Key Points
Integrin signaling promotes proliferative signals in AML cells that are mediated by the kinase Syk and the transcription factors STAT3 and STAT5.
Spleen tyrosine kinase (Syk) induces cell survival and proliferation in a high proportion of acute myeloid leukemia (AML) blasts, but the underlying molecular events of Syk signaling have not been investigated. Proteomic techniques have allowed us to identify the multiprotein complex that is nucleated by constitutively active Syk in AML cells. This complex differs from the B-lymphoid Syk interactome with respect to several proteins, especially the integrin receptor Mac-1, the Fc-γ receptor I (FcγRI), and the transcription factors STAT3 and STAT5. We show in several AML cell line models that tonic signals derived from the Fc-γ chain lead to Syk-dependent activation of STAT3 and STAT5, which in turn induces cell survival and proliferation. Moreover, stimulation of Mac-1 or FcγRI intensifies the constitutive Syk-mediated STAT3/5 activation in AML cells, a scenario likely to take place in the bone marrow niche. In accordance with these findings, we observed that β2 integrins, including Mac-1, trigger proliferation of AML cells in an AML cell/stroma coculture model. Taken together, we identified an oncogenic integrin/Syk/STAT3/5 signaling axis that might serve as a therapeutic target of AML in the future.
Introduction
Constitutive activation of kinase-dependent signal transduction events has been shown to be a common event in acute myeloid leukemia (AML). Most frequently, mutational activation of Flt3 by internal tandem duplications (FLT3-ITD) results in the aberrant activation of several signaling pathways, often the activation of STAT3 and STAT5 proteins.1 Recent studies showed clinical response in patients with AML who were treated with endothelial growth factor receptor inhibitors in early clinical trials.2,3 Subsequently, it was shown in a ribonucleic acid interference approach that, in certain AML cell line models, gefitinib targeted spleen tyrosine kinase (Syk) and that these cells were dependent on Syk activity in vitro as well as in vivo.4 Syk is a nonreceptor tyrosine kinase that is expressed most in hematopoietic cells. Syk-mediated signaling is crucial for many immune and nonimmune biological responses and, furthermore, the aberrant expression or function of Syk is linked to a wide range of malignancies in hematopoietic and nonhematopoietic cell types.5,-7 Interestingly, the role of Syk is disease specific. For example, it has been shown that Syk acts as an oncoprotein in diffuse large B-cell lymphomas.6,8 In contrast, loss of Syk function contributes to the onset of pro–/pre–B-cell leukemias by causing a differentiation block in the B-cell lineage.9
Upon receptor stimulation, a phosphorylated immunoreceptor tyrosine-based activation motif (ITAM)/Syk-(SH2)2 interaction induces an allosteric activation of Syk by changing the conformation of Syk from a closed, inactive structure to an open, active form.10 Syk activity is furthermore controlled by the phosphorylation of distinct tyrosine residues within different domains of the kinase, a process that can be either driven by Src kinases or by Syk itself.11 In particular, the phosphorylation of tyrosines 348 and 352 is essential for conformational Syk activation. Several other phosphotyrosines within Syk serve as inducible docking sites for the SRC homology 2 (SH2) domains of other signaling proteins. These molecules can either regulate Syk function or, possibly, serve as substrates of Syk.10
In B cells, Syk is capable of regulating development and proliferation. This is achieved through the Syk-dependent activation of key signaling cascades downstream of the B-cell antigen receptor that include phosphatidylinositol 3-kinase, nuclear factor κB, and calcium ion pathways.12 Although Syk is studied in the context of myeloid effector cells such as mast cells or macrophages,13 very little is known about Syk’s role in myeloid progenitors. Only in 2009 was it shown that Syk is expressed in a large proportion of AML blasts. The kinase Syk triggers proliferation of AML blasts, and the pharmacological inhibition of Syk results in either apoptosis or differentiation into myeloid effector cells both in vitro and in vivo.4
However, the molecular mechanism of Syk function in AML cells has not been studied. Therefore, we characterized the Syk signaling network in AML cells by applying quantitative proteomic strategies and uncovered a functional link between the ITAM-bearing Fc-γ chain, β2 integrins, Syk, and the oncogenic transcription factors STAT3 and STAT5. Moreover, our functional analysis revealed that this constitutively activated signaling axis contributes to the survival and proliferation of AML cells. Because activation of STAT3 and STAT5 seems to be an important event for AML initiation and maintenance, as well as for leukemic stem cell self-renewal,14,,-17 and because it is frequently encountered in FLT3-ITD–negative cases,18 information about druggable upstream activators of STATs in AML is of interest.
Materials and methods
Cells and reagents
Peripheral blood samples were obtained from patients with AML at diagnosis or relapse with the approval of the ethics committee of the Johann Wolfgang Goethe University, Frankfurt, Germany (statement date August 28, 2002, renewed on January 22, 2009; statement file number 4/09). (For details, see supplemental Materials and methods). This study was conducted in accordance with the Declaration of Helsinki.
Metabolic labeling of KG-1 cells with stable isotopes of arginine and lysine (SILAC) was performed as described in Oellerich et al19 with the exception that the final concentrations of arginine and lysine were 60 mg/L and 120 mg/L, respectively.
To stimulate Fc-γ receptor I (FcγRI)), cells were treated with monoclonal antibodies for indicated time points as described previously in Melendez et al.20 To induce Mac-1 clustering, cells were treated with immobilized anti-αM F(ab′)2 fragments of LPM19c for indicated durations as described previously in Xue et al.21 Antibodies and further reagents are outlined in supplemental Materials and methods. The STAT3 complementary DNA was manipulated by site-directed mutagenesis in a way that upon expression cysteine residues substituted A661 and N663 of STAT3.22 The complementary DNA encoding for constitutively active STAT5A harbors point mutations resulting in the replacement of S711 by phenylalanine on the protein level.23 Retroviral transductions were performed as described previously.24
Immunoprecipitation experiments and immunoblotting
Syk, SLP65, and FcRγ were immunoprecipitated from 107 KG-1 cells using 2 μg of the respective antibodies per sample as described previously in Oellerich et al.24 In addition, Syk was immunoprecipitated from 5 × 106 cells of the primary AML cell lines. Immunoblotting was performed as described previously in Oellerich et al.24
Mass spectrometric analysis, database search, and MaxQuant-based protein quantification
Proteins were separated by one-dimensional polyacrylamide gel electrophoresis (4%-12% NuPAGE Bis-Tris Gel; Invitrogen) and the entire lane of the Coomassie blue-stained gel was cut into 23 slices. All slices were processed as detailed in supplemental Materials and methods.
In vitro kinase assays
To test for the ability of Syk to phosphorylate tyrosine residues 705 and 694 of STAT3 and STAT5, respectively, a kinase assay was performed using the biotinylated peptides EHPEADPGSAAPYLKTKFIC for STAT3 and TPVLAKAVDGYVKPQIK for STAT5 (Pierce). The kinase assay was performed as detailed in supplemental Materials and methods.
Flow cytometry
GFP expression of transduced KG-1 cells as well as CD34 expression was assessed by flow cytometry on a FACSCanto II instrument.
Cell proliferation assay
Cell proliferation was measured by an XTT-based method according to the protocol of the Cell Proliferation Kit II (Roche), as outlined in supplemental Materials and methods.
Results
Elucidation of the Syk activation state
Syk was reported earlier to be expressed in AML blasts.4 To decipher the Syk activation status in detail, we performed MS-based analysis of the Syk phosphorylation state in cells derived from 2 Syk expressing primary human AML cell cultures (FFM05 and FFM12) and from the human AML cell lines KG-1 and U937 (Figure 1A). The primary AML characteristics of patients, including the cytogenetic profiles, are outlined in supplemental Table 1. The phosphoproteomic analysis identified 25 phosphorylation sites (p-sites) within Syk in both primary AML and KG-1 cells (Figure 1B-C; supplemental Figure 1; and supplemental Table 2). In U937 cells, we identified 17 p-sites (supplemental Figure 1; supplemental Table 2). All activation-inducing tyrosines including Tyr 348 and Tyr 352 were found phosphorylated in all tested AML cells, indicating constitutive Syk activation in these cells. Interestingly, the proportion of tyrosine phosphorylation within Syk in AML cells is significantly higher compared with Syk in activated B-lymphoid cells. A comparison of Syk’s site-specific phosphorylation patterns in AML cells and activated B-cells reveals an overlap of 20 p-sites, while 5 p-sites were AML-specific and 12 p-sites were specific for B cells (Figure 1D and supplemental Table 3). Moreover, we tested the biological impact of Syk inhibition on AML cells. Both the primary human AML cell cultures and the KG-1 cells showed significantly reduced proliferation rates in the presence of pharmacological Syk inhibition or upon short hairpin RNA (shRNA)–based silencing of Syk (Figure 1E). Furthermore, reduced Syk expression resulted in the differentiation of FFM05 and FFM12 cells as monitored by a strong reduction of CD34 expression that was measured 7 days after Syk knockdown, while in the respective control cells CD34 expression did not change (Figure 1F). In contrast to the primary AML cell cultures, CD34 expression was not altered in KG-1 cells after knockdown of Syk (Figure 1F). Taken together, the results indicate significant tonic signaling in AML cells that involves Syk activation and leads to proliferation.
P-sites in myeloid Syk. (A) Cleared cellular lysates of Syk-deficient DT40 cells (lane 1), KG-1 cells (lane 2), primary AML cells (lanes 3 and 4), and wild-type DT40 cells (lane 5) were subjected to western blot analyses with antibodies against Syk (upper panel). Protein loading was monitored by anti-actin immunoblotting (lower panel). (B) Shown is the domain structure of human Syk, including the p-sites that were identified by mass spectrometry upon TiO2-based phosphopeptide enrichment. Phosphorylated tyrosine residues are marked in red, phosphorylated serine/threonine residues in blue. (C) The amino acid sequence of human Syk is shown with p-sites labeled as follows: All p-sites, which were identified in human AML cells, are labeled with red, bold, lowercase letters. Red, bold, capital letters indicate activation-inducing tyrosines that were identified. Bold, italic letters are used if a p-site could not be allocated to 1 of 2 adjacent amino acids. (D) Cell type–specific phosphorylation patterns are outlined. The red circle represents p-sites that were identified in AML cells in this study, while the black circle represents the total number of p-sites that were identified in B cells in previous studies.26 (E) Shown are growth curves of KG-1, FFM05, and FFM12 cells that were left untreated (upper panel, black lines) or were treated with the Syk inhibitor Bay 61-3606 at a final concentration of 250 nM (upper panel, gray lines). Shown are growth curves of KG-1, FFM05, and FFM12 cells that were treated with either control shRNA (lower panel, black lines) or Syk-specific shRNA (lower panel, gray lines). Proliferation was monitored by XTT-based assay. (F) FFM05, FFM12, and KG-1 cells were treated with control- or Syk-specific shRNA and were analyzed for CD34 expression by flow cytometry 7 days after transduction. FITC, fluorescein isothiocyanate; nsp, nonspecific; Syk kd, Syk knockdown.
P-sites in myeloid Syk. (A) Cleared cellular lysates of Syk-deficient DT40 cells (lane 1), KG-1 cells (lane 2), primary AML cells (lanes 3 and 4), and wild-type DT40 cells (lane 5) were subjected to western blot analyses with antibodies against Syk (upper panel). Protein loading was monitored by anti-actin immunoblotting (lower panel). (B) Shown is the domain structure of human Syk, including the p-sites that were identified by mass spectrometry upon TiO2-based phosphopeptide enrichment. Phosphorylated tyrosine residues are marked in red, phosphorylated serine/threonine residues in blue. (C) The amino acid sequence of human Syk is shown with p-sites labeled as follows: All p-sites, which were identified in human AML cells, are labeled with red, bold, lowercase letters. Red, bold, capital letters indicate activation-inducing tyrosines that were identified. Bold, italic letters are used if a p-site could not be allocated to 1 of 2 adjacent amino acids. (D) Cell type–specific phosphorylation patterns are outlined. The red circle represents p-sites that were identified in AML cells in this study, while the black circle represents the total number of p-sites that were identified in B cells in previous studies.26 (E) Shown are growth curves of KG-1, FFM05, and FFM12 cells that were left untreated (upper panel, black lines) or were treated with the Syk inhibitor Bay 61-3606 at a final concentration of 250 nM (upper panel, gray lines). Shown are growth curves of KG-1, FFM05, and FFM12 cells that were treated with either control shRNA (lower panel, black lines) or Syk-specific shRNA (lower panel, gray lines). Proliferation was monitored by XTT-based assay. (F) FFM05, FFM12, and KG-1 cells were treated with control- or Syk-specific shRNA and were analyzed for CD34 expression by flow cytometry 7 days after transduction. FITC, fluorescein isothiocyanate; nsp, nonspecific; Syk kd, Syk knockdown.
The Syk interactome in AML cells
To gain insight into Syk-driven signaling pathways, we characterized the Syk complex in AML cells. In order to identify the interaction partners of Syk, we performed a protein interaction screening by quantitative immunoprecipitation combined with knockdown (QUICK) analysis.25 The experimental setup is outlined in Figure 2A. KG-1 cells were left unlabeled (“light,” or L) or were metabolically labeled via SILAC with “heavy” (H) stable isotopes of lysine and arginine. Syk was subsequently knocked down by a Syk-specific shRNA in the L–KG-1 cell batch. The efficiency of the Syk knockdown is demonstrated in Figure 2B. Upon immunoprecipitation of Syk from both the L- and the H-labeled KG-1 batches, precipitated proteins were eluted, combined in a 1:1 ratio, and subsequently analyzed by mass spectrometry. This setup allowed for the discrimination of interacting proteins (ratio of H/L > 5) and unspecific background proteins (ratio of H/L ∼1; Figure 2C).
The Syk interactome in AML cells. (A) Shown is a schematic representation of the protein purification and identification process. KG-1 cells that were treated with shRNA specific for Syk were cultured in light SILAC medium and served as the negative control. Wild-type KG-1 cells were grown in heavy SILAC medium. Following immunoprecipitation of Syk from both cell batches, precipitated proteins were mixed in equimolar amounts, separated by 1D-PAGE, digested by trypsin, and then analyzed by mass spectrometry. (B) An immunoblot of cleared cellular lysates derived from wild-type KG-1 cells (lane 1) and KG-1 Syk knockdown cells (lane 2) with antibodies against Syk (upper panel) is shown. Protein loading was monitored by anti-actin immunoblotting (lower panel). (C) All identified proteins were plotted according to their signal intensities and their heavy versus light ratio of enrichment (H/L) on logarithmic scales. Red dots indicate proteins with an H/L ratio > 5 that were identified as interaction partners of Syk. A complete list of proteins and statistics is listed in supplemental Table 2. (D) KG-1 cells (left panel) and primary AML cells (middle and right panels) were lysed and subjected to immunoprecipitations (IPs) using anti-Syk antibodies (lanes 2, 4, and 6) or isotype-matched control antibodies (C; lanes 1, 3, and 5). Obtained proteins were analyzed by immunoblotting with antibodies directed against STAT3 and STAT5 (upper and middle panels). Effective IP of Syk was confirmed by immunoblotting using Syk-specific antibodies (lower panels). (E) Recombinant GST or GST-STAT3-SH2 proteins were subjected to lysates of KG-1 cells. Upon affinity purification, eluates were collected and analyzed by immunoblot analysis using antibodies specific for either Syk (upper panel) or GST (lower panel).
The Syk interactome in AML cells. (A) Shown is a schematic representation of the protein purification and identification process. KG-1 cells that were treated with shRNA specific for Syk were cultured in light SILAC medium and served as the negative control. Wild-type KG-1 cells were grown in heavy SILAC medium. Following immunoprecipitation of Syk from both cell batches, precipitated proteins were mixed in equimolar amounts, separated by 1D-PAGE, digested by trypsin, and then analyzed by mass spectrometry. (B) An immunoblot of cleared cellular lysates derived from wild-type KG-1 cells (lane 1) and KG-1 Syk knockdown cells (lane 2) with antibodies against Syk (upper panel) is shown. Protein loading was monitored by anti-actin immunoblotting (lower panel). (C) All identified proteins were plotted according to their signal intensities and their heavy versus light ratio of enrichment (H/L) on logarithmic scales. Red dots indicate proteins with an H/L ratio > 5 that were identified as interaction partners of Syk. A complete list of proteins and statistics is listed in supplemental Table 2. (D) KG-1 cells (left panel) and primary AML cells (middle and right panels) were lysed and subjected to immunoprecipitations (IPs) using anti-Syk antibodies (lanes 2, 4, and 6) or isotype-matched control antibodies (C; lanes 1, 3, and 5). Obtained proteins were analyzed by immunoblotting with antibodies directed against STAT3 and STAT5 (upper and middle panels). Effective IP of Syk was confirmed by immunoblotting using Syk-specific antibodies (lower panels). (E) Recombinant GST or GST-STAT3-SH2 proteins were subjected to lysates of KG-1 cells. Upon affinity purification, eluates were collected and analyzed by immunoblot analysis using antibodies specific for either Syk (upper panel) or GST (lower panel).
In AML cells, the interactome analysis revealed 18 proteins as Syk ligands (Table 1; supplemental Table 4). When we compared the Syk interactomes in AML and B-lymphoid cells, we observed an overlap regarding Vav, phosphatidylinositol 3-kinase, Cbl, dynein, tubulin, and 14-3-3 proteins.26,27 However, we identified several novel Syk interactors in AML cells that have not been described to interact with Syk in other cell types. Interestingly, we found that in all AML cells tested, the transcription factors STAT3 and STAT5 interacted with Syk. Constitutive activation of STAT3 and STAT5 has been reported to play a key role in leukemic transformation and uncontrolled proliferation of myeloid progenitor cells.28 In addition to STAT transcription factors, we found that in AML cells, Syk interacts with 2 cell surface receptors: the ITAM-bearing Fc-γRI and the integrin receptor Mac-1. These receptors are well-known activators of Syk in several cell types including macrophages, lymphocytes, and platelets.29
Syk interactome in AML cells
| Ligand . | IPI no. . | Function . | Found in B cells . |
|---|---|---|---|
| STAT5A | 00030783 | Transcription factor | No |
| STAT3 | 00784414 | Transcription factor | No |
| Cbl | 00884054 | E3 ubiquitin ligase | Yes45 |
| Vav | 00011696 | Guanosine exchange factor | Yes46 |
| PI3K p85 | 00807573 | Regulatory subunit of PI3-kinase | Yes47 |
| PIK3AP1 | 00783519 | Adaptor for PI3-kinase | No |
| 14-3-3-γ | 00220642 | Adaptor protein | Yes26 |
| 14-3-3-ζ/-δ | 00982101 | Adaptor protein | Yes26 |
| Ibtk | 00792333 | Inhibitor of Btk | No |
| FcγRI | 00023502 | High affinity Fc receptor | No |
| Integrin β2 | 00291792 | Subunit of Mac-1 | No |
| Integrin, α M | 00645887 | Subunit of Mac-1 | No |
| Tubulin α-4a | 00007750 | Cytoskeleton component | Yes26,27 |
| Tubulin α-1C | 00218343 | Cytoskeleton component | Yes26,27 |
| Tubulin β-2C | 00007752 | Cytoskeleton component | Yes26,27 |
| Tubulin β chain | 01019113 | Cytoskeleton component | Yes26,27 |
| Dynein 1 intermediate chain-2 | 00744015 | Motor protein | Yes26,27 |
| Dynein 1 heavy chain-1 | 00456969 | Motor protein | Yes26,27 |
| Ligand . | IPI no. . | Function . | Found in B cells . |
|---|---|---|---|
| STAT5A | 00030783 | Transcription factor | No |
| STAT3 | 00784414 | Transcription factor | No |
| Cbl | 00884054 | E3 ubiquitin ligase | Yes45 |
| Vav | 00011696 | Guanosine exchange factor | Yes46 |
| PI3K p85 | 00807573 | Regulatory subunit of PI3-kinase | Yes47 |
| PIK3AP1 | 00783519 | Adaptor for PI3-kinase | No |
| 14-3-3-γ | 00220642 | Adaptor protein | Yes26 |
| 14-3-3-ζ/-δ | 00982101 | Adaptor protein | Yes26 |
| Ibtk | 00792333 | Inhibitor of Btk | No |
| FcγRI | 00023502 | High affinity Fc receptor | No |
| Integrin β2 | 00291792 | Subunit of Mac-1 | No |
| Integrin, α M | 00645887 | Subunit of Mac-1 | No |
| Tubulin α-4a | 00007750 | Cytoskeleton component | Yes26,27 |
| Tubulin α-1C | 00218343 | Cytoskeleton component | Yes26,27 |
| Tubulin β-2C | 00007752 | Cytoskeleton component | Yes26,27 |
| Tubulin β chain | 01019113 | Cytoskeleton component | Yes26,27 |
| Dynein 1 intermediate chain-2 | 00744015 | Motor protein | Yes26,27 |
| Dynein 1 heavy chain-1 | 00456969 | Motor protein | Yes26,27 |
Two independent SILAC-based liquid chromatography-tandem mass spectrometry analyses were performed for identification of Syk interactors in untreated KG-1 cells. Mass spectrometric data sets were analyzed by MaxQuant software. The proteins listed showed an at least fivefold enrichment of heavy peptides compared with light ones. The protein functions listed were obtained by a literature search. In addition the international protein index number (IPI no.) is listed for each identified protein. Detailed statistics including total numbers of all identified and quantified peptides are outlined in supplemental Table 4.
By co-immunoprecipitation experiments and subsequent immunoblot analysis, we confirmed the interaction of Syk and STAT3 and STAT5 in KG-1 cells as well as in AML cells derived from the primary cultures mentioned above (Figure 2D). Furthermore, we detected an interaction between Syk and the SH2 domain of STAT3 in a glutathione S-transferase (GST) affinity purification experiment (Figure 2E). Taken together, our analysis revealed that in AML cells, constitutively active Syk assembles a multiprotein complex that differs from the B-lymphoid Syk complex. Interestingly, signal intermediates that have been shown to be important mediators of AML transformation and maintenance are recruited to this AML-specific Syk complex.
Syk activates STAT3 and STAT5
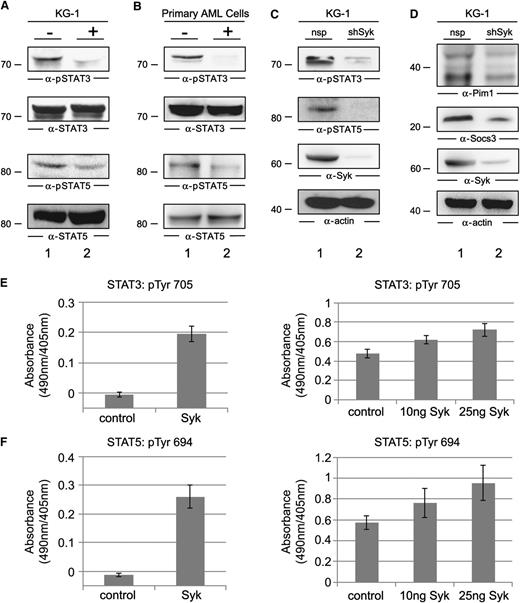
Next we investigated the biochemical and functional interplay of the identified AML-specific binding partners with Syk. First we tested whether STAT3 and STAT5 are active and whether their activation is Syk dependent. Therefore, we analyzed the activation state of STAT3 and STAT5 in the presence or absence of Syk inhibitors. Immunoblots with phosphosite-specific antibodies revealed the constitutive phosphorylation of Tyr 705 and Tyr 694 of STAT3 and STAT5, respectively, in KG-1 cells as well as in AML cells derived from the primary cultures. In the presence of the Syk inhibitor Bay 61-3606, we observed a profound decrease of STAT3 and STAT5 phosphorylation (Figure 3A-B). To exclude the possibility that the observation was due to off-target effects of the kinase inhibitor, we knocked down Syk in KG-1 cells using a specific shRNA and obtained similar results as with the small molecule inhibitor in KG-1 cells (Figure 3C). Moreover, we observed strongly reduced expression of the STAT3/5 target genes pim1 and socs3 in the Syk knockdown cells (Figure 3D).
Functional characterization of the Syk/STAT interplay. (A) Immunoblot analysis was performed with anti-pSTAT3 and anti-pSTAT5 antibodies (upper panels) or anti-STAT3 and anti-STAT5 antibodies (lower panels) of untreated KG-1 cells (−; lane 1) or KG-1 cells that were treated with the Syk inhibitor Bay 61-3606 at a final concentration of 250 nM for 1 hour (+; lane 2). (B) Immunoblot analysis was performed using the same antibodies as described in (A) of untreated AML cells derived from the primary culture FFM05 (−; lane 1) and AML cells from the same culture, but they were treated with the Syk inhibitor as described in (A) (+; lane 2). (C) Cleared cellular lysates of KG-1 cells treated with unspecific shRNA (lane 1) or Syk-specific shRNA (lane 2) were analyzed by immunoblotting with phosphosite-specific antibodies against the activatory tyrosines of STAT3 and STAT5 (two upper panels). Syk expression and protein loading were monitored by immunoblotting with antibodies against Syk or actin (two lower panels). (D) Three days after shRNA transduction, KG-1 cells were tested for their expression levels of Pim1 and Socs3 by immunoblotting using specific antibodies (upper and two middle panels). Protein loading was monitored by anti-actin immunoblotting (bottom panel). (E-F) Phosphorylation of Tyr 705 of STAT3 and Tyr 694 of STAT5 was monitored using an in vitro kinase assay. For this purpose, the biotinylated peptides encompassing the amino acids EHPEADPGSAAPYLKTKFIC for STAT3 and TPVLAKAVDGYVKPQIK for STAT5 were used as substrates for the Syk obtained from untreated KG-1 cells (left panels) or for the enzymatically active recombinant kinase domain of Syk (right panels). Tyrosine phosphorylation of these peptides was monitored by antiphosphotyrosine staining with enzyme-linked immunosorbent assay.
Functional characterization of the Syk/STAT interplay. (A) Immunoblot analysis was performed with anti-pSTAT3 and anti-pSTAT5 antibodies (upper panels) or anti-STAT3 and anti-STAT5 antibodies (lower panels) of untreated KG-1 cells (−; lane 1) or KG-1 cells that were treated with the Syk inhibitor Bay 61-3606 at a final concentration of 250 nM for 1 hour (+; lane 2). (B) Immunoblot analysis was performed using the same antibodies as described in (A) of untreated AML cells derived from the primary culture FFM05 (−; lane 1) and AML cells from the same culture, but they were treated with the Syk inhibitor as described in (A) (+; lane 2). (C) Cleared cellular lysates of KG-1 cells treated with unspecific shRNA (lane 1) or Syk-specific shRNA (lane 2) were analyzed by immunoblotting with phosphosite-specific antibodies against the activatory tyrosines of STAT3 and STAT5 (two upper panels). Syk expression and protein loading were monitored by immunoblotting with antibodies against Syk or actin (two lower panels). (D) Three days after shRNA transduction, KG-1 cells were tested for their expression levels of Pim1 and Socs3 by immunoblotting using specific antibodies (upper and two middle panels). Protein loading was monitored by anti-actin immunoblotting (bottom panel). (E-F) Phosphorylation of Tyr 705 of STAT3 and Tyr 694 of STAT5 was monitored using an in vitro kinase assay. For this purpose, the biotinylated peptides encompassing the amino acids EHPEADPGSAAPYLKTKFIC for STAT3 and TPVLAKAVDGYVKPQIK for STAT5 were used as substrates for the Syk obtained from untreated KG-1 cells (left panels) or for the enzymatically active recombinant kinase domain of Syk (right panels). Tyrosine phosphorylation of these peptides was monitored by antiphosphotyrosine staining with enzyme-linked immunosorbent assay.
To investigate whether Syk directly activates STAT3 and STAT5, we performed an in vitro kinase assay using biotinylated peptides encompassing the activation-inducing tyrosine motifs of STAT3 and STAT5. Affinity-purified Syk from untreated KG-1 cells was subjected to this kinase assay. Both STAT3- and STAT5-derived peptides were phosphorylated by Syk in vitro (Figure 3E-F, left panels). We conclude from these experiments that Syk activates STAT3 and STAT5 by phosphorylating the Tyr 705 of STAT3 and the Tyr 694 of STAT5, presumably through direct interaction. To exclude, finally, the possibility that a kinase potentially copurified with Syk might contribute to the phosphorylation of the STAT peptides, we subjected both peptides to the action of a recombinant enzymatically active kinase domain of Syk. As shown in Figure 3E-F (right panels), we detected again tyrosine phosphorylation of Tyr 705 and Tyr 694, thus confirming that Syk is able to phosphorylate STAT3 and STAT5 directly.
Constitutively active STAT3 and STAT5 counteract the effects of Syk inhibition
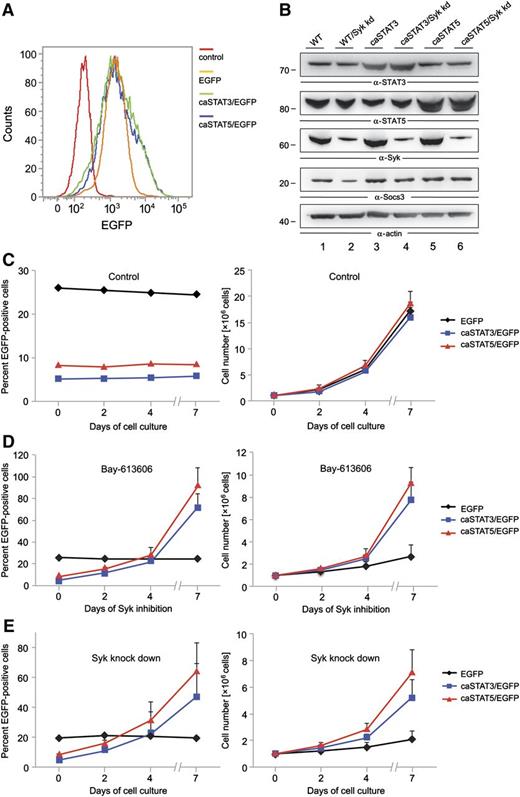
Our biochemical analyses identified STAT3 and STAT5 as substrates of Syk in AML cells. Both transcription factors are known as oncogenic effectors in AML.30 Thus, we tested whether the oncogenic effect of Syk in AML is caused by Syk-mediated activation of STAT3 and STAT5. For this purpose, we retrovirally transduced KG-1 cells using IRES-GFP vectors to express GFP alone or constitutive active forms of either STAT3/GFP (caSTAT3) or STAT5/GFP (caSTAT5). We subsequently treated these transduced cell batches with either the Syk inhibitor Bay 61-3606 for 7 days or Syk-specific shRNAs. To test in our experimental setup whether the caSTAT3 and caSTAT5 variants are constitutively active (as reported before in Bromberg et al22 and Moriggl et al 23 ), we enriched the cells expressing the respective caSTAT variants by fluorescence-activated cell sorting and examined the expression of the STAT target gene Socs3 (Figure 4A-B). To prove the Syk independency of the STAT variants, we knocked down Syk within the sorted cell batches (Figure 4B). We observed that in Syk knockdown cells, the expression of Socs3 was reduced in the control cells, while Socs3 expression was not altered in the cells expressing either caSTAT3 or caSTAT5, indicating the constitutive activity of both transcription factors (Figure 4B).
STAT3 and STAT5 function as oncogenic effectors of Syk. KG-1 cells were subjected to retroviral transduction with IRES-EGFP vectors inducing stable expression of EGFP, caSTAT3 and EGFP, or caSTAT5 and EGFP. (A) After fluorescence-activated cell sorting for EGFP, the percentage of GFP-expressing transduced KG-1 cells was monitored by flow cytometry. (B) Syk was knocked down by shRNA treatment in the sorted KG-1 cell batches that were enriched for EGFP-expressing cells. Expression of STAT3, STAT5, Syk, Socs3, and actin was monitored by immunoblot analyses 24 hours after lentiviral shRNA treatment. The transduced cell batches were left untreated (C) or were treated with the Syk inhibitor Bay 61-3606 at a final concentration of 250 nM for up to 7 days (D). (E) The transduced cell batches were left untreated or were treated with shRNAs specific for Syk. After confirmation of successful knockdown of Syk, cells were cultured for up to 7 days. The proportion of EGFP-positive cells was analyzed by flow cytometry (C-E, left panels). Absolute cell numbers are shown in the right panels. Wild-type (WT) KG-1 cells were used as the negative control in all flow cytometric measurements (data not shown).
STAT3 and STAT5 function as oncogenic effectors of Syk. KG-1 cells were subjected to retroviral transduction with IRES-EGFP vectors inducing stable expression of EGFP, caSTAT3 and EGFP, or caSTAT5 and EGFP. (A) After fluorescence-activated cell sorting for EGFP, the percentage of GFP-expressing transduced KG-1 cells was monitored by flow cytometry. (B) Syk was knocked down by shRNA treatment in the sorted KG-1 cell batches that were enriched for EGFP-expressing cells. Expression of STAT3, STAT5, Syk, Socs3, and actin was monitored by immunoblot analyses 24 hours after lentiviral shRNA treatment. The transduced cell batches were left untreated (C) or were treated with the Syk inhibitor Bay 61-3606 at a final concentration of 250 nM for up to 7 days (D). (E) The transduced cell batches were left untreated or were treated with shRNAs specific for Syk. After confirmation of successful knockdown of Syk, cells were cultured for up to 7 days. The proportion of EGFP-positive cells was analyzed by flow cytometry (C-E, left panels). Absolute cell numbers are shown in the right panels. Wild-type (WT) KG-1 cells were used as the negative control in all flow cytometric measurements (data not shown).
The initial transduction efficiency was about 25% for the empty vector control and about 5% for caSTAT3/5, and the proportion of EGFP-positive cells did not change over 7 days of cell culture (Figure 4C). Whereas we observed a significant increase in the proportion of cells that express either caSTAT3 or caSTAT5 upon 7 days of continuous Syk inhibition, the percentage of cells expressing GFP alone did not change over time (Figure 4D, left panel). To rule out that off-target effects of the Syk inhibitor might be responsible for this phenomenon, we knocked down Syk expression in the KG-1 cells mentioned above and observed the same effect as with the Syk inhibitor (Figure 4E, left panel). Both Syk inhibition and Syk knockdown decreased the proliferation rate of the control cells to a much greater extent than the cells expressing caSTAT3 or caSTAT5 (Figure 4D-E, right panels). We conclude from this experiment that Syk can act as a proliferation-inducing oncoprotein in AML cells by activating STAT3 and STAT5, although we cannot exclude that other Syk-dependent pathways also contribute to Syk’s proliferation-inducing signaling capacity.
Activation mechanisms of Syk/STAT signaling in AML cells
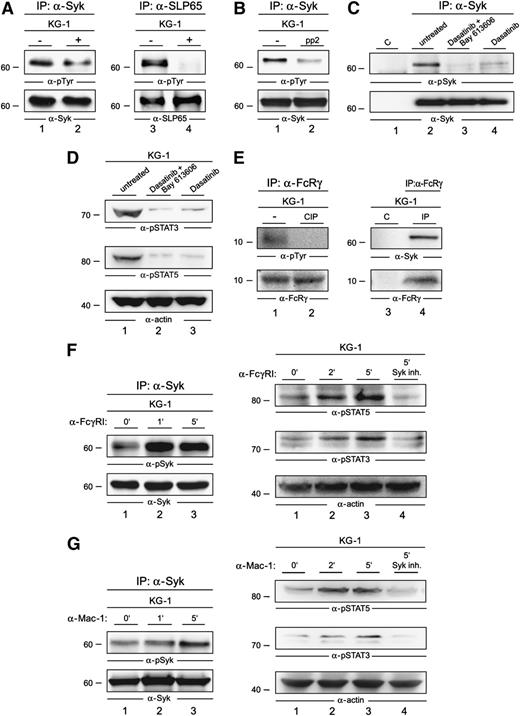
The constitutive activation of Syk in AML cells might be driven by autophosphorylation, by other kinases like Src family members, or by both mechanisms in parallel. To test which of these mechanisms are operational in AML cells, we examined the phosphorylation state of Syk in the presence or absence of a Syk inhibitor. We observed only a moderate decrease of Syk phosphorylation in the presence of the Syk inhibitor Bay 61-3606 at a final concentration of 250 nM (Figure 5A, left panel). Under these conditions, known Syk substrates like SLP65 were not detectably phosphorylated (Figure 5A, right panel). This indicates that kinases other than Syk contribute to Syk activation. Kinases of the Src family are known upstream regulators of Syk function in other cell types. Thus, we investigated the potential of Src kinase inhibition to suppress Syk activation in AML cells. Upon treatment of KG-1 cells with Src kinase inhibitors pp2 as well as dasatinib, a strong decrease of the activation-inducing tyrosine phosphorylation of Syk (Figure 5B-C) was observed. In line with this observation, our proliferation assay revealed that both KG-1 cells and the primary human AML cell cultures critically depend on Src kinase activity regarding their proliferation capacity (supplemental Figure 2). Moreover, dasatinib treatment likewise Syk inhibitor treatment led to diminished activation of STAT3 and STAT5 (Figure 5D).
Activation mechanisms of Syk in AML cells. (A) Lysates of KG-1 cells that were left untreated (−; lanes 1 and 3) or treated with Bay 61-3606 for 1 hour at a final concentration of 250 nM (+; lanes 2 and 4) were subjected to IP with antibodies against Syk (left panel) or SLP65 (right panel). Subsequently, immunoblot analyses of the obtained proteins were performed by using antiphosphotyrosine antibodies (4G10) (upper panels) and, in addition, antibodies against Syk and SLP65 (lower panels). (B) KG-1 cells were left untreated (−) or were treated with the Src-kinase inhibitor pp2 for 1 hour at a final concentration of 1 µM. The respective lysates were subjected to IP with antibodies against Syk, followed by immunoblot analyses of the proteins obtained; the immunoblot employed antibodies against phosphotyrosine (upper panel) or Syk (lower panel). (C) KG-1 cells were left untreated (lane 2) or were treated with dasatinib (4 nM) and Bay 61-3606 (250 nM) (lane 3), or dasatinib (4 nM) (lane 4) for 1 hour. The respective lysates were subjected to IP with antibodies against Syk, followed by immunoblot analysis of the proteins obtained; the immunoblot employed antibodies against phospho-Syk (pSyk; upper panel) or Syk (lower panel). Isotype-matched antibodies were used as the control (lane 1). (D) Immunoblot analysis was performed with anti-pSTAT3, anti-pSTAT5, and anti-actin antibodies of lysates derived from untreated KG-1 cells (lane 1), KG-1 cells that were treated with either dasatinib (4 nM) and Bay 61-3606 (250 nM) (lane 2), or dasatinib (4 nM) for 1 hour (lane 3). (E) Lysates of KG-1 cells were left untreated (left panel, lane 1; right panel, lanes 3 and 4) or were treated with the phosphatase CIP for 30 minutes (left panel, lane 2) and subsequently subjected to IP with antibodies specific for the Fc-γ chain (left panel, lanes 1 and 2; right panel, lane 4) or nonspecific control antibodies of the same isotype (right panel, lane 3). The proteins obtained were analyzed by antiphosphotyrosine antibodies (left panel, upper immunoblot) or by antibodies against Syk (right panel, upper immunoblot). Effective immunoprecipitation of the Fc-γ chain was confirmed by anti–Fc-γ chain immunoblotting (left and right panels, lower immunoblots). (F) Lysates of KG-1 cells that were left untreated (left panel, lane 1) or were stimulated for 1 or 5 minutes through FcγRI (left panel, lanes 2 and 3) were subjected to IP with antibodies against Syk, followed by immunoblotting using antibodies against pTyr525/526 of Syk (left, upper panel) and Syk (left, lower panel). Shown are immunoblot analyses of lysates derived from KG-1 cells that were left untreated (right panel, lane 1), stimulated via FcγRI for 2 or 5 minutes (right panel, lanes 2 and 3, respectively), or stimulated for 5 minutes after a 1-hour treatment with the Syk inhibitor Bay 61-3606 (right panel, lane 4). Immunoblotting was performed by using phosphosite-specific antibodies against pTyr 694 of STAT5 and pTyr 705 of STAT3 (right, upper and middle panels). Protein loading was monitored by the immunoblotting of actin (right, lower panel). (G) Lysates of KG-1 cells that were left untreated (left panel, lane 1) or were stimulated for 1 or 5 minutes through the integrin receptor Mac-1 (left panel, lanes 2 and 3, respectively) were treated as described in (D). Shown are immunoblot analyses of lysates derived from KG-1 cells that were left untreated (right panel, lane 1), stimulated by the integrin receptor Mac-1 for 2 or 5 minutes (right panel, lanes 2 and 3, respectively), or stimulated for 5 minutes after a 1-hour treatment with the Syk inhibitor Bay 61-3606 (right panel, lane 4). Immunoblotting was performed as described in (D).
Activation mechanisms of Syk in AML cells. (A) Lysates of KG-1 cells that were left untreated (−; lanes 1 and 3) or treated with Bay 61-3606 for 1 hour at a final concentration of 250 nM (+; lanes 2 and 4) were subjected to IP with antibodies against Syk (left panel) or SLP65 (right panel). Subsequently, immunoblot analyses of the obtained proteins were performed by using antiphosphotyrosine antibodies (4G10) (upper panels) and, in addition, antibodies against Syk and SLP65 (lower panels). (B) KG-1 cells were left untreated (−) or were treated with the Src-kinase inhibitor pp2 for 1 hour at a final concentration of 1 µM. The respective lysates were subjected to IP with antibodies against Syk, followed by immunoblot analyses of the proteins obtained; the immunoblot employed antibodies against phosphotyrosine (upper panel) or Syk (lower panel). (C) KG-1 cells were left untreated (lane 2) or were treated with dasatinib (4 nM) and Bay 61-3606 (250 nM) (lane 3), or dasatinib (4 nM) (lane 4) for 1 hour. The respective lysates were subjected to IP with antibodies against Syk, followed by immunoblot analysis of the proteins obtained; the immunoblot employed antibodies against phospho-Syk (pSyk; upper panel) or Syk (lower panel). Isotype-matched antibodies were used as the control (lane 1). (D) Immunoblot analysis was performed with anti-pSTAT3, anti-pSTAT5, and anti-actin antibodies of lysates derived from untreated KG-1 cells (lane 1), KG-1 cells that were treated with either dasatinib (4 nM) and Bay 61-3606 (250 nM) (lane 2), or dasatinib (4 nM) for 1 hour (lane 3). (E) Lysates of KG-1 cells were left untreated (left panel, lane 1; right panel, lanes 3 and 4) or were treated with the phosphatase CIP for 30 minutes (left panel, lane 2) and subsequently subjected to IP with antibodies specific for the Fc-γ chain (left panel, lanes 1 and 2; right panel, lane 4) or nonspecific control antibodies of the same isotype (right panel, lane 3). The proteins obtained were analyzed by antiphosphotyrosine antibodies (left panel, upper immunoblot) or by antibodies against Syk (right panel, upper immunoblot). Effective immunoprecipitation of the Fc-γ chain was confirmed by anti–Fc-γ chain immunoblotting (left and right panels, lower immunoblots). (F) Lysates of KG-1 cells that were left untreated (left panel, lane 1) or were stimulated for 1 or 5 minutes through FcγRI (left panel, lanes 2 and 3) were subjected to IP with antibodies against Syk, followed by immunoblotting using antibodies against pTyr525/526 of Syk (left, upper panel) and Syk (left, lower panel). Shown are immunoblot analyses of lysates derived from KG-1 cells that were left untreated (right panel, lane 1), stimulated via FcγRI for 2 or 5 minutes (right panel, lanes 2 and 3, respectively), or stimulated for 5 minutes after a 1-hour treatment with the Syk inhibitor Bay 61-3606 (right panel, lane 4). Immunoblotting was performed by using phosphosite-specific antibodies against pTyr 694 of STAT5 and pTyr 705 of STAT3 (right, upper and middle panels). Protein loading was monitored by the immunoblotting of actin (right, lower panel). (G) Lysates of KG-1 cells that were left untreated (left panel, lane 1) or were stimulated for 1 or 5 minutes through the integrin receptor Mac-1 (left panel, lanes 2 and 3, respectively) were treated as described in (D). Shown are immunoblot analyses of lysates derived from KG-1 cells that were left untreated (right panel, lane 1), stimulated by the integrin receptor Mac-1 for 2 or 5 minutes (right panel, lanes 2 and 3, respectively), or stimulated for 5 minutes after a 1-hour treatment with the Syk inhibitor Bay 61-3606 (right panel, lane 4). Immunoblotting was performed as described in (D).
In most studied cell systems, activation of Src kinases and Syk has been shown to be controlled by cell surface receptors.31 As described above, we identified an interaction of both cell surface receptors FcγRI and Mac-1 with Syk (Table 1). Both receptors are transmembrane proteins that are intracellularly associated with the ITAM-containing, signal transducing Fc-γ chain (FcRγ).32 To elucidate the mechanism that leads to the constitutive activation of Syk, we tested whether FcRγ is constitutively activated in AML cells. Indeed, we observed tyrosine phosphorylation of the ITAM-bearing Fc-γ chain, indicating the existence of tonic receptor signaling that might contribute to constitutive activation of Syk (Figure 5E, left panel). To further analyze this, we performed co-immunoprecipitation experiments and detected an interaction of the Fc-γ chain with Syk; this also supported the presence of a constitutively active ITAM-Syk complex in AML cells (Figure 5E, right panel). As both the FcγRI and the integrin receptor Mac-1 can transduce signals through the Fc-γ chain, it is likely that both receptors are involved in the constitutive activation of Syk.
Next we investigated, whether the engagement of either FcγRI or the integrin receptor Mac-1 can boost the activation of the Syk/STAT signaling axis. We observed an increase of Syk activation upon FcγRI stimulation (Figure 5F, left panel) and, furthermore, we detected a Syk-dependent boost of STAT3 and STAT5 activation (Figure 5F, right panel). The same phenomenon occurred upon Mac-1 stimulation (Figure 5G). In summary, we have identified a constitutively active Syk/STAT signaling axis that can be boosted by stimulation of FcγRI and Mac-1.
β2 integrins and the Fc-γ chain induce AML cell proliferation through Syk
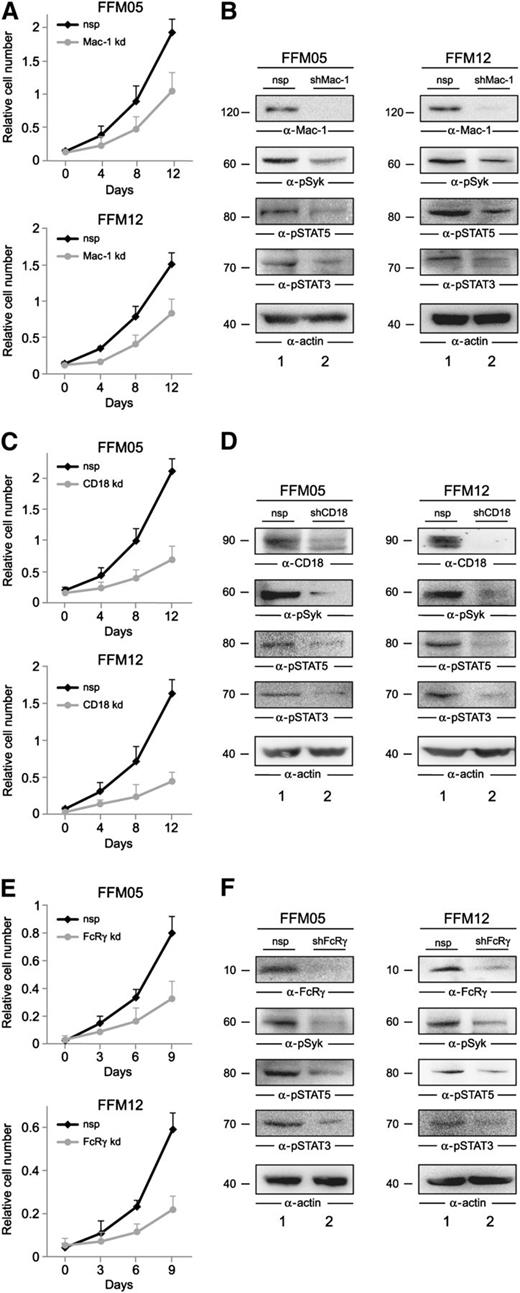
Having identified the 2 cell surface receptors Mac-1 and FcγRI as putative triggers of Syk-dependent AML cell proliferation, we set out to test the hypothesis that both receptors might contribute to AML cell proliferation via Syk in an AML cell/stroma coculture model. First we knocked down Mac-1 in the primary AML cell cultures FFM05 and FFM12 by using Mac-1–specific shRNAs. In the second step, we cultured the Mac-1 knockdown cells in the presence of a confluent layer of KM-102 bone marrow−derived stroma cells. FFM05/12 cells that were treated with the respective nonspecific control shRNAs were cocultured in the same way. We monitored cell proliferation of the shRNA-treated cell batches by XTT-based assays and observed that the knockdown of Mac-1 reduces cell proliferation significantly in the coculture model (Figure 6A). Moreover, we harvested the shRNA-treated FFM05/12 cells after 2 days of coculture and analyzed the activation state of Syk, STAT3, and STAT5 by immunoblot analysis. The purity of the AML cells was controlled by flow cytometric immunophenotyping using pan-leukocyte and myeloid-associated markers as described in Weir et al.33 We found that both Syk and STAT3/5 phosphorylation were reduced but still detectable in Mac-1 knockdown cells (Figure 6B). Thus, we hypothesized that other β2 integrins might redundantly activate Syk. To test this, we knocked down CD18 in FFM05 and FFM12 cells and analyzed proliferation as well as the activation state of the Syk-STAT signaling axis in the coculture model mentioned above. We observed that proliferation of CD18 knockdown cells, which express only a very low amount of β2 integrins, is reduced to a greater extent than Mac-1 knockdown cells (Figure 6C). Moreover, activation of Syk, STAT3, and STAT5 is barely detectable in CD18 knockdown cells, supporting our hypothesis regarding redundant functions of β2 integrin family members (Figure 6D). Finally, we tested whether the ITAM-containing Fc-γ chain that was shown to link integrins to Syk is needed to promote proliferation in the AML/stroma coculture model. This analysis revealed a strong reduction of proliferation in AML cells with diminished Fc-γ chain expression and, again, the activation of the Syk-STAT signaling axis was significantly reduced (Figure 6E-F). Of note, we also investigated the role of FcγRI in the regulation of proliferation in the coculture model, because we have also identified this receptor as part of the Syk interactome. We did not observe any effect on proliferation when we analyzed FcγRI knockdown cells using the assays outlined above (data not shown). In summary, these results support our biochemical data and show that β2 integrins promote proliferation of AML cells by activating Syk and its downstream targets.
β2 integrins induce proliferation of AML cells by the activation of Syk. (A-E) Shown are growth curves of FFM05 and FFM12 cells that were treated with shRNAs against (A) Mac-1, (C) CD18, and (E) FcRγ (all shown as gray lines) or the respective control shRNAs (black lines). The shRNA-treated FFM05/12 cells were cultured on confluent layers of KM-102 bone marrow–derived stroma cells. Proliferation was monitored by XTT-based assay. After 48 hours of AML/stroma cell coculture, FFM05/12 cells were harvested, and lysates of FFM05 and FFM12 cells that were treated with (B, D, F) control shRNAs (lane 1) or (B) Mac-1–, (D) CD18-, or (F) FcRγ-specific shRNAs (all shown in lane 2) were subjected to immunoblot analyses using antibodies against Mac-1, CD18, FcRγ, pSTAT3, pSTAT5, pSyk, and actin.
β2 integrins induce proliferation of AML cells by the activation of Syk. (A-E) Shown are growth curves of FFM05 and FFM12 cells that were treated with shRNAs against (A) Mac-1, (C) CD18, and (E) FcRγ (all shown as gray lines) or the respective control shRNAs (black lines). The shRNA-treated FFM05/12 cells were cultured on confluent layers of KM-102 bone marrow–derived stroma cells. Proliferation was monitored by XTT-based assay. After 48 hours of AML/stroma cell coculture, FFM05/12 cells were harvested, and lysates of FFM05 and FFM12 cells that were treated with (B, D, F) control shRNAs (lane 1) or (B) Mac-1–, (D) CD18-, or (F) FcRγ-specific shRNAs (all shown in lane 2) were subjected to immunoblot analyses using antibodies against Mac-1, CD18, FcRγ, pSTAT3, pSTAT5, pSyk, and actin.
Discussion
Syk is expressed in more than 80% of AML cases, and its inhibition causes a strong inhibition of AML cell growth and can induce apoptosis through a yet-undefined molecular mechanism.4 In this study, we elucidated the oncogenic Syk signaling network in AML cells by proteomic techniques. AML-specific interactome analysis revealed that, apart from proteins that are known interaction partners of Syk in B cells, other proteins complexed with Syk, like the transcription factors STAT3 and STAT5. Activation of both proteins is frequently found in AML cells and has been shown to potentially represent one of the key mechanisms in AML initiation and progression.15,28 Interestingly, STAT5 activation seems to be highly important for hematopoietic stem cell maintenance, both in normal and leukemic hematopoiesis.34
In the majority of AML cases, the upstream mediators inducing STAT3/5 activation are not known. We and others have shown that FLT3-ITD aberrantly and effectively activates STAT5 in AML blasts.1,35 Also, we showed that STAT5 targets like the Pim kinases are important mediators of transformation and collaborate with other oncogenes to induce AML in animal models.36 In 2011, it was shown that small molecule-mediated inhibition of STAT3 may effectively reduce AML cell survival and proliferation.37 However, constitutive STAT activation is not restricted to AML cells expressing FLT3-ITD.18 Our data show that Syk directly interacts with these STAT proteins and induces their phosphorylation at the decisive tyrosines needed for their activation. Because Syk is readily druggable, these findings raise immediate hypotheses for possible therapeutic strategies involving Syk kinase inhibitors in AML.
In B cells, Syk interactors have been extensively studied.26 Interestingly, no interaction between Syk and STAT3 or STAT5 was detected in B cells, although they express STAT3 and STAT5 proteins at significant levels.26 In contrast, a recent study showed that Syk phosphorylates STAT3 in acute lymphoblastic leukemia cells when they are exposed to oxidative stress.38 Thus it appears that different posttranslational modification patterns of Syk are 1 reason for differences in the composition of the Syk interactomes in different cell types. We found that the Syk phosphorylation state in AML cells differs from the B-lymphoid one26 ; this phenomenon of differential cell type–specific phosphorylation might be 1 possible explanation for the existence of the Syk/STAT complex in AML cells. Using a proteomic approach, we identified the β2 integrin receptor Mac-1 and FcγRI as Syk binding partners. Both are known regulators of Syk in other cell types, and it was reported that integrin receptors can activate Syk via the ITAM-bearing Fc-γ chain,32 which was originally identified as the signal transducer element of several Fc receptors including FcγRI.39 We show here that stimulation of either Mac-1 or FcγRI is sufficient to activate a Syk/STAT signaling axis in AML cells. In vivo, AML cells interact with both stromal cells and the ECM of the bone marrow niche, a scenario that has emerged as an important mechanism of AML cell survival and chemoresistance and that is still poorly characterized on the molecular level.40,41 To elucidate whether Mac-1 and FcγRI are relevant to promoting AML cell growth, we used an AML cell/stroma coculture model to mimic cellular interactions that are present in the bone marrow niche. Using this experimental setup, we observed that β2 integrins including Mac-1 trigger proliferation, while FcγRI did not influence the proliferation rate of AML cells.
Flow cytometric expression analyses in the past revealed that both Mac-1 and FcγRI are widely expressed in the primary AML cells of different Fab types.42,43 Interestingly, several studies reported that the expression of Mac-1 in AML is associated with a poor prognosis,42,44 which might be, at least in part, caused by Mac-1–mediated activation of the proproliferative Syk/STAT3/STAT5 pathways.
Taken together, our AML-specific characterization of the Syk signaling network identified apart from known signaling intermediates45-47 novel signaling pathways that promote AML cell growth and might therefore serve as therapeutic targets in the future.
The online version of this article contains a data supplement.
The publication costs of this article were defrayed in part by page charge payment. Therefore, and solely to indicate this fact, this article is hereby marked “advertisement” in accordance with 18 USC section 1734.
Acknowledgments
The authors thank Lisa Neuenroth, Monika Raabe, Astrid Eichler, Anja Vogel, and Uwe Plessman for their technical support. The authors also thank Richard Moriggl and Christian Wichmann for providing valuable reagents and Claudia Haferlach for the genetic characterization of AML cells.
This work was supported by the Landes-Offensive zur Entwicklung Wissenschaftlich-ökonomischer Exzellenz (LOEWE) initiative Oncogenic Signaling Frankfurt.
Authorship
Contribution: T.O. designed and supervised experiments, performed proteomic experiments, and wrote the manuscript; M.F.O. contributed to immunoblot analysis; M.E. performed in vitro kinase assays; H.-H.H. and J.C. contributed to proteomic analysis; S. Münch, S. Mohr, M.N., J.Z., and H.B. established and analyzed AML knockdown cells and contributed to biochemical analysis; T.B. performed flow cytometric analysis; G.B. contributed the primary AML cell cultures; M.A.R., J.W., and C.B. provided reagents; H.U. supervised proteomic analysis; and H.S. designed experiments, supervised the project, and wrote the paper.
Conflict-of-interest disclosure: The authors declare no competing financial interests.
Correspondence: Hubert Serve, Department of Hematology/Oncology, Johann Wolfgang Goethe University, Theodor-Stern-Kai 7, 60590 Frankfurt am Main, Germany; e-mail: serve@em.uni-frankfurt.de.