | GCK, B4GALT2 |
| 450 | .04708 | 5.119341564 | 2 | 26 | Seleno amino acid metabolism | AHCY, CTH |
IHC reveals overexpression of target proteins of suppressed miRNAs in NKTL
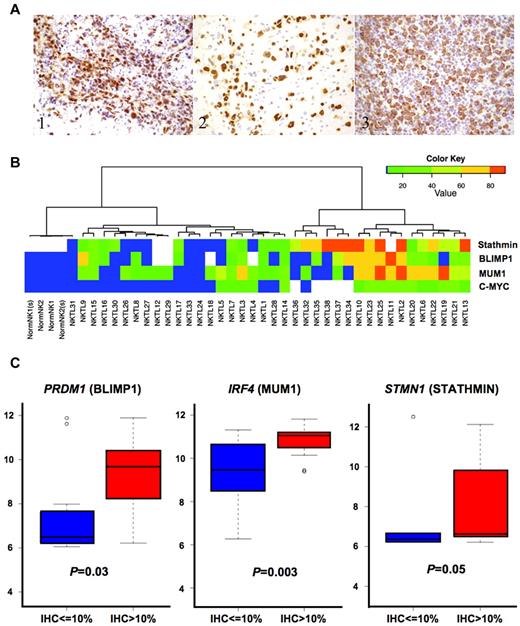
To further validate our gene expression results, we performed IHC for selected target proteins of the deregulated miRNAs in NKTL, including MUM1/IRF4, BLIMP1, and STMN1 on TMA sections containing 33 samples of NKTL and whole paraffin sections of 5 cases that were not included in the TMAs. In corroboration with the MEP findings, we observed a significant percentage of our NKTL cases showing positive expression for MUM1/IRF4 (20 of 38, 53%), BLIMP1 (17 of 34, 50%), and STMN1 (20 of 35, 57%; Figure 4A-B; supplemental Table 6A). In contrast, normal NK cells show minimal (≤ 5%) to absent expression of the 3 target proteins (supplemental Table 6B). Similarly, these proteins were aberrantly expressed in NKTL cell lines (supplemental Figure 4). In addition, cases with greater percentage (> 10%) of positive staining cells for the 3 proteins, had higher expression of the corresponding mRNA (Figure 4C). The expression of these mRNAs is also significantly inversely correlated with the expression of their regulating miRNAs (supplemental Figure 3), further suggesting that the overexpression of MUM1, BLIMP1, and STMN1 in NKTL may be driven by abnormal expression of their regulating miRNAs.
Expression of protein targets of deregulated miRNA. (A) IHC showing overexpression of (i) BLIMP1, (ii) MUM1, and (iii) STATHMIN in the tumor cells. BLIMP1 and MUM1 are expressed in the nuclei, whereas STATHMIN is expressed in the cytoplasm of the neoplastic lymphoid cells. All photographs were taken with a DP20 Olympus camera (Olympus) using an Olympus BX41 microscope (Olympus); images were acquired using DP Controller 2002 (Olympus) and processed using Adobe Photoshop Version 7.0 (Adobe Systems). Original magnifications ×600. (B) The percentage of tumor cells staining for the different protein markers are represented in the form of a heat map. The color scale corresponding to the percentage of positive staining cells is appended in the left upper corner. Samples with results represented by white indicate that the stain was not done because of inadequate material. Cases with the highest expression of STATHMIN, MUM1, and BLIMP1 also have the highest expression of MYC. (C) The expression of mRNA corresponding to these proteins was higher in those samples where > 10% of tumor cells are staining positive for each protein marker. For BLIMP1 and MUM1, it is statistically significant.
Expression of protein targets of deregulated miRNA. (A) IHC showing overexpression of (i) BLIMP1, (ii) MUM1, and (iii) STATHMIN in the tumor cells. BLIMP1 and MUM1 are expressed in the nuclei, whereas STATHMIN is expressed in the cytoplasm of the neoplastic lymphoid cells. All photographs were taken with a DP20 Olympus camera (Olympus) using an Olympus BX41 microscope (Olympus); images were acquired using DP Controller 2002 (Olympus) and processed using Adobe Photoshop Version 7.0 (Adobe Systems). Original magnifications ×600. (B) The percentage of tumor cells staining for the different protein markers are represented in the form of a heat map. The color scale corresponding to the percentage of positive staining cells is appended in the left upper corner. Samples with results represented by white indicate that the stain was not done because of inadequate material. Cases with the highest expression of STATHMIN, MUM1, and BLIMP1 also have the highest expression of MYC. (C) The expression of mRNA corresponding to these proteins was higher in those samples where > 10% of tumor cells are staining positive for each protein marker. For BLIMP1 and MUM1, it is statistically significant.
Mechanism of miRNA dysregulated in NKTL
Next, we investigated the possible mechanisms of miRNA dysregulation in NKTL. A number of the dysregulated miRNAs are located within host gene sequences. However, only 3 miRNAs, miR-152, miR-598, and miR-378, have correlated expression with their host genes. The eregulation of these miRNA and gene expression is not associated with known deletion or amplification of the genomic locus based on a previous publication of array comparative genomic hybridization analysis.17 Two underexpressed miRNAs, miR-186 and miR-101, are located within a genomic locus that is commonly deleted in NKTL, chromosome 1p21.3-p31.2 (Table 3).17 The miRNA signature of NKTL from our analysis is one associated with mainly down-regulation of miRNAs. Recently, it has been shown that MYC can cause extensive repression of miRNA expression.18 Indeed, in our cohort, tumor samples with increased expression of BLIMP1, MUM1, and STMN1 proteins, regulated by their underexpressed miRNAs, showed higher MYC nuclear expression, consistent with MYC activation (Figure 4B). EBV infection is universal in NKTL. Indeed, 4 of the deregulated miRNAs that we observed have been reported to be down-regulated (let 7g, let-7 and let-7c)19 and up-regulated (miR-155)20 on EBV infection (Table 3), suggesting that EBV infection may also have an effect on miRNA deregulation in NKTL.
Potential mechanisms of miRNA deregulation in NKTL
| Dysregulated miRNA . | Dysregulated in NKTL . | Genomic loci . | Affected by EBV infection . | Host gene . | Correlation with host gene expression . | Abnormality in genomic loci . | aCGH band . |
|---|---|---|---|---|---|---|---|
| hsa-miR-342–5p | − | Chromosome 14 | EVL | ||||
| hsa-miR-26b | − | Chromosome 2 | CTDSP1 | ||||
| hsa-miR-363 | − | Chromosome X | |||||
| hsa-miR-150 | − | Chromosome 19 | |||||
| hsa-miR-28–5p | − | Chromosome 3 | LPP | ||||
| hsa-miR-152 | − | Chromosome 17 | + | COPZ2 | Yes | ||
| hsa-miR-361–3p | − | Chromosome X | CHM | ||||
| hsa-miR-22* | − | Chromosome 17 | C17orf91 | ||||
| hsa-miR-340 | − | Chromosome 5 | RNF130 | ||||
| hsa-miR-598 | − | Chromosome 8 | XKR6 | Yes | |||
| hsa-miR-181a-2* | − | Chromosome 9 | NR6A1 | ||||
| hsa-miR-132 | − | Chromosome 17 | |||||
| hsa-miR-194 | − | Chromosome 1 | IARS2 | ||||
| hsa-miR-768–3p | − | Chromosome 16 | AP1G1 | ||||
| hsa-miR-873 | − | Chromosome 9 | |||||
| hsa-miR-338–3p | − | Chromosome 17 | AATK | ||||
| hsa-miR-215 | − | Chromosome 1 | IARS2 | ||||
| hsa-miR-186 | − | Chromosome 1 | ZRANB2 | − | 1p21.3-p31.2 | ||
| hsa-miR-140–3p | − | Chromosome 16 | WWP2 | ||||
| hsa-miR-140–5p | − | Chromosome 16 | WWP2 | ||||
| hsa-miR-374b | − | Chromosome X | NCRNA00182 | ||||
| hsa-miR-26a | − | Chromosome 12 | CTDSP2 | ||||
| hsa-let-7g | − | Chromosome 3 | − | WDR82 | |||
| hsa-miR-342–3p | − | Chromosome 14 | EVL | ||||
| hsa-miR-101 | − | Chromosome 1 | − | 1p21.3-p31.2 | |||
| hsa-miR-192 | − | Chromosome 11 | |||||
| hsa-miR-374a | − | Chromosome X | NCRNA00182 | ||||
| hsa-miR-876–5p | − | Chromosome 9 | |||||
| hsa-miR-22 | − | Chromosome 17 | C17orf91 | ||||
| hsa-miR-10a | − | Chromosome 17 | |||||
| hsa-miR-590–5p | − | Chromosome 7 | EIF4H | ||||
| hsa-miR-30b | − | Chromosome 8 | |||||
| hsa-miR-181c | − | Chromosome 19 | |||||
| hsa-miR-142–5p | − | Chromosome 17 | |||||
| hsa-let-7a | − | Chromosome 11 | − | LOC399959 | |||
| hsa-miR-155 | + | Chromosome 21 | + | MIR155HG | |||
| hsa-let-7c | − | Chromosome 21 | − | C21orf34 | |||
| hsa-miR-378 | + | Chromosome 5 | PPARGC1B | Yes | |||
| hsa-miR-181a | − | Chromosome 1 | |||||
| hsa-miR-142–3p | − | Chromosome 17 | |||||
| hsa-miR-15a | − | Chromosome 13 | DLEU2 |
| Dysregulated miRNA . | Dysregulated in NKTL . | Genomic loci . | Affected by EBV infection . | Host gene . | Correlation with host gene expression . | Abnormality in genomic loci . | aCGH band . |
|---|---|---|---|---|---|---|---|
| hsa-miR-342–5p | − | Chromosome 14 | EVL | ||||
| hsa-miR-26b | − | Chromosome 2 | CTDSP1 | ||||
| hsa-miR-363 | − | Chromosome X | |||||
| hsa-miR-150 | − | Chromosome 19 | |||||
| hsa-miR-28–5p | − | Chromosome 3 | LPP | ||||
| hsa-miR-152 | − | Chromosome 17 | + | COPZ2 | Yes | ||
| hsa-miR-361–3p | − | Chromosome X | CHM | ||||
| hsa-miR-22* | − | Chromosome 17 | C17orf91 | ||||
| hsa-miR-340 | − | Chromosome 5 | RNF130 | ||||
| hsa-miR-598 | − | Chromosome 8 | XKR6 | Yes | |||
| hsa-miR-181a-2* | − | Chromosome 9 | NR6A1 | ||||
| hsa-miR-132 | − | Chromosome 17 | |||||
| hsa-miR-194 | − | Chromosome 1 | IARS2 | ||||
| hsa-miR-768–3p | − | Chromosome 16 | AP1G1 | ||||
| hsa-miR-873 | − | Chromosome 9 | |||||
| hsa-miR-338–3p | − | Chromosome 17 | AATK | ||||
| hsa-miR-215 | − | Chromosome 1 | IARS2 | ||||
| hsa-miR-186 | − | Chromosome 1 | ZRANB2 | − | 1p21.3-p31.2 | ||
| hsa-miR-140–3p | − | Chromosome 16 | WWP2 | ||||
| hsa-miR-140–5p | − | Chromosome 16 | WWP2 | ||||
| hsa-miR-374b | − | Chromosome X | NCRNA00182 | ||||
| hsa-miR-26a | − | Chromosome 12 | CTDSP2 | ||||
| hsa-let-7g | − | Chromosome 3 | − | WDR82 | |||
| hsa-miR-342–3p | − | Chromosome 14 | EVL | ||||
| hsa-miR-101 | − | Chromosome 1 | − | 1p21.3-p31.2 | |||
| hsa-miR-192 | − | Chromosome 11 | |||||
| hsa-miR-374a | − | Chromosome X | NCRNA00182 | ||||
| hsa-miR-876–5p | − | Chromosome 9 | |||||
| hsa-miR-22 | − | Chromosome 17 | C17orf91 | ||||
| hsa-miR-10a | − | Chromosome 17 | |||||
| hsa-miR-590–5p | − | Chromosome 7 | EIF4H | ||||
| hsa-miR-30b | − | Chromosome 8 | |||||
| hsa-miR-181c | − | Chromosome 19 | |||||
| hsa-miR-142–5p | − | Chromosome 17 | |||||
| hsa-let-7a | − | Chromosome 11 | − | LOC399959 | |||
| hsa-miR-155 | + | Chromosome 21 | + | MIR155HG | |||
| hsa-let-7c | − | Chromosome 21 | − | C21orf34 | |||
| hsa-miR-378 | + | Chromosome 5 | PPARGC1B | Yes | |||
| hsa-miR-181a | − | Chromosome 1 | |||||
| hsa-miR-142–3p | − | Chromosome 17 | |||||
| hsa-miR-15a | − | Chromosome 13 | DLEU2 |
– indicates down-regulated; +, upregulated; and empty fields, no abnormalities detected.
Discussion
NKTL is a highly aggressive tumor, and a better understanding of the molecular abnormalities underlying this condition will provide important insights into the biology of this disease and potential new therapeutic avenues. To the best of our knowledge, this is the first comprehensive genome-wide study of miRNA expression profiling using the microarray platform on FFPE NTKL samples. The validity of our results was supported by quantitative PCR validation as well as corroboration of our in silico functional analysis with IHC in a larger dataset, showing good correlation between MEP, GEP, IHC, and RT-PCR results. In the present study, we characterized the miRNA signature of NKTL compared with normal NK cells. Our results identified the dysregulated miRNAs in NKTL, target genes involved, and activation of signaling pathways that may be relevant to the pathophysiology of the disease and could potentially serve as therapeutic targets.
We found that the predominant changes are down-regulation of miRNAs. We validated that the expression of a number of these down-regulated miRNAs, such as miR-101, miR-26a, miR26b, miR-30b, miR-28–5, and miR-363, affects growth of the NK-YS cell line. In addition, they modulated the expression of their predicted target genes, suggesting that these miRNAs are of functional relevance, and their suppression could lead to increased expression of a number of genes implicated in oncogenesis. Indeed, we confirmed, for the first time, that miR-101 directly regulate STMN1.
A recent study by Paik et al reported that miRNA-146a is down-regulated in NKTL and may function as a tumor suppressor in NK/T-cell lymphoma.21 In line with this study, we also detected down-regulation of miR-146a in FFPE NKTL compared with normal NK cells but not between NK cell lines and normal NK cells (see supplemental Tables 4 and 5). Only 2 miRNAs (miR-155 and miR-378) were up-regulated in both NKTL and NK cell lines compared with normal NK cells. Overexpression of miR-155 induced activation of AKT signaling pathway in NK cell lymphoma,7 whereas overexpression of miR-378 has been found to enhance cell survival, reduce caspase-3 activity, and promote tumor growth and angiogenesis.22 In corroboration with data from GEP studies in NKTL, targets of dysregulated miRNA in NKTL are significantly enriched for genes involved in the cell cycle-related pathway, p53 pathway, and MAPK signaling pathway.1-3 This suggests that oncogenic pathways activated in NKTL may be in part mediated by miRNA dysregulation.
BLIMP1 and MUM1/IRF4, 2 of the up-regulated targets identified in this study with corresponding protein overexpression, are of interest as they have been previously implicated in T- and NK-cell malignancies.23 Our study reveals that the up-regulation of BLIMP1 and IRF4 in NKTL may be driven by the suppression of their regulating miRNAs. Besides being a master regulator of terminal B-cell differentiation, BLIMP-1 also plays a role in the later stages of T-cell differentiation.24,25 Recently, BLIMP1 was shown to be required for NK-cell maturation and for regulating their proliferative potential.26 We found that BLIMP1 expression is significantly higher and aberrant in a subset of NKTL compared with normal NK cells. Expression of BLIMP1 is also associated with chemoresistance and poorer disease outcome in T-cell malignancies. The multiple myeloma oncogene-1 (MUM1/IRF4) encodes a transcription factor thought to play a central role in the development of lymphoid cells. Besides being expressed in B cells and plasma cells, IRF4 is known to be expressed in activated T-cell malignancies and to regulate T-cell transformation. Expression of IRF4 is also associated with inferior overall survival in peripheral T-cell lymphoma, and this association was observed across PTCL subtypes.27 Recently, MUM1/IRF4 expression in PTCLs, including NKTL, was linked to expression of BLIMP1. PTCL cell lines treated in vitro with the proteasome inhibitor bortezomib down-regulated MUM1/IRF4, an effect dependent on NF-κB inhibition and associated with BLIMP1 down-regulation.28 Given that IRF4 overexpression is oncogenic in vitro,29 and because NKTL lacks good treatment options, MUM1/IRF4 might represent a potential therapeutic target in patients with NKTL.
miRNAs are often encoded in fragile sites in the genome, where their expression can be altered by events, such as genomic amplification, loss of heterozygosity, viral integration, or genomic rearrangement.30 Our analysis revealed that 5% of the deregulated miRNAs in NKTL correlated with their host gene expression and may be deregulated as a result of abnormalities affecting the host genes. However, except for miR-101 and miR-186, which are located on 1p21.3-p31.2 that has been previously reported to be deleted in NKTL,17 chromosomal alteration appears to be an unlikely mechanism contributing to the deregulation of miRNAs in NKTL.
EBV infection has been described to regulate the expression of miRNAs in Burkitt lymphoma.31 Our results reveal the down-regulation of let-7g, let-7a, and let-7c, and up-regulation of miR-155 in both NK cell lines and FFPE NKTL samples. miRNAs let-7g, let-7a, and let-7c have been demonstrated in other studies to be down-regulated by EBV.19 Similarly, overexpression of miR-155 has been demonstrated in EBV-infected B lymphocytes displaying type III latency,20 and this is because of EBV gene expression and not epigenetic differences in cell lines tested.32 It is plausible that EBV may play a role in the dysregulation of these miRNAs in NKTL. Our result is also consistent with the study by Yamanaka et al, which demonstrated that overexpression of miR-155 resulted in the activation of AKT signaling pathway in NK cell lymphoma.7
As most of the dysregulated miRNAs are down-regulated, one important mechanism driving changes in miRNA may be MYC activation, which has been shown to repress a large number of miRNAs in tumor development.18 We have shown previously that MYC is activated in a substantial number of NKTL,3 and here we show correlation between MYC activation and overexpression of target proteins of down-regulated miRNA, suggesting that MYC activation may be one of the mechanisms in the deregulation of miRNA in NKTL.
In conclusion, our study indicates that the deregulation of miRNAs is of functional relevance, providing an additional mechanism by which some of the oncogenic pathways may be deregulated and hence contribute to the pathogenesis of NKTL. Furthermore, miRNAs may themselves be potential therapeutic targets that can be exploited in the future.33,34
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
W.-J.C. was supported by the National Medical Research Council Clinician Scientist Investigator Award. This work is supported in part by the Singapore Cancer Syndicate and the Research Center of Excellence Program (funded by the Singapore National Research Foundation and the Ministry of Education).
Authorship
Contribution: S.-B.N. designed experiments, performed IHC and scoring, and wrote the manuscript; J.Y. J.L.-S.T., and J.T. performed miRNA functional studies; G.H. performed bioinformatics analysis; V.S. and B.L. performed experiments; C.B. performed microarray experiments; Y.-L.K., N.S., and K.A. provided cell lines and approved the final manuscript; and W.-J.C. designed experiments, performed analysis, and wrote the manuscript.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Wee-Joo Chng, Department of Haematology-Oncology, National University Hospital, 5 Lower Kent Ridge Road, Main Building, Level 3, Singapore 119074; e-mail: mdccwj@nus.edu.sg; and Siok-Bian Ng, Department of Pathology, National University Hospital, 5 Lower Kent Ridge Road, Main Building, Level 3, Singapore 119074; e-mail: patnsb@nus.edu.sg.
References
Author notes
J.Y. and G.H. contributed equally to this study.